Lus10004018 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10004018 pacid=23153351 polypeptide=Lus10004018 locus=Lus10004018.g ID=Lus10004018.BGIv1.0 annot-version=v1.0
ATGGGCGAGTATGAAACTGCGGAGTCACAGCCAGCAAGTGTGCTCTTGCCCAACGGGTTACTGCCTGATGAAGCAGCCTCCGTGATGCAAGTGCTTGACT
CGGCCAGATGGTCGAAAGCTGAAGAGAGAACTGCTGAACTCATAGAGTGCATTAAGCCAAATGAACGTTCGGAGAATCAACGTAAGGCCGTGGCATCTTA
TGTGCAGCAGCTCATTGCCAAGTGCGTTCCTTGCCAGGTTTTCACTTTCGGGTCTGTTCCGCTTAAAACTTATTTGCCAGATGGAGATATTGACTTGACA
GCCTTCAGTAAGAATCAGACTTTGAAAGAGACTTGGGCTCATCAGGTTAGGGATATGTTGGAGAACGAGGAGAAGAATGAGAATGCCGAATTTTGTGTAA
AGGAAGTGCAGTACATTCAGGCTGAGGTGAAACTGGTAAAGTGCCTTGTGGAGAACATCGTCGTGGATATATCGTTTAACCAGCTTGGTGGGCTATGCAC
GCTCTGTTTCCTTGAAGAGGTTGATCATAAGATAGGAAATGATCATTTATTCAAGCGGAGCATTATTCTTATAAAGGCGTGGTGTTATTATGAGAGCCGT
ATATTGGGTGCTCACCATGGTCTGATATCCACTTATGCGTTGGAAACTCTGGTTCTCTATATCTTTCATGTTTTCAACAATTCCTTTGCTGGACCACTTG
AGGTCCTCTATCGTTTCCTTGAGTTCTTCAGTAAATTTGATTGGGATAACTTCTGTGTTAGCCTCTGGGGCCCTGTACCGATCAGTTCACTCCCAAACGT
GACAGCTGAAGCACCCCGAAAGGATGATGGAGATTTACTACTCGAAAAATCATTTCTTGATGCCTGTGGCTTGGCATATGCAGTTTTTCCCAATAGTCAA
GAAAATAACGGCTCCCCGTTCATCTCCAAGCACTTCAATGTCATTGATCCTTTGCGTGTTAACAACAATCTGGGTCGCAGTGTCAGCAAAGGTAACTTCT
TCAGGATTCGTAGTGCATTTGCATTTGGGGCAAAGAAGCTTGCTAGGTTACTTGAATGTGACAGCGAGAACATACACGCTGAAGTGAATCAGTTTTTCCA
GAATACCTGGGAAAGACATGGAAGTGGTCACCGTCCTGATGCTCCAATCAATGCTTTGGGGCAGCTGATTGGGTCAAGGAATGAGAATGTCATAAACACT
TCAAATCGAGAAAGTCAAAATGATATATCACAGCATGGTAGTCGTACCTTAGAAAGCACGCCACAGAGTAGTGATATTGCTACAGCATTTCGAACACACA
TTCAAACGATGAACAACTCCAGTTTGAGGATATCTGACGAGTCTAGACCAGAAATCAATTCAGATAATGGTGTTCACGCTGAAAATGGTCCTAGAACTTC
TAAACCTGACAGTATGTTGAGCGGTTTACAAGGAAGGTATATATTTGCAAGGACACGCTCAAGTCCTGAGCTTACTGATACCTATAATGAAGCATCCTCA
CAAACGAGACAGAATAGAGCACCAGAAGCAAAGGGTGAGGCCTCGGCAGTGATGTTGGACAGCATCAAGAGACAAAACACTGAACATGAAAAGTCATCTA
GGCATGGTACTGGATCTTTAGAAGATGATCCTTCTGTTACACGCAGCTCAATGCACCAGAATCTTGATGCTGTAAGTAACTATAGTGATGATTCAGCCCT
TTTTGCTGCAAGTGAGGAATTCACTTCTATCCTGGGTACACAAGGGATGTATCAGGATGTCGTGAACATGATGGCTTCTTCGAGTGGACATGGCTTTAAT
GGACAAGTCCATCTTCCCCTGAATTTAGCTTCTGGTCAGATGCCCTTTCCCATACCACCTTCCGCTCTAGCTTCTATGGGATATTTTCAACGGAATTTGG
GTGGAATGGTACCACCAAATACTTCATTGTTGGAGACGCATTGGGGTCCAAGTATGCAATTTCCGCAGGGTTTGTTTCCTTCTCAGTCGAATCACTTTTT
TCCCACGGTAGGATTGGCATCAAGTCCAGAAGGTTCAGTTGAACATTCAGATGAGAACTTAAGTTCTTTGGAATCAAATGTTGGGGTGGCTGGCAACAAT
TTCTTGCATGAACATGAAAGGGCCTCTATTGATGGTCTTGAACTTTATAATGCCAATTCCCTGATGCATCAGTCTGACGAGAAACATCAATCAAATAGTA
ATCACTTCGTTCCTTTGTCTAGAACAGGTGGCTCTAGGAACTCTTCTAGTGTTCAACCAAGGGAGTCTCGACATTCAAGTAGGCGGGATCATATGGAAAC
TTATCCTTACTCGGAAAACAGAGGTACCGAAGTGCTCACTGAGGAAAACCTTGCTAGCTCAAGGCCTTCGTCAAGTAATTCTTGGAGAAGTAAGCCTGCA
TCTGAGAGCTCATCTGATGGATTATCTGCAAAGACAAAGACAATGAGGGAAAAGAGAGGGAGGAAGACTTCGACAGCTGTTGCTTATGGGATAGGCAGGG
GAATCAATGAGAATGGCTCCCTTCACATGGATGATGACACGCAGCAGCTCTCCTCAAGCCCTGAGACTGTTGAAAGGGGCTTCGGGCCTCCGGCTCCTAG
TTCTTTGCATGTTTCAAGGCATAATAATGTGCCTCGATTTGATCTGGCGCAGACAGGTGAATCTGATTCATTAGCATCCATTCCTCCGGTGCAATTAGGT
CATGGTGCACGGTTTTATCCTACGGGGCCACCTGTTCCATTTGTTACGATGCTACCCATTTACAACTTCCCAACAGAGAGGGGAATTTCTGAAGCATCAA
CTAGCCAGCATAGCTTTGAGGAGATCATAGATAACAATGTTCCACGTCAGAATGGTTATTCAATGAGCGTTTCGGTCATTGATGAGCCATTTGAACACAA
GTTTGACATCCTCAATAGCGATTTTGCTAGTCATTGGCAGAACTTGCAATTTGGAAGGTTTTGCCAAGATCCTCCGTCGCTCTCACCGGTGACCAGTCCT
TCACCGGTTGTTGTACCGCCGGTGTATTTACAGGGCCGTTTCCCATGGGAAGGTCCTGGAAGGCCCATTTCACCTAATATGAATCTTTTCTCTCAGTTGA
TGGGTTTCGGGCCTCGTATCCTTCCAGTTGCTCCTGTTCCAGCTGTATCAAATAGACCTGCTGCCGTCTACCACCAGCCTTTCGTTGATGAGGTGCCTAG
ATACCGCGGTGGAACTGGAACCTACCTTCCAAATCCTAATGTGGCTGTCCGAGATACTCCAAACTCGAGAAAAACAAACTATAACTTTGATAGAAGTGAC
TACCATGGTGATCGGGAGGGGAACTGGAATAATTCAAAGTCGAGAGCCCCCGGGCGCAGTCAGAGTCGTAACCAGGCAGATAAGTCGAGATCTGACCGGC
CATGGGTTTCACACAGACACGACACTTCCGTTACTTATCAGTCTCAAAACGGGCCAAGCCGAACAAATGCCTCTCTAGTTGGTTCTGCGGCCAATACGAA
TTATGGCATGTATCCAATAACGTCTATGAATTCGAATGGACAGGTTGTCATGATGTATCCTTTTGACCAAAACGCTGCTGGTTATGTCTCTCCAGCGGAC
CAGATTGAGTTTGGCTCGTTTGTTCCTCCCGGAAACTCTGGTGTAAATGATCTTAATGAGGGAAGTAGACCCGGCGAGGGATTCGAGGACCAAAGATTTC
ACAGTAGTGCAGCAGTTCAACTGTCTTCCCCGGATCCGCCTTCTTCTCCTCGCTTTCAGAGCTGCGCCAAGAATCCCGAGTTGAAGGACTCGGTGGGAAT
CCTACCGCTGGCTGCGGGGACCAGGTCGGTTACACATATCTCGGTTGGCCAGAACAAAGAACTTTGGTTCGATTTGATTCAGCTCTCAGATCAGCCTTCC
ATGATAGCTGCCAGTTCTTGCGAATTAAGCAGAAGAAAGAGACGAAACTCGGGCGATGAGCCATTGCAGGACTTGGGTAACTAG
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10004018 pacid=23153351 polypeptide=Lus10004018 locus=Lus10004018.g ID=Lus10004018.BGIv1.0 annot-version=v1.0
MGEYETAESQPASVLLPNGLLPDEAASVMQVLDSARWSKAEERTAELIECIKPNERSENQRKAVASYVQQLIAKCVPCQVFTFGSVPLKTYLPDGDIDLT
AFSKNQTLKETWAHQVRDMLENEEKNENAEFCVKEVQYIQAEVKLVKCLVENIVVDISFNQLGGLCTLCFLEEVDHKIGNDHLFKRSIILIKAWCYYESR
ILGAHHGLISTYALETLVLYIFHVFNNSFAGPLEVLYRFLEFFSKFDWDNFCVSLWGPVPISSLPNVTAEAPRKDDGDLLLEKSFLDACGLAYAVFPNSQ
ENNGSPFISKHFNVIDPLRVNNNLGRSVSKGNFFRIRSAFAFGAKKLARLLECDSENIHAEVNQFFQNTWERHGSGHRPDAPINALGQLIGSRNENVINT
SNRESQNDISQHGSRTLESTPQSSDIATAFRTHIQTMNNSSLRISDESRPEINSDNGVHAENGPRTSKPDSMLSGLQGRYIFARTRSSPELTDTYNEASS
QTRQNRAPEAKGEASAVMLDSIKRQNTEHEKSSRHGTGSLEDDPSVTRSSMHQNLDAVSNYSDDSALFAASEEFTSILGTQGMYQDVVNMMASSSGHGFN
GQVHLPLNLASGQMPFPIPPSALASMGYFQRNLGGMVPPNTSLLETHWGPSMQFPQGLFPSQSNHFFPTVGLASSPEGSVEHSDENLSSLESNVGVAGNN
FLHEHERASIDGLELYNANSLMHQSDEKHQSNSNHFVPLSRTGGSRNSSSVQPRESRHSSRRDHMETYPYSENRGTEVLTEENLASSRPSSSNSWRSKPA
SESSSDGLSAKTKTMREKRGRKTSTAVAYGIGRGINENGSLHMDDDTQQLSSSPETVERGFGPPAPSSLHVSRHNNVPRFDLAQTGESDSLASIPPVQLG
HGARFYPTGPPVPFVTMLPIYNFPTERGISEASTSQHSFEEIIDNNVPRQNGYSMSVSVIDEPFEHKFDILNSDFASHWQNLQFGRFCQDPPSLSPVTSP
SPVVVPPVYLQGRFPWEGPGRPISPNMNLFSQLMGFGPRILPVAPVPAVSNRPAAVYHQPFVDEVPRYRGGTGTYLPNPNVAVRDTPNSRKTNYNFDRSD
YHGDREGNWNNSKSRAPGRSQSRNQADKSRSDRPWVSHRHDTSVTYQSQNGPSRTNASLVGSAANTNYGMYPITSMNSNGQVVMMYPFDQNAAGYVSPAD
QIEFGSFVPPGNSGVNDLNEGSRPGEGFEDQRFHSSAAVQLSSPDPPSSPRFQSCAKNPELKDSVGILPLAAGTRSVTHISVGQNKELWFDLIQLSDQPS
MIAASSCELSRRKRRNSGDEPLQDLGN
|
DESeq2's median of ratios [FLAX]
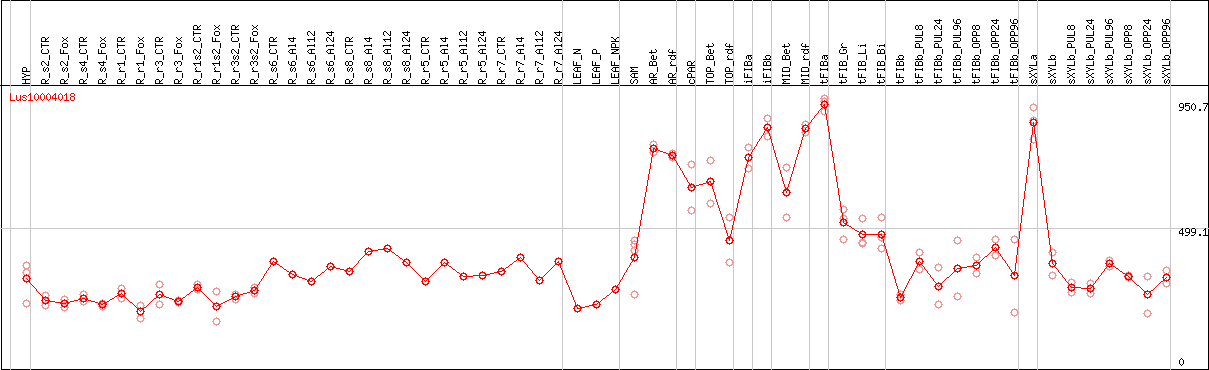
Coexpressed genes
Lus10004018 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
