Lus10004153 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10004153 pacid=23153304 polypeptide=Lus10004153 locus=Lus10004153.g ID=Lus10004153.BGIv1.0 annot-version=v1.0
ATGGGGGACACGGATAGCTGTAGCAGCAGAGCCGTGGAGTTCGTTCCGGCTATGAATAGAAATCGGAGGCAAAAAGTAGAGGTTTACAATGAAGTTCTCC
GCCGGCTCAGGAATTCCAATGACCAAGAAGCTAGTCTTCCCGGCTTCGAAGACGAGCTCTGGAATCATTTCCTTCGGCTTCCTACTAGATATGCACTGGA
AGTGAACGCAGAAAGGGCACATGACGTCCTTATGCACAAGAGATTGCTGCAGATGGCTCATAATCCTGCTACTAGGCCGGCAATTGAGGTCCGGCTCGTG
CAGCAATGGCAGTTTGCTTTGGACTCGGTTGCTTTTAAGGTTCAAATTGTTTTTATTCAAGCTGTTTTCACGCAGGTTTATTCTGCAGCCCATGGGCATT
ATGGTGACTCTAGTGAAACTGAATTTTCTATGGAAAAGCTATCTCAATTTTCTGATCAACCCTGGAAGCAGAGGCATGGATGGGTTTATCCACATACTGT
TAAGGTTTCATTTGCTTTCAGGGTAGTTCTAATTAAAAAGCTGACATGTTTGGACTCTTCCTCTGATTTCAGTCTCCATCCACCACCTGCCTTTGGTTCA
TTGCCTGACCTAGACCTTGCATATGGAGCTCAAGATAAGGATACTGGCCATGCAATTTATTCCCGCCTTATGCACGAGATTACAATATCCACAAGTGATA
AGCCAAAGCTTCTCAGTCAGTTGACTTCCTTGCTCTCTGAGATTGAGCTAAATATACAAGAAGCGCTTGCTTTTTCTACAGCTGATGGGTACTCATTAGA
TGTGTTTGTTGTTGACAATTGGAGTCCTGAGGAGTTGGAGCAACTTAGGGTAGTGCTGGAGCGTGAAATACCAAAGATTGAGCAAGACCAAAGAGGAACC
AATTATAGATCTTCTGATATGAGCATTCCTACGGCGTCAATCGATGTTTGGGAAATAGATGCAACATTATTGAAGTATGAATTCAGAATTGCATCTGGAC
CCTATGGTGACATGTATAAAGGTACCTTCTGTGGTCAAGATGTGGCTATTAAGCACTTCAGGGCGGAGCACCTAAGTGGTAAACTTCGAAAGGAATTTGC
TCAAGAAATCTTTATTCTGAGGAAAGTCCGGCACAAGAATGTTGTTCAATTTGTCGGTGCATGTACCAGACCTCCAAACCTCTGTATCCTGACAGAATTC
ATGTCTGGTGGAAGCATGTACGACTATCTGCATAAAAGAAAGGAAATGTTTACATGTCAGTCCTTGGTTAAAGTGGCATCTGATGTATCCAGGGGAATGA
ACTACCTGCACAGAAATAGTATAATACACAGAGACCTGAAAGCTGCCAATCTTCTGATGGATGAAAATGGAATAGTCAAGGTGGCTGATTTTGGTGTTGC
CAGAATGCAAGATCATTCAGGTGTGATGACTGCAGAAACTGGAACGTACCGTTGGATGGCTCCAGAGGTGATGGCGGTGGAGGCAGAGCTCGTGTTAAGT
GACGGCGGCAGAGGAGACTGCTGTCGAGAAGTACCAGAAAAGATGAAGGTTATTGGACACAAGCCTTACAATCATAAAGCTGATGTATTCAGCTTTGGTG
TTGTGTTGTGGGAATTGCTTACAGGGAAGATTCCTTATGAAAACTTAACCCCATTACAAGCTGCAGTTGGTGTTGTACAACAGGGCCTGAGACCATCCGT
TCCGAGGAATACCCATCCCAAAGTGGTTGAACTTCTTGAGAGATGCTGGGACCAGGACCCATGTTTGAGGCCTGAATTCTCTGAAATTTTGGAACTCCTA
CAAAATTTACAGAAGATGAGGCGCACTTGTGCAAGGAACGTCGGTGGAGGCTCTGATCACATGGACACTGAAGCATCTACAAGGAAAAGGTACTTGGATG
TTAGATTATTTGGACTTAGACAAGAGGGCCTTTCTACAATGGAAAGTCTGAGAAGCTCCTTGAAATCTTGCTGTTCTCATGCCAACCTGCAACGACAAGA
TTATCTTGACCAAGAAAAAAAGGAAGCTCTCTTGAACCAGATCGACCGCAATTCCCATCCGGTCAGGATATCTAATTTTCCTGGCAAGGAAGACCAGTTG
GTAGTTGACATCCATAATGCCACGCCCTGCAAAGGAACAAAGGAGACTGCTTCAAAGCACTCTAATCCGGCAAAGCCTAAGCTTTCCTTTGCTGAAATTG
CAGAGGAAGCTATGTGGTTCAAATCAGATAAAGAAGAAAGCCAACCATCACAGAATGATCCTTTCCGCCGTAACCCATGGAGGTCACTTATGATCAGAAC
AAGGTCCAGACTAATCGAAGACCTTCCAGATGAAGAATACATCACAAGTGATTCTGAGAAAGAAGAAGTTTTAGAAGAAGAAGATGACCATGATGAAGAT
GTCTCATACAAGAACAGAGGAGGCAACACTGTCAGAATGCTACTTTATCTGGCTCTTATCTCTACTCTTGCCGCCTTAATTTGCACGCTTTGCGTCTCTG
CAATGAGGAAGATAACACTCTGGGACCTTCCACTGTGGAAATGGGAGATAATGGTTTTGGCACTCATGGGTGGCAATTTGGTCTCCAGTTGTGGGGTTAA
GCTCGTCGTGATCCTCATTGAGCGTAGCTTCCTTCTGCAGAAACGAGTTCTGTACTTCGTTTACGGGCTGAAGAAGCCAGTTCAGAACTGTATCTGGCTG
GGTTTGGTTTTGCTCGTTTGGCACTGGGTTTTCAATGACAAGATCAAGGAACAGACTCATAGTATAATATTGCCTTATGGGACCAAGATTTTAGCTTGTC
TCTTGGTTGGAACGTTGATCTGGCTGGTGAAGACCCTCTTTGTCAAAGGTCTTGCTTCTTCTTTCCACGAGAACACTTTCTTTGAACGAATCCAGGTGGC
TCTCTTCAACCAGTACGTAATCGAGACACTTTCTGGTCCACCAGTCTTTGAAGTCGAAAGTGAATCGAGCAGTAAAGAGACGAAGCTACAAAAGTGCTCC
ACCATCGGTGGAAAGTTGTCAAGGTTGTCTAGGACAGTTTCTAGCAGAAGGAATGAAGAGATCACAATAGACGAGCTGCAGACTCTGAGCAAGAAGAACA
TTTCGGCTTGGACAATGAGAAGAATGATCAACATCGTTCGCAAAGGTTCATTGTCTACTCTGGATGAACACATACTTGATTCAGACATGAAAGACGAGTC
TTCGTTGCACATCAGGAGTGTTTGCCAAGCAAAGGAAGCAGCCAAGAAAATATTTGCTAATGTGGCTGCTCCAGGATTCCAGCATATCTTCATTGCCGAC
TTGCTGCGCTTCATGGCTAAAGAAGAGGCTTTGAGAGCCATGCATCACTTTGGACCAGGATCTGAGAATGGAGGGATTAGCAGATCATTACTCACTGAGT
GGCTGGTTAATGCATTCAGGGACAGGCAAGCGCTTGCACTGTCACTTAACGACACGAAAACAGCTGTCGACGAACTCCACAACATGCTCAACTTCATTGT
AGCAGTTATCATCATAATAATCTGGCTTGTCATGCTGGGAATCCCCATTACTCACTTCCTGGTTCTCTTAAGCTCACAGCTTCTTCTGGTGGCTTTCATT
TTTGGCAATTCCTGCAAAACGACCTTTGAAGCCATCATTTTTTTGTTCGTCATGCACCCTTTCGATGTCGGAGACCTTTGTGAAGTTGATGGAACCCAGA
TGAGAGTGGAGGAGATGAACATCTTAACCACAGTGTTCCTCAAGTCCAATTACCAGAAGATAACATTCCCAAACAGTGTTCTCTCCACCAAACCGATCTC
CAATTTCTACCGCAGCCCAGATATGACGGAGGCAATCCACTTTTGTGTCCACATCTCAACTCCAATGGACAAGATCTCCGCTATGAACGACCGGCTAAAA
GCGTACGTGGAGCGGCATAAGCAGTACTGGTACCCAACGGTGTCGGTGTTTATAAATGAGGTGGAAGACATGAACAAATTGCAGATGCAAGTGGTTGCGA
GGCACAGGATGAACTTCCAGAATATGGGAGATAGATTTACGAGGAGGGGGCATCTGATTAAAGAACTCATCACGATGTTTCGAGACTTAGAACTCGAGTA
CCGAATGCTTCCACTTGATGTCAACGTGCGAAATCTGCCGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10004153 pacid=23153304 polypeptide=Lus10004153 locus=Lus10004153.g ID=Lus10004153.BGIv1.0 annot-version=v1.0
MGDTDSCSSRAVEFVPAMNRNRRQKVEVYNEVLRRLRNSNDQEASLPGFEDELWNHFLRLPTRYALEVNAERAHDVLMHKRLLQMAHNPATRPAIEVRLV
QQWQFALDSVAFKVQIVFIQAVFTQVYSAAHGHYGDSSETEFSMEKLSQFSDQPWKQRHGWVYPHTVKVSFAFRVVLIKKLTCLDSSSDFSLHPPPAFGS
LPDLDLAYGAQDKDTGHAIYSRLMHEITISTSDKPKLLSQLTSLLSEIELNIQEALAFSTADGYSLDVFVVDNWSPEELEQLRVVLEREIPKIEQDQRGT
NYRSSDMSIPTASIDVWEIDATLLKYEFRIASGPYGDMYKGTFCGQDVAIKHFRAEHLSGKLRKEFAQEIFILRKVRHKNVVQFVGACTRPPNLCILTEF
MSGGSMYDYLHKRKEMFTCQSLVKVASDVSRGMNYLHRNSIIHRDLKAANLLMDENGIVKVADFGVARMQDHSGVMTAETGTYRWMAPEVMAVEAELVLS
DGGRGDCCREVPEKMKVIGHKPYNHKADVFSFGVVLWELLTGKIPYENLTPLQAAVGVVQQGLRPSVPRNTHPKVVELLERCWDQDPCLRPEFSEILELL
QNLQKMRRTCARNVGGGSDHMDTEASTRKRYLDVRLFGLRQEGLSTMESLRSSLKSCCSHANLQRQDYLDQEKKEALLNQIDRNSHPVRISNFPGKEDQL
VVDIHNATPCKGTKETASKHSNPAKPKLSFAEIAEEAMWFKSDKEESQPSQNDPFRRNPWRSLMIRTRSRLIEDLPDEEYITSDSEKEEVLEEEDDHDED
VSYKNRGGNTVRMLLYLALISTLAALICTLCVSAMRKITLWDLPLWKWEIMVLALMGGNLVSSCGVKLVVILIERSFLLQKRVLYFVYGLKKPVQNCIWL
GLVLLVWHWVFNDKIKEQTHSIILPYGTKILACLLVGTLIWLVKTLFVKGLASSFHENTFFERIQVALFNQYVIETLSGPPVFEVESESSSKETKLQKCS
TIGGKLSRLSRTVSSRRNEEITIDELQTLSKKNISAWTMRRMINIVRKGSLSTLDEHILDSDMKDESSLHIRSVCQAKEAAKKIFANVAAPGFQHIFIAD
LLRFMAKEEALRAMHHFGPGSENGGISRSLLTEWLVNAFRDRQALALSLNDTKTAVDELHNMLNFIVAVIIIIIWLVMLGIPITHFLVLLSSQLLLVAFI
FGNSCKTTFEAIIFLFVMHPFDVGDLCEVDGTQMRVEEMNILTTVFLKSNYQKITFPNSVLSTKPISNFYRSPDMTEAIHFCVHISTPMDKISAMNDRLK
AYVERHKQYWYPTVSVFINEVEDMNKLQMQVVARHRMNFQNMGDRFTRRGHLIKELITMFRDLELEYRMLPLDVNVRNLP
|
DESeq2's median of ratios [FLAX]
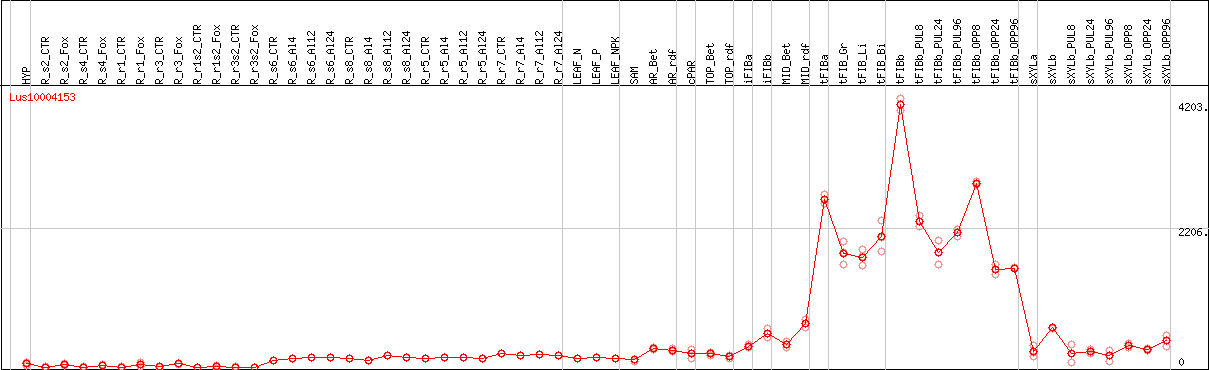
Coexpressed genes
Lus10004153 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
