External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G62500 114 / 1e-30
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
AT2G10940 92 / 5e-22
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1.2)
AT3G22142 82 / 5e-18
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
AT3G22120 77 / 1e-16
CWLP
cell wall-plasma membrane linker protein (.1)
AT4G15160 75 / 1e-16
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1.2)
AT1G12100 64 / 4e-13
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
AT4G12470 62 / 5e-12
AZI1
azelaic acid induced 1 (.1)
AT4G12510 61 / 5e-12
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
AT4G12520 61 / 5e-12
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
AT1G12090 60 / 2e-11
ELP
extensin-like protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10028930
134 / 8e-39
AT1G62500 141 / 4e-41
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Lus10032262
121 / 2e-33
AT1G62500 162 / 3e-48
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Lus10010482
82 / 2e-20
AT3G22142 162 / 5e-48
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Lus10003801
84 / 3e-19
AT3G22142 168 / 7e-47
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Lus10027704
82 / 3e-19
AT2G10940 135 / 2e-39
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1.2)
Lus10001493
69 / 8e-15
AT1G62510 111 / 2e-32
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Lus10032259
68 / 2e-14
AT4G12520 133 / 2e-41
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Lus10004346
66 / 9e-14
AT4G12520 161 / 2e-52
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Lus10024625
65 / 2e-13
AT4G12520 158 / 5e-51
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G111300
121 / 1e-33
AT1G62500 125 / 2e-34
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Potri.006G065500
98 / 4e-25
AT2G10940 147 / 8e-44
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1.2)
Potri.018G126000
98 / 6e-24
AT2G10940 109 / 2e-27
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1.2)
Potri.006G008500
84 / 5e-20
AT3G22120 80 / 6e-18
cell wall-plasma membrane linker protein (.1)
Potri.016G015500
87 / 9e-20
AT3G22142 161 / 1e-42
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Potri.006G008300
82 / 8e-19
AT3G22142 113 / 2e-28
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Potri.018G025900
66 / 6e-14
AT2G45180 127 / 8e-39
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Potri.001G121900
67 / 8e-14
AT1G62510 99 / 5e-27
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Potri.001G121800
66 / 1e-13
AT1G12100 143 / 5e-45
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
Potri.003G111400
65 / 3e-13
AT1G62510 93 / 7e-25
Bifunctional inhibitor/lipid-transfer protein/seed storage 2S albumin superfamily protein (.1)
PFAM info
Representative CDS sequence
>Lus10004349 pacid=23161718 polypeptide=Lus10004349 locus=Lus10004349.g ID=Lus10004349.BGIv1.0 annot-version=v1.0
ATGGCTTCTTCTTCAAAACTAATAGCCTTGATATTCATCTCCATGCATTTCATTCTAATCTCCTTGCCTCCTATTTACTCTTGTGTCCCTTGCACTCACC
CAAAACCAAAACCAAACCCAAACCCACCCTCTCACTCTCACTCTCCTCCATCATCCGGCCACCGTCCACATCCCCCCACCACCAGCCCACCCCACCACGG
CGGGGGCCGCGGTGGTCACCATGGTAAAGGCAAAGGCAAAGGCAAAGGAGGTGGCGGAGGTAAAGGGCATAATCGTCCTCCACCAATAGTTTCACCACCC
ATCATAGTCAACCCTCCTCCGGTTACATACCCTCCTCCGGTGGTGACGCCTCCCGTGATAAACCCTCCACCTCCTTCTTCCGTGCCATGTCCTCCTCCGC
CAGCTACGCAACCCACGTGTCCGATCGGAGGGCTGAAGCTAGGAGCGTGCGTGGACGTGCTGGGAGGGTTGGTCCACGTAGGGGTAGGGAACCCGGTGGA
GAATGTGTGCTGTCCGGTGATTAAAGGGCTGCTGGAGATTGAAGCCGCCGTTTGTTTGTGCACGTCCATAAGGCTGAAGCTTCTTAACATCAACATCTTC
ATCCCTTTAGCACTTCAGGCTCTCATCACCTGCGGCAAGACTTCCCCCCCTGGCTTCGTTTGCCCTCCACTTTGA
AA sequence
>Lus10004349 pacid=23161718 polypeptide=Lus10004349 locus=Lus10004349.g ID=Lus10004349.BGIv1.0 annot-version=v1.0
MASSSKLIALIFISMHFILISLPPIYSCVPCTHPKPKPNPNPPSHSHSPPSSGHRPHPPTTSPPHHGGGRGGHHGKGKGKGKGGGGGKGHNRPPPIVSPP
IIVNPPPVTYPPPVVTPPVINPPPPSSVPCPPPPATQPTCPIGGLKLGACVDVLGGLVHVGVGNPVENVCCPVIKGLLEIEAAVCLCTSIRLKLLNINIF
IPLALQALITCGKTSPPGFVCPPL
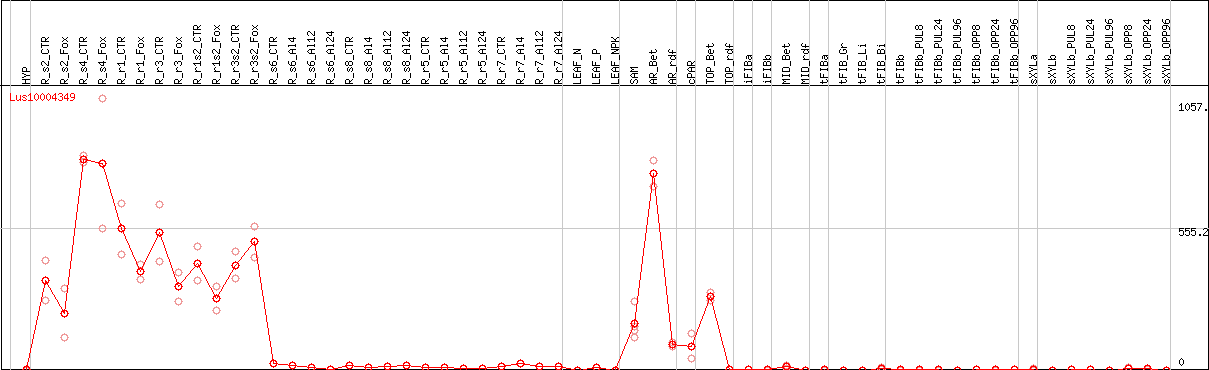
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10004349 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.