External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G45170 179 / 4e-59
ATATG8E
AUTOPHAGY 8E (.1.2)
AT4G16520 177 / 2e-58
ATG8F
autophagy 8f, Ubiquitin-like superfamily protein (.1.2)
AT1G62040 174 / 4e-57
ATG8C
autophagy 8c, Ubiquitin-like superfamily protein (.1.2)
AT3G60640 171 / 5e-56
ATG8G
AUTOPHAGY 8G, Ubiquitin-like superfamily protein (.1)
AT2G05630 169 / 3e-55
ATG8D
Ubiquitin-like superfamily protein (.1.2)
AT4G21980 167 / 3e-54
ATG8A, APG8A
AUTOPHAGY-RELATED 8A, AUTOPHAGY 8A, Ubiquitin-like superfamily protein (.1.2)
AT4G04620 163 / 4e-53
ATG8B
autophagy 8b, Ubiquitin-like superfamily protein (.1.2)
AT3G15580 121 / 2e-36
APG8H, ATG8I
AUTOPHAGY 8I, AUTOPHAGY 8H, Ubiquitin-like superfamily protein (.1)
AT3G06420 116 / 2e-34
ATG8H
autophagy 8h, Ubiquitin-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10028933
197 / 4e-66
AT4G16520 215 / 6e-74
autophagy 8f, Ubiquitin-like superfamily protein (.1.2)
Lus10008507
185 / 1e-61
AT4G16520 216 / 3e-74
autophagy 8f, Ubiquitin-like superfamily protein (.1.2)
Lus10000733
183 / 7e-61
AT4G16520 217 / 1e-74
autophagy 8f, Ubiquitin-like superfamily protein (.1.2)
Lus10015563
172 / 3e-56
AT1G62040 230 / 1e-79
autophagy 8c, Ubiquitin-like superfamily protein (.1.2)
Lus10027186
168 / 6e-55
AT2G05630 222 / 1e-76
Ubiquitin-like superfamily protein (.1.2)
Lus10039656
164 / 2e-52
AT2G05630 203 / 6e-68
Ubiquitin-like superfamily protein (.1.2)
Lus10038046
120 / 9e-36
AT3G15580 199 / 1e-67
AUTOPHAGY 8I, AUTOPHAGY 8H, Ubiquitin-like superfamily protein (.1)
Lus10009987
112 / 2e-29
AT3G62240 625 / 0.0
RING/U-box superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G136040
187 / 2e-62
AT4G16520 197 / 1e-66
autophagy 8f, Ubiquitin-like superfamily protein (.1.2)
Potri.003G110901
187 / 2e-62
AT4G16520 197 / 1e-66
autophagy 8f, Ubiquitin-like superfamily protein (.1.2)
Potri.001G122700
186 / 4e-62
AT4G16520 191 / 2e-64
autophagy 8f, Ubiquitin-like superfamily protein (.1.2)
Potri.002G144600
181 / 4e-60
AT3G60640 192 / 8e-65
AUTOPHAGY 8G, Ubiquitin-like superfamily protein (.1)
Potri.014G060300
180 / 1e-59
AT4G16520 214 / 1e-73
autophagy 8f, Ubiquitin-like superfamily protein (.1.2)
Potri.002G228800
171 / 6e-56
AT2G05630 224 / 1e-77
Ubiquitin-like superfamily protein (.1.2)
Potri.014G153800
169 / 2e-55
AT2G05630 202 / 6e-69
Ubiquitin-like superfamily protein (.1.2)
Potri.004G013700
164 / 2e-53
AT1G62040 216 / 3e-74
autophagy 8c, Ubiquitin-like superfamily protein (.1.2)
Potri.011G004300
162 / 4e-52
AT4G21980 219 / 1e-74
AUTOPHAGY-RELATED 8A, AUTOPHAGY 8A, Ubiquitin-like superfamily protein (.1.2)
Potri.008G099400
121 / 2e-36
AT3G15580 187 / 7e-63
AUTOPHAGY 8I, AUTOPHAGY 8H, Ubiquitin-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0072
Ubiquitin
PF02991
Atg8
Autophagy protein Atg8 ubiquitin like
Representative CDS sequence
>Lus10004352 pacid=23161741 polypeptide=Lus10004352 locus=Lus10004352.g ID=Lus10004352.BGIv1.0 annot-version=v1.0
ATGCTCCCCTCTTCGAATTATAGTAGGGGTGACTCTAATAAAACTGAATCACTAGTCAGCTGGATGATTCAGCAGCTTGAGCACCAAGATAGAGAGAAGA
GAAAGGCTGAGGCTGCCAGGATAAGGGAGAAGTACCCAGATAGAATTCCGGTCATTGTGGAGAAGGCGGAGAGAAGTGAAATACCAAACATAGACAAGAA
AAAGTACCTAGTCCCATCTGACTTGACTGTCGGACAGTTTGTTTACGTAATCCGAAAGAGGATCAAATTGAGTGCTGAAAAGGCAATCTTCATATTCGTG
GACAATGTCCTGCCACCAACCGGTGCTATAATGTCTGCAATATATGAAGAAAAGAAGGATGAAGACGGATTTCTCTATGTTACTTACAGCGGCGAGAATA
CTTTCGGAGTGGAGATTCATCAGTAG
AA sequence
>Lus10004352 pacid=23161741 polypeptide=Lus10004352 locus=Lus10004352.g ID=Lus10004352.BGIv1.0 annot-version=v1.0
MLPSSNYSRGDSNKTESLVSWMIQQLEHQDREKRKAEAARIREKYPDRIPVIVEKAERSEIPNIDKKKYLVPSDLTVGQFVYVIRKRIKLSAEKAIFIFV
DNVLPPTGAIMSAIYEEKKDEDGFLYVTYSGENTFGVEIHQ
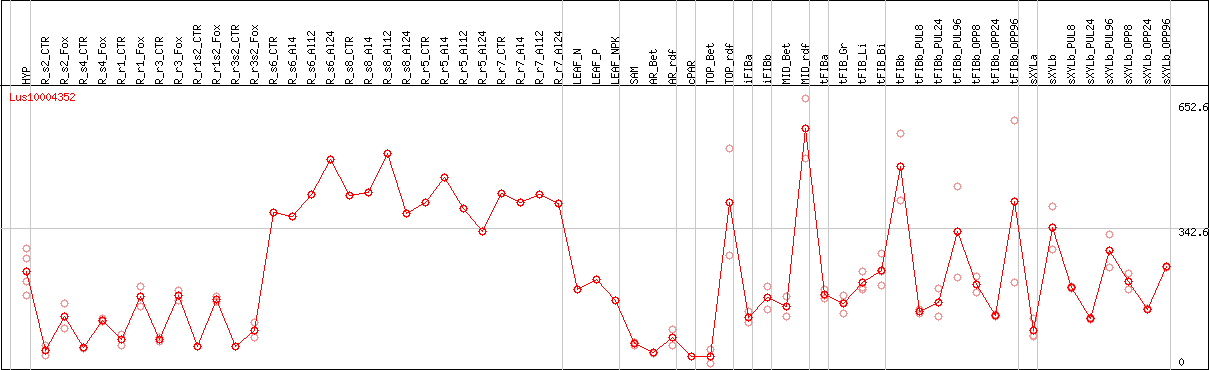
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10004352 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.