External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G36730 355 / 1e-120
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT2G29760 229 / 1e-69
OTP81
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G21065 220 / 2e-68
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT2G02980 222 / 6e-68
OTP85
ORGANELLE TRANSCRIPT PROCESSING 85, Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT5G56310 213 / 2e-65
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G13410 211 / 8e-65
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G11290 217 / 1e-64
CRR22
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G49170 216 / 5e-64
EMB2261
embryo defective 2261, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G20540 210 / 6e-64
MEF21
mitochondrial editing factor 21 (.1)
AT5G59200 207 / 9e-63
OTP80
ORGANELLE TRANSCRIPT PROCESSING 80, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10040171
513 / 0
AT2G36730 456 / 4e-159
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10040680
244 / 2e-75
AT2G29760 926 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10018223
238 / 5e-73
AT2G29760 931 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10031430
218 / 2e-68
AT5G06540 350 / 1e-115
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10010901
218 / 6e-68
AT3G15930 360 / 2e-118
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10010320
218 / 4e-66
AT2G34400 658 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Lus10028837
211 / 3e-65
AT1G11290 650 / 0.0
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10031989
217 / 4e-65
AT3G26782 865 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10025490
213 / 3e-64
AT2G33760 734 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.017G083900
412 / 5e-143
AT2G36730 552 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.013G103600
238 / 8e-74
AT1G08070 477 / 7e-160
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.004G125500
228 / 3e-70
AT3G29230 829 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.003G031600
229 / 8e-70
AT3G15930 784 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.010G168800
225 / 3e-69
AT2G02980 860 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 85, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.017G001000
224 / 2e-68
AT4G21065 793 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.003G223900
222 / 2e-68
AT5G56310 508 / 1e-176
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.015G034900
221 / 2e-68
AT4G21065 370 / 2e-122
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.005G012600
221 / 5e-67
AT2G29760 540 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.018G071800
220 / 1e-66
AT1G08070 606 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10004373 pacid=23151916 polypeptide=Lus10004373 locus=Lus10004373.g ID=Lus10004373.BGIv1.0 annot-version=v1.0
ATGAGGGGTTCAGGATTCGAACCGGACGAAGCGACCATGGTGGTCATGCTTTCGCTTTCGGCAGAAGTAGGCAACTTAGGGTTAGGGAAATGGATTCATT
CACAAGTGATTAAGGGGAGAATGGAATTGAATTACCAGCTGGGCACAGCCATTGTTGACATGTACGCAAAGAGTGGGAGTGTAGGATATGCAATGTCGGT
TTTCGGAAGCATGGAGAAGAGAAATGTGTGGACTTGGAGTGCTATGATTTTGGGGTTGGCTCAACACGGGTTCGCAAAGGAAGGGCTCGAACTATTTCAT
CAGATGATCGAGACGTCGTCTATCCGCCCTAATTACGTGACCTTCCTCGGAGTCCTTTGCGCTTGTAGCCATGCTGGTCTGGTAGAGGACGGGTTTCGAT
ACTTCCACGAAATGGAGCACCAGCACGGGATCAAACCCACGGTGGTGCACTACGGTGCAATGGTTGATATACTTGCTCGTGCAGGCCACCTGAGGGAAGC
TTACAACTTTGTAACACATATGCCTCACCAGAACGACCCGACAGTGTTAAGAACATTGTTAGGAGCTTGCAGCAACCGCAGCGAGAATGACGACGATGGA
GTTGCGGATAAGGTAAGGAAGCGCTTGCTGGAAGTGGAACCGAGGAGAAGTGGGAACCTTGTTATGGTTGCAAAGATGTATGCTGATGCAGGGATGTGGC
AGCGAGTTGCAGATGTAAGGAAGGTTATGAGGGATGGCGGTCTGAAGACAAGAGGTGGCGAGAGTTGTCTCGAATTAGATGGCTTGATTCATCGGTTCTT
CTCTGGATATAGCTGGCAAGAATATCCTGAAATTCTACACCAGTTTCTAACTGCATTGGACCTGCACCTTAAGACGGTAAACATTGATTGA
AA sequence
>Lus10004373 pacid=23151916 polypeptide=Lus10004373 locus=Lus10004373.g ID=Lus10004373.BGIv1.0 annot-version=v1.0
MRGSGFEPDEATMVVMLSLSAEVGNLGLGKWIHSQVIKGRMELNYQLGTAIVDMYAKSGSVGYAMSVFGSMEKRNVWTWSAMILGLAQHGFAKEGLELFH
QMIETSSIRPNYVTFLGVLCACSHAGLVEDGFRYFHEMEHQHGIKPTVVHYGAMVDILARAGHLREAYNFVTHMPHQNDPTVLRTLLGACSNRSENDDDG
VADKVRKRLLEVEPRRSGNLVMVAKMYADAGMWQRVADVRKVMRDGGLKTRGGESCLELDGLIHRFFSGYSWQEYPEILHQFLTALDLHLKTVNID
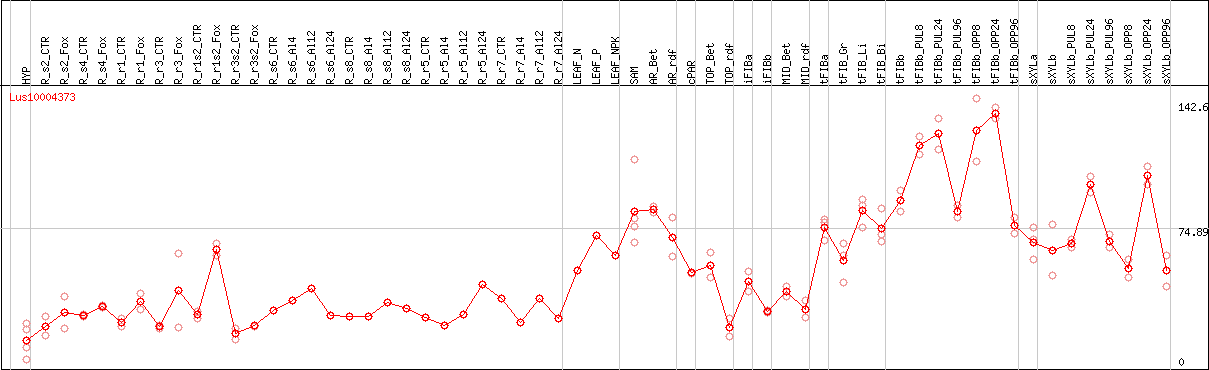
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10004373 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.