External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G19040 322 / 8e-110
ATIPT5
Arabidopsis thaliana ISOPENTENYLTRANSFERASE 5, isopentenyltransferase 5 (.1)
AT3G23630 293 / 1e-98
ATIPT7
ARABIDOPSIS THALIANA ISOPENTENYLTRANSFERASE 7, isopentenyltransferase 7 (.1)
AT3G63110 260 / 2e-85
ATIPT3
isopentenyltransferase 3 (.1)
AT1G68460 205 / 7e-64
ATIPT1
Arabidopsis thaliana isopentenyltransferase 1, isopentenyltransferase 1 (.1)
AT3G19160 184 / 4e-56
PGA22, ATIPT8
isopentenyltransferase 8, ATP/ADP isopentenyltransferases (.1)
AT1G25410 178 / 2e-53
ATIPT6
ARABIDOPSIS THALIANA ISOPENTENYLTRANSFERASE 6, isopentenyltransferase 6 (.1)
AT4G24650 158 / 7e-46
ATIPT4
ARABIDOPSIS THALIANA ISOPENTENYLTRANSFERASE 4, isopentenyltransferase 4 (.1)
AT2G27760 97 / 3e-22
IPPT, ATIPT2
tRNAisopentenyltransferase 2 (.1)
AT5G20040 73 / 2e-14
ATIPT9
ARABIDOPSIS THALIANA ISOPENTENYLTRANSFERASE 9, isopentenyltransferase 9 (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10034025
490 / 2e-176
AT5G19040 378 / 7e-132
Arabidopsis thaliana ISOPENTENYLTRANSFERASE 5, isopentenyltransferase 5 (.1)
Lus10028015
258 / 1e-84
AT3G63110 327 / 3e-111
isopentenyltransferase 3 (.1)
Lus10012372
257 / 5e-84
AT3G63110 332 / 5e-113
isopentenyltransferase 3 (.1)
Lus10034334
181 / 5e-55
AT1G68460 267 / 3e-88
Arabidopsis thaliana isopentenyltransferase 1, isopentenyltransferase 1 (.1)
Lus10034333
176 / 1e-52
AT1G68460 258 / 1e-83
Arabidopsis thaliana isopentenyltransferase 1, isopentenyltransferase 1 (.1)
Lus10034332
177 / 2e-52
AT1G68460 304 / 3e-101
Arabidopsis thaliana isopentenyltransferase 1, isopentenyltransferase 1 (.1)
Lus10006281
99 / 5e-23
AT2G27760 493 / 2e-172
tRNAisopentenyltransferase 2 (.1)
Lus10020566
98 / 1e-22
AT2G27760 491 / 1e-171
tRNAisopentenyltransferase 2 (.1)
Lus10012051
64 / 5e-11
AT5G20040 505 / 1e-177
ARABIDOPSIS THALIANA ISOPENTENYLTRANSFERASE 9, isopentenyltransferase 9 (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G202200
335 / 7e-115
AT5G19040 396 / 9e-139
Arabidopsis thaliana ISOPENTENYLTRANSFERASE 5, isopentenyltransferase 5 (.1)
Potri.010G030500
332 / 6e-114
AT5G19040 365 / 1e-126
Arabidopsis thaliana ISOPENTENYLTRANSFERASE 5, isopentenyltransferase 5 (.1)
Potri.014G139300
275 / 2e-91
AT3G63110 356 / 7e-123
isopentenyltransferase 3 (.1)
Potri.008G033300
234 / 3e-76
AT5G19040 235 / 3e-76
Arabidopsis thaliana ISOPENTENYLTRANSFERASE 5, isopentenyltransferase 5 (.1)
Potri.004G150900
219 / 2e-69
AT5G19040 257 / 6e-84
Arabidopsis thaliana ISOPENTENYLTRANSFERASE 5, isopentenyltransferase 5 (.1)
Potri.008G121500
191 / 2e-58
AT1G68460 348 / 6e-119
Arabidopsis thaliana isopentenyltransferase 1, isopentenyltransferase 1 (.1)
Potri.010G123801
172 / 6e-51
AT1G68460 338 / 4e-115
Arabidopsis thaliana isopentenyltransferase 1, isopentenyltransferase 1 (.1)
Potri.009G147600
100 / 1e-23
AT2G27760 544 / 0.0
tRNAisopentenyltransferase 2 (.1)
Potri.001G376600
73 / 3e-14
AT5G20040 568 / 0.0
ARABIDOPSIS THALIANA ISOPENTENYLTRANSFERASE 9, isopentenyltransferase 9 (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF01715
IPPT
IPP transferase
Representative CDS sequence
>Lus10004428 pacid=23154273 polypeptide=Lus10004428 locus=Lus10004428.g ID=Lus10004428.BGIv1.0 annot-version=v1.0
ATGATGAATTCCACTTTGTCGGGAGCTAATTACAGGCAAGTCCAATTCCCGACCATGAACAATTTCAACAACAAGTCCTTCACCATGGATCATCCTCCCT
TCTTCCGCCGTAAAGACAAGGTGGTCTTCGTGGTGGGACCCACTGGCACTGGCAAGTCCAGGCTGGCGATCGACCTGGCAACCCGCTTCCCTGCGGAGGT
GGTCAACTCTGACAAAATGCAAGTCTACAAGGGACTCGAGATCGCCACCAATAAGGTCACCGACGACGAGTGCCGCGGCGTCCCCCACCATTTGCTCGGT
GTCGTACTCGACCCCGCCGCTGATTATACCTCCGACGACTTCCGTCGCCACGCCTCCCATGTCGTCGACTCCGTCGTCTCCCGCGACCGCCTCCCCGTTG
TGGTGGGAGGGTCAAACTCCTTTGTCGACGCCCTGGTCAACGCTGACATTGACTTCCGTTCCCGCTACGACTTCTGCTTTCTCTGGGTCGACGTGGCGCT
CCCGGTGCTCTCCAAGTTCATTTCCGCTCGGGTTGACCGGATGGTGAAGGCGGGTTTGGTCGACGAGGTCGGTGCCGTTTTCGACCCGAATGACATGGAT
TACTCTCGGGGGATTCGCCGCGCGATCGGAGTTCCTGAATTGGACGCTTATTTTAGGTGCGGCGGCGGCGCCGCTGCTCTGGACGCGGCGATTGAGGAGA
TAAAGAAGAACACGTGCTTATTGGCTTGTCAGCAGCTGAAGAAGATCCAGCGGCTGCACAATCAGTGGAGCTGGAGTATGCATCGGGTCGACGCCACTGA
GGTTTTTCTCCGCCGAGGAATTGAAGCCGACGAAATCTGGGAGGAGCTTGTGGCGGGACCCAGCGTTTCGATTGTCGACCAATTCCTTCATCGGGAATTG
GATCTGGTGTCCGCGGCGACGGCCGTGGAAATGAAGGACTAA
AA sequence
>Lus10004428 pacid=23154273 polypeptide=Lus10004428 locus=Lus10004428.g ID=Lus10004428.BGIv1.0 annot-version=v1.0
MMNSTLSGANYRQVQFPTMNNFNNKSFTMDHPPFFRRKDKVVFVVGPTGTGKSRLAIDLATRFPAEVVNSDKMQVYKGLEIATNKVTDDECRGVPHHLLG
VVLDPAADYTSDDFRRHASHVVDSVVSRDRLPVVVGGSNSFVDALVNADIDFRSRYDFCFLWVDVALPVLSKFISARVDRMVKAGLVDEVGAVFDPNDMD
YSRGIRRAIGVPELDAYFRCGGGAAALDAAIEEIKKNTCLLACQQLKKIQRLHNQWSWSMHRVDATEVFLRRGIEADEIWEELVAGPSVSIVDQFLHREL
DLVSAATAVEMKD
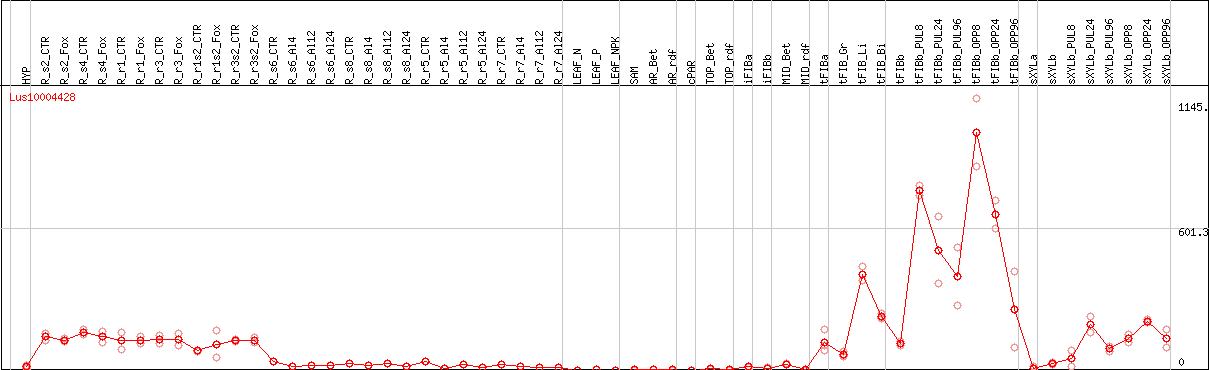
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10004428 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.