External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G27670 152 / 8e-48
HTA7
histone H2A 7 (.1)
AT5G59870 148 / 3e-46
HTA6
histone H2A 6 (.1)
AT5G02560 145 / 5e-45
HTA12
histone H2A 12 (.1.2)
AT1G08880 133 / 1e-40
HTA5 ,G-H2AX ,GAMMA-H2AX ,H2AXA
histone H2A 5, gamma histone variant H2AX, GAMMA H2AX, Histone superfamily protein (.1)
AT1G54690 133 / 1e-40
HTA3 ,G-H2AX ,GAMMA-H2AX ,H2AXB
histone H2A 3, GAMMA H2AX, gamma histone variant H2AX (.1)
AT4G27230 132 / 3e-40
HTA2
histone H2A 2 (.1.2)
AT1G51060 132 / 3e-40
HTA10
histone H2A 10 (.1)
AT3G20670 132 / 4e-40
HTA13
histone H2A 13 (.1)
AT5G54640 131 / 4e-40
ATHTA1, HTA1, RAT5
RESISTANT TO AGROBACTERIUM TRANSFORMATION 5, histone H2A 1, Histone superfamily protein (.1)
AT1G52740 85 / 1e-21
HTA9
histone H2A protein 9 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G018200
158 / 3e-50
AT5G27670 160 / 4e-51
histone H2A 7 (.1)
Potri.005G026500
156 / 2e-49
AT5G27670 145 / 2e-45
histone H2A 7 (.1)
Potri.006G082300
153 / 3e-48
AT5G02560 143 / 2e-44
histone H2A 12 (.1.2)
Potri.005G040700
136 / 7e-42
AT1G08880 183 / 2e-60
histone H2A 5, gamma histone variant H2AX, GAMMA H2AX, Histone superfamily protein (.1)
Potri.013G028900
136 / 1e-41
AT1G54690 220 / 4e-75
histone H2A 3, GAMMA H2AX, gamma histone variant H2AX (.1)
Potri.005G040800
135 / 1e-41
AT1G54690 221 / 2e-75
histone H2A 3, GAMMA H2AX, gamma histone variant H2AX (.1)
Potri.013G028800
135 / 2e-41
AT1G08880 190 / 2e-63
histone H2A 5, gamma histone variant H2AX, GAMMA H2AX, Histone superfamily protein (.1)
Potri.011G131400
132 / 3e-40
AT1G51060 200 / 1e-67
histone H2A 10 (.1)
Potri.001G415700
132 / 1e-39
AT1G51060 174 / 1e-56
histone H2A 10 (.1)
Potri.004G031300
130 / 2e-39
AT1G08880 150 / 6e-48
histone H2A 5, gamma histone variant H2AX, GAMMA H2AX, Histone superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0012
Histone
PF00125
Histone
Core histone H2A/H2B/H3/H4
Representative CDS sequence
>Lus10004502 pacid=23169706 polypeptide=Lus10004502 locus=Lus10004502.g ID=Lus10004502.BGIv1.0 annot-version=v1.0
ATGGATTCCGCGACAGAGACGACAACAGCTGCTGTAACAGCGACGCCGAAGGGTGCCGGCGGGAGAGGCCGAGGCGACCGTAAAAAGGCGGTTTCCAAAT
CGACGAAAGCAGGGCTCCAGTTCCCCGTCGGCCGCATTTTCCGCTTCCTCAAGAGAGGCCGCTACTCCCAGCGATACGGCGCCGGCGCCCCGATCTACCT
TGCCGCCGTCCTCGAGTATCTCGCCGCCGAGGTGCTGGAACTTGCCGGTAATGCAGCGAGGGATAACAAGAAGAACAGGATAAACCCAAGGCACGTTCTG
TTAGCAGTGAGGAACGACGAGGAACTTGGGAAGCTTCTTGCAGGAGTCACGATTGCAAGCGGAGGTGTTCTGCCGAACATCAATCCGGTGCTTTTGCCGA
AGAAGAAGATCACACCGGAGGGTAGTTCTTCTCCTGCTGCTGAACCCAAATCCCCAAAGAAGAAGAAAACTACCTAA
AA sequence
>Lus10004502 pacid=23169706 polypeptide=Lus10004502 locus=Lus10004502.g ID=Lus10004502.BGIv1.0 annot-version=v1.0
MDSATETTTAAVTATPKGAGGRGRGDRKKAVSKSTKAGLQFPVGRIFRFLKRGRYSQRYGAGAPIYLAAVLEYLAAEVLELAGNAARDNKKNRINPRHVL
LAVRNDEELGKLLAGVTIASGGVLPNINPVLLPKKKITPEGSSSPAAEPKSPKKKKTT
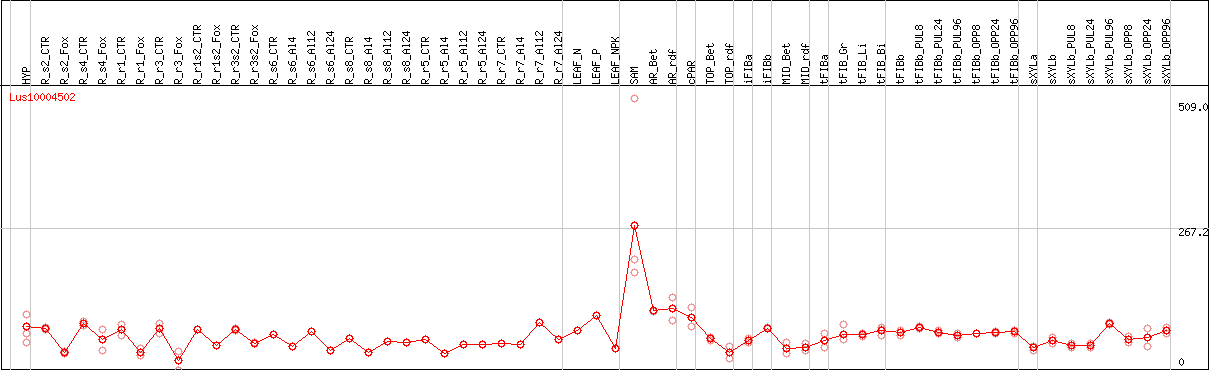
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10004502 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.