External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G17020 232 / 1e-74
ATSRG1, SRG1
senescence-related gene 1 (.1)
AT4G25300 220 / 5e-70
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
AT1G78550 217 / 5e-69
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G25310 215 / 4e-68
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G17010 213 / 2e-67
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT3G21420 140 / 3e-39
LBO1
LATERAL BRANCHING OXIDOREDUCTASE 1, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT2G38240 139 / 9e-39
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT5G05600 125 / 2e-33
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT3G55970 121 / 5e-32
ATJRG21
jasmonate-regulated gene 21 (.1)
AT3G11180 121 / 9e-32
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10011979
422 / 2e-149
AT1G17020 374 / 4e-129
senescence-related gene 1 (.1)
Lus10011980
389 / 3e-136
AT1G17020 382 / 1e-132
senescence-related gene 1 (.1)
Lus10011981
342 / 7e-118
AT1G17020 375 / 2e-129
senescence-related gene 1 (.1)
Lus10015252
276 / 5e-92
AT1G17020 375 / 1e-129
senescence-related gene 1 (.1)
Lus10026173
234 / 5e-75
AT1G17020 443 / 5e-156
senescence-related gene 1 (.1)
Lus10022292
211 / 2e-66
AT1G17020 449 / 5e-159
senescence-related gene 1 (.1)
Lus10004581
201 / 3e-64
AT4G25300 159 / 4e-49
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Lus10011985
194 / 7e-60
AT1G17020 367 / 4e-126
senescence-related gene 1 (.1)
Lus10032574
191 / 1e-58
AT4G25300 419 / 2e-146
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G355100
258 / 1e-84
AT1G17020 439 / 1e-154
senescence-related gene 1 (.1)
Potri.001G382400
248 / 6e-81
AT1G17020 446 / 1e-157
senescence-related gene 1 (.1)
Potri.001G381700
234 / 1e-75
AT1G17020 436 / 2e-153
senescence-related gene 1 (.1)
Potri.009G025900
206 / 2e-64
AT4G25300 410 / 3e-143
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Potri.009G022800
188 / 9e-58
AT1G17020 302 / 6e-101
senescence-related gene 1 (.1)
Potri.001G355200
171 / 5e-51
AT1G17020 329 / 3e-111
senescence-related gene 1 (.1)
Potri.010G201000
159 / 2e-46
AT3G21420 290 / 5e-96
LATERAL BRANCHING OXIDOREDUCTASE 1, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.010G200900
153 / 5e-44
AT3G21420 288 / 4e-95
LATERAL BRANCHING OXIDOREDUCTASE 1, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.010G023600
149 / 9e-43
AT3G21420 511 / 0.0
LATERAL BRANCHING OXIDOREDUCTASE 1, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.016G117100
134 / 7e-37
AT5G05600 491 / 4e-175
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
PFAM info
Representative CDS sequence
>Lus10004582 pacid=23149755 polypeptide=Lus10004582 locus=Lus10004582.g ID=Lus10004582.BGIv1.0 annot-version=v1.0
ATGAGAGAGATCATGGAACAAGAGCTAACAAAGCTTGGAAGCTCCCTGCCAGTGCCATGCGTTCAGGAACTAGCCAAATATCCCCTCCTCACCTCCGTCC
CTCCACCACGTTTCATCCGCCATGATCCACACCGTCCACACAACGATACCACTATCGAATCTACGGCTCAAGTTCCCTTAATCGGTTTGGAGAAACTACT
ATCCTCGGATCAAGAGCTGGATAGGCTCCACGACGCTTGCAAACATTGGGGTTTCTTCCAGGTGATTAATCATGGGGTGAGCAATTCATTAGTGGAGAGA
GTGAAGAAGGAGATTCAAGAATGGTTCAACATTCCAATGGAAGAGAAGAAGAAGTTCTGGCAGAGATGTGGAGATTTGGAAGGTTTTGGGCAAGCCTTTG
TGGTTTCTGAAGAGCAAAAGCTTGATTGGGGAGATATGTTCTACCTTACTAGCCTCCCTATACATGAGAGGAGACCTTATCTCTTCCCTTTGTTGCCTCT
CACACTCGGGTATTGTTTGAGTACTCAGCAACTGTTTGAATCGGGATACGCTGGAAGAATATTCAGTGGCACTGAAGAGTCTGGCCATGAAGATTCTGAA
CCCGATGGCCAAGCTCTAGGGATGGACCAAAATGACTTGAATGTGTTGTTTGATGAAGAAGGGTGGCAGTTATTCCGGATGAACTATTATCCACCGTGTC
CGCAGCCGGAGTTAGTGATGGGGCTCAACTCTCACTCCGACATCGTCGGACTCACCATCCTGCTGGAAGTTACCGCCAATACTCCAGGGTTACCGGTTAA
AAAAAGATGGATATTGGGTTCCGGTCACGCCTCTCCCTAA
AA sequence
>Lus10004582 pacid=23149755 polypeptide=Lus10004582 locus=Lus10004582.g ID=Lus10004582.BGIv1.0 annot-version=v1.0
MREIMEQELTKLGSSLPVPCVQELAKYPLLTSVPPPRFIRHDPHRPHNDTTIESTAQVPLIGLEKLLSSDQELDRLHDACKHWGFFQVINHGVSNSLVER
VKKEIQEWFNIPMEEKKKFWQRCGDLEGFGQAFVVSEEQKLDWGDMFYLTSLPIHERRPYLFPLLPLTLGYCLSTQQLFESGYAGRIFSGTEESGHEDSE
PDGQALGMDQNDLNVLFDEEGWQLFRMNYYPPCPQPELVMGLNSHSDIVGLTILLEVTANTPGLPVKKRWILGSGHASP
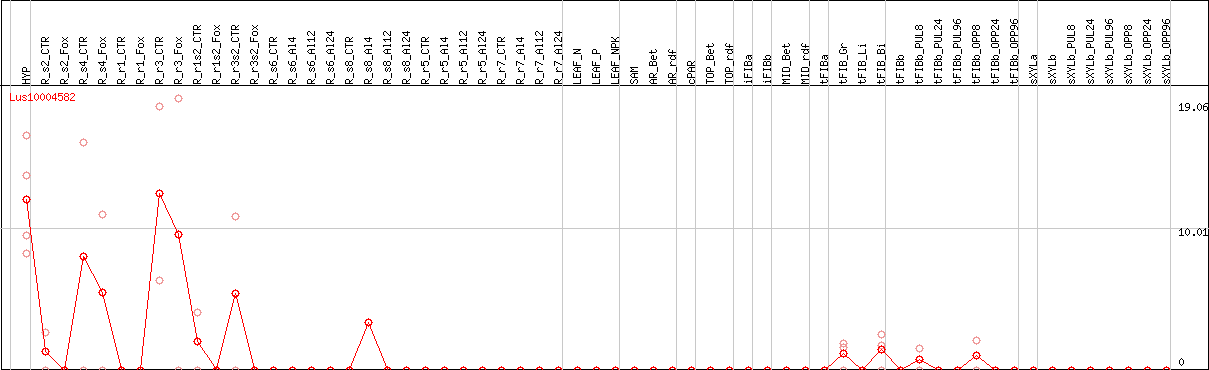
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10004582 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.