Lus10004730 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10004730 pacid=23162948 polypeptide=Lus10004730 locus=Lus10004730.g ID=Lus10004730.BGIv1.0 annot-version=v1.0
ATGGATTATGATGACAATGATTTTCAAAGCCAGAATCTTCATTTAGCCGGTGAAGGGAGCAACAAATTTCCTCCTGTCATAAACCCATATGCTCTTCCTA
AGTTTGAGTTTGACGACAATATTCATGTACCATTACGCTACGATAGCCTAGTTGAAACTGAGGTTTTTCTTGGCATTGAAAACAACGAAGATAGCCAGTG
GATTGAGGATTATTCTCGGGGTAGCAACACAATACAATTTGGTTCAAGTGCAGCAGAGTCTTGCTGCATTTTGAGACAGAACAATGTTTGGTCTGAGGCA
ACTTCCTCAGAATCTGTTGAGATGCTGTTAAGATCTGTCGGTAACGAAGAAAGCTTTCCGTTACAAGCTAGTGCTAAGAAGTCAGATGCATGTGATGAGC
TGGGCTGCATGATTAGCCAGATGGAACCTAGCTCAAAGCAGGATAGTAATTTACCTATTAGGCCAGTAGATTTTGCAGATGTGCAGCAGACTTTACAATC
TGCTGATATTGCTGAGAGATTCCCCACATTAAATGCCGAGGCTGGGGCCCAGCAGCCTCTGGATCAGGGTGTTTCAGAGGGTCATGATGGTAACAACTGT
TTTGAAAATGGTTTAGGTGATTTAACTGCTGTTGCGGCAGGCGCCGAGTTTTCTGTTGCAGTGGAGAACGAATTTATCGACGGTAAATGTGACAGTGTCG
GCCAAAGTGCTGGTAATACTGTTATTTATGAATCTTTAGCTATAAGGACAGTAGAAGCTGATGGTTCAGAGGTGCAAGTTGTTGACATAGTGGATACTGA
AAATATCAGCATTAGCAATATAAGGGCTGAGAATGAAGACGTCTTGAATCGGACCACGAATTCAGGTGATGGGATAAAACATACAGTAGAAATCGACACT
GATGACTGTAAAGAAGAAAGGGATGTATTGGGCGAAGCTGAAATCGGTGTATCCCAAGAAATTTCAGATGCAACCTTGGTTGAAACTGGTAGTAAAGAGT
TGCAGAGTCCTTTCTGTTCTGCCCCTGTGGAAAATGCAAGTGAAGCAAATATAAGCGGCAGTAACTTGAGCAATATGGAAGGACCTTATACCTTCAGGAC
AGAGACAGAGACAGACTTGTCTGAGAACCTAGAGGATGCTGATGTATCTAGAGGAAGTGAAAGTATACTATTAGTACATGTCGATAATAGGGTGCAGGAT
GGGAGTAGCAGCGTTTCCCCGTTGCTGACAGATAGCTTTGTCACAAGCTCTGAGGTACTTCCGACATCTGAGGAGAAAGAACCTTTTGCTGATGGGAATG
CAAGCATTGAGGTCTCTCATGCTGATGATAGTTCTGATTATAATATTCTTGCTACAAAAGCTTGCGAGGGTACTCCACCATCCTTTCCGGCTGAATCTCC
GCAAGGGCATGGGAGAGAAATAGTCCCTCAAGAGGCTGAATCTCCCCAAGGGCATTTGAGAGAAATAGTCCCTGAAGAGACTGAATCTCCGCAAGGGCAT
GAGAGAGAAATAGACCCTCAAGAGGATGAAGATCGTGGATTAGCACTAGATGGTCTTGTTCAGCAAAATGAAAGGTCAGAGTCACCTTCTAATGGCGCTA
CTGTCAACAATGAGGTTCAGAACTCTCCTATAACAGATAGGAACATTGGGACCTTGCCTTCAGCGGGTGTAGATGACAAGGTAATGGGTACAGAATACCC
TGTTGATCTGGCTGCTGGCGATGCTACAGCGAGGACTGTTGATCCTAGCATTTCCGAAGTCTCCTCCGTATCAGCTTTGGGCTCTGCTCAAATATCAGAA
GAAGATGGCAGAGATATTCCAATCTCAGGGTTATCTGAAGGAACAGCAGCTATTGTAGGCCCTGTCTTAGAATTGGAAAAGGGGGTTTCTGGAGCATCTG
GACAAGGTTTAAATTCAAATGTGGATGAGTCTTTATCAATGGTTGATGAGTGTAGTCAAAAAGACCATATTGATTCTGAAGCAGCCTCTATGAATGAAGA
TGCTCCAGAATCTTCAAAGGAAACCGAAGAATGCTCGATTCGTCCGACTTCTTCGCTGAAGGAAAACAATGCTATGATTGTTGAAGAAACCGAGGAGAAG
GCTTCATTGAAAGATCAAGATAATGTTTCAAAGCAGAATGCTAGTCAGAAGGGGCATGTGGAAGATGAACCTGTCCTAACCATTAATACTAATGACAGAC
AGGATTCTATGACAGACAGAAGTGATAATGTTTCAAAGCAGGATGCTAGTCAGAAGGGGCATGGGGAAGATGGAGCTGTCCTAGCCATTGATACTAATGA
CAAACAGGATTCTATGACGGACAGAAGTGAATGTGAGACAACTGCTGCTGATCTGGAGAAGCAATCTGTTGCTTCGTCTGTAGTCGCACCAACTACTGAA
CTCTCTGAAGCTCTTCCAACGGAGGAAATCAATAGATATCCACATTTTATCCCTTCAACTTCAGTTGACGCCAACAAGTCTTTCTCTCATGATTCCAAGA
AGGGCACACCCTCAAAAGATGATGGTAGCTTCACTTTTCAGGTCACCCCTTTGACAGAGTCGCCACAAGTGGAAAATACGATGAAATTGCAGCACCTGCA
GACTGTGGAGCCCAGTAAAGTATCTAAGCCTATGGGTGCTTCTCCATCGATGCCTGTTTTAGGCCAATTAGATGGCAAAATTGTCCAAGATGTCTCACAT
CAGAGTCCTAAGGTTTCTGATGTAACCATTCGTCAAGTTGGTGCAAAAGGAAATTCTGAGCGTAAGCCCAGACGAAGGGCAACTCCAAAAGAGTCTGCTA
AGAGGGTAAAGACTCCAAAGGAGATAACACCTGCAAAGATGGAAAGGGGTAACAAAGCAATTATTACACCGAGTTCATCAGTAGTTGCTGAACGTGCACA
GTCAAATGAGATTCAGCGATATGGGCCTCTTGTTGGGACATCGATGGCTGGTTTGCCAGATTTGAATTCTTCAGCCTCTTCCATGGTGGTGTTTCAACAA
CCGTTCACGGATATGCAACAAGTTCAATTACGTGCTCAGATCTTTGCATATGGAGCTTTGATTCAAGGAACAGCCCCAGAGGAGGCATATATGATATCTG
CATTTGGAGGCCCTGATGGTGGTAAGAGCGCTTGGGAGAAACCATGGCGTTTGTTTGTAGAAAGACTGAATAGTCAAATAACCCATCTCAACACCCCGGG
GACTCCAGTGCAGTCTTTCTCTGGTGCTAGAGTTCCAGTTCAAGATCAAAGTAGTAAGCAAAGCACACGTAAAGTTAAGGCCGGGTCCTCTACTGGTGCT
AGATCCAGCAGCAAGGCAATAACGCCAATTACAAGTCCCACACTACCTCTTTCGACACCGCTTTGGAATGTCTCAACTCCTTCGGATCGCATGCAATCAA
GTTCCGTAGCAAGGAGTCCTGTTCTGGACTTCCAGAGGACACTTTCTCCATTGCCGTCTCATCAGCCCCCAGCCATGAGGAATTTGGTTGGACATAATCC
TTGGATCTCTCCGGGTCATTTTGGTCCCTGGATGACTTCGCCACAAACCTCTGCAGTCGATACTGGCAGTCGTTTTACCGTACAAATGCCTCATACAGAG
CCCGTTCACCTGACTCCCGTAAAAGACTCATCTTTGCCTCTCCCCTCTGCTATGAAGCATGTTTCTACTCCACATGTGACTTATACAGATCCATCTGGCA
ACTTGTTAGGCAATACTTCCATGCCTGAGGTTAGCAAGATGGTAATAGCATCTGGTCAACAGTCCACCATTCCGAAGCCTAGAAAAAGGAAGAAGTCGTC
TGTGCCAGAGAATACCGTTAACATGAGTTTACTCCACCAGCCTTCACAAGAATTCGTTCCTTCTGTTGCCATCACAACCCCCGTCAGCTTTGTTCCTAAA
CCGCCTGCGGAAATTCCTCTTCCTTCTACTTCTCCTGTGACTACCAATTTCACGAAAGCGGACCTGTTTGCAGACCAGAGGGTTAACTCCTCTGAAGAAT
CACTCAGCCTAGTGAAAGATGCTAGACATCATGCAGAGGATGCTGCTACTCTTGCTGCTACTGCTATTAGTCAAAGCCAAGAACTATGGAGTAAATTGGA
TAAGCAGAAAAATGCCGGCTTGTCACCCGACGTGGAAGCAAAACTATCTTCTGCAGCCGTGGCAATAGCTGCTGCTTCTGCCGTCGCAAAAGCCGCAGCC
GCAGCTGCTAAAGTTGCGGCGGATGCCGCATTGCAAGCTCGGCTAATGGCAGACGAGGCGGTTCAGACTTCTGTAATCTCTTTCTCTGATGAGATGAAGA
ATTTGGGAAATGCAACACCAGCTACGATATTGAGACGTGACGACGGAACCAGCAGTTCCAGTTCGATCATTCTCGCTGCGAGAGAAACAGCAAGAAAAAG
GGTCGAAGCAGCCTCTGCTGCATCAAAACGAGCTGAAAATATTGATGCCATTATTAGAGCTGCGGAGCTTGCTGCAGAAGCTGTTTCTCAAGCTGGAAAG
ATTGTAGCTATGGGCGATCCTTTGCCCTTGAGTGTGCTAGCCTTAACCGGTCCAGAGGGCTACTGGAAAGATCCCCAAGTGTCTCCCCAGCGCGTCTGTA
AAGCCAATGAGCTCATCACTGTCAATTTGGATGTTCCTAGTAGAGGAAGAGATGTTTCTGGCGATTCTGCTAGAAGTAGGACCAAGGAAGGCTTGTCTAG
CAGCCAAACTGGGATGCCAGGTGGCAAGGAAGCCAAAGAACTCAAAGGTCCTAAATATTCAGAGACAACCTTGGAAGTGGTTCTTCTGGAAGATGGAAAT
GGATTGAAATCTCCGACCACTTCAGAGACTGAAAATATAGAAGCTGTTGGGAGTTCAAAAGAAAGCGTCATCAGAGAAGGGAGCAGTGTAGAGATCTTGA
GGGATGGAAATGGATTCAAAGCGGCCTGGTTTGCAGCAAAAGTATTGAATCTGCGCGATGACTTAGCTTATGTAAGCTATGATGAACTTACATCAGGGGA
AGGCTCAGAGAAGTTAACTGAGTGGGTAGCCCTTCGAGGCGAAGGAGCAGAGGCTCCAAAAATACGCATTTCTCGCCCTGTTATTGCCCTGCCATATGAA
GGAACTAGGAAGAGGCGGAGATCTGCCGTAAACGACTATAACTGGTCGGTTGGCGACCGAGTGGATGCATGGTTAAGGGAAAGTTGGTGGGAGGGAGTTG
TCACAGAAAAGAACAAGAAAGATGAGACCCACTTGACGGTTTATTTTCCAGTACAAGGAGAAACATCGGTAGTAAGAGCCTGGCATCTTCGTCCTTCTCT
TCTATGGAAAGATGGGGAATGGATGGAGTGGTCGAATTTACAGCAGAAGAATGTCTCGTTCAATACGGGAGAAACGCCGAAACAGAAACGGTCCAGAATG
CAGAGCCCTGTAGTAGAATCCAAAGACAAGGACAAGGTGTCTGAAAATCCGATTGTCAAAGAGTCCGGTAAACATGACGAGTCAACGTCGCTGAATTTGG
CAGCTGGTGAGGCGGTATTTAACGTTGGTAAGGCTACTGGAGATGGAAGCAGAACGGCTCGAACTGGTTTGCAGAAGGAAGGATCGAAGGTCGTCTTTGG
CATACCGAAGCCGGGAAAGAAACGGAAGTTCATGGAAGTGAGCAAACACTACGTTGCTGGCCGAACCGACAAGACGAATGAAGCAAACGATCCGGCGAAA
TCTGCGAAATATTCGACGCCTCAAGTATCAGGTCCTCGGGGATGGAGAAGTACACTGAAAACAGACACCGCTGCCGAGAAACGAACAACAACAGCAGCGT
TTGCTGCACCCAAGTCCAAGCTTCCCAAGCCCGGTAATCGACCTAATCCTTCCAGTAGGACAATTCCGCAGAAGGAGAACTCGTCGAACAGCAGCAGTGG
TGCTGCGTTTACAGACAATAGTGTGAAGGCTAAAACCGGTGGAGAAGCTTCTTCAAACGCACTTGAGAAAGAGAGTTCGATAGACCGTTCCTTCTCCAGC
TCCGGGGAAGCAACAGACCCCCCGATTGTATTTTCTGCAGTTGCTCGACGATCAGATGCCATCCCCTCTAAGAAACTGTCGACGACAAACGCAAGAAGCG
TGCGGGGTAACAAGGGAAAATTCGCTCCTGCTGCAGGCAGATTGGGCAAAGTTGAAGAGGGCAAAGTCTTCAACAGCGCTGCTTCGTCCACTTCAGGACA
GTCTTCCTCAGCCAACGAGCCTCGTAGGTCCGTCCGTCGAATCCAGCCAACTTCAAGACTACTGGAAGGGCTGCAGAGCTCGTTGATGGTCTCGAAAATA
CCGTCCTCAGTCTCTCACGATAGGAGCCACAAGAGTAGGCCGGCCTTTAGAGGCAATAACAACGGTTGA
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10004730 pacid=23162948 polypeptide=Lus10004730 locus=Lus10004730.g ID=Lus10004730.BGIv1.0 annot-version=v1.0
MDYDDNDFQSQNLHLAGEGSNKFPPVINPYALPKFEFDDNIHVPLRYDSLVETEVFLGIENNEDSQWIEDYSRGSNTIQFGSSAAESCCILRQNNVWSEA
TSSESVEMLLRSVGNEESFPLQASAKKSDACDELGCMISQMEPSSKQDSNLPIRPVDFADVQQTLQSADIAERFPTLNAEAGAQQPLDQGVSEGHDGNNC
FENGLGDLTAVAAGAEFSVAVENEFIDGKCDSVGQSAGNTVIYESLAIRTVEADGSEVQVVDIVDTENISISNIRAENEDVLNRTTNSGDGIKHTVEIDT
DDCKEERDVLGEAEIGVSQEISDATLVETGSKELQSPFCSAPVENASEANISGSNLSNMEGPYTFRTETETDLSENLEDADVSRGSESILLVHVDNRVQD
GSSSVSPLLTDSFVTSSEVLPTSEEKEPFADGNASIEVSHADDSSDYNILATKACEGTPPSFPAESPQGHGREIVPQEAESPQGHLREIVPEETESPQGH
EREIDPQEDEDRGLALDGLVQQNERSESPSNGATVNNEVQNSPITDRNIGTLPSAGVDDKVMGTEYPVDLAAGDATARTVDPSISEVSSVSALGSAQISE
EDGRDIPISGLSEGTAAIVGPVLELEKGVSGASGQGLNSNVDESLSMVDECSQKDHIDSEAASMNEDAPESSKETEECSIRPTSSLKENNAMIVEETEEK
ASLKDQDNVSKQNASQKGHVEDEPVLTINTNDRQDSMTDRSDNVSKQDASQKGHGEDGAVLAIDTNDKQDSMTDRSECETTAADLEKQSVASSVVAPTTE
LSEALPTEEINRYPHFIPSTSVDANKSFSHDSKKGTPSKDDGSFTFQVTPLTESPQVENTMKLQHLQTVEPSKVSKPMGASPSMPVLGQLDGKIVQDVSH
QSPKVSDVTIRQVGAKGNSERKPRRRATPKESAKRVKTPKEITPAKMERGNKAIITPSSSVVAERAQSNEIQRYGPLVGTSMAGLPDLNSSASSMVVFQQ
PFTDMQQVQLRAQIFAYGALIQGTAPEEAYMISAFGGPDGGKSAWEKPWRLFVERLNSQITHLNTPGTPVQSFSGARVPVQDQSSKQSTRKVKAGSSTGA
RSSSKAITPITSPTLPLSTPLWNVSTPSDRMQSSSVARSPVLDFQRTLSPLPSHQPPAMRNLVGHNPWISPGHFGPWMTSPQTSAVDTGSRFTVQMPHTE
PVHLTPVKDSSLPLPSAMKHVSTPHVTYTDPSGNLLGNTSMPEVSKMVIASGQQSTIPKPRKRKKSSVPENTVNMSLLHQPSQEFVPSVAITTPVSFVPK
PPAEIPLPSTSPVTTNFTKADLFADQRVNSSEESLSLVKDARHHAEDAATLAATAISQSQELWSKLDKQKNAGLSPDVEAKLSSAAVAIAAASAVAKAAA
AAAKVAADAALQARLMADEAVQTSVISFSDEMKNLGNATPATILRRDDGTSSSSSIILAARETARKRVEAASAASKRAENIDAIIRAAELAAEAVSQAGK
IVAMGDPLPLSVLALTGPEGYWKDPQVSPQRVCKANELITVNLDVPSRGRDVSGDSARSRTKEGLSSSQTGMPGGKEAKELKGPKYSETTLEVVLLEDGN
GLKSPTTSETENIEAVGSSKESVIREGSSVEILRDGNGFKAAWFAAKVLNLRDDLAYVSYDELTSGEGSEKLTEWVALRGEGAEAPKIRISRPVIALPYE
GTRKRRRSAVNDYNWSVGDRVDAWLRESWWEGVVTEKNKKDETHLTVYFPVQGETSVVRAWHLRPSLLWKDGEWMEWSNLQQKNVSFNTGETPKQKRSRM
QSPVVESKDKDKVSENPIVKESGKHDESTSLNLAAGEAVFNVGKATGDGSRTARTGLQKEGSKVVFGIPKPGKKRKFMEVSKHYVAGRTDKTNEANDPAK
SAKYSTPQVSGPRGWRSTLKTDTAAEKRTTTAAFAAPKSKLPKPGNRPNPSSRTIPQKENSSNSSSGAAFTDNSVKAKTGGEASSNALEKESSIDRSFSS
SGEATDPPIVFSAVARRSDAIPSKKLSTTNARSVRGNKGKFAPAAGRLGKVEEGKVFNSAASSTSGQSSSANEPRRSVRRIQPTSRLLEGLQSSLMVSKI
PSSVSHDRSHKSRPAFRGNNNG
|
DESeq2's median of ratios [FLAX]
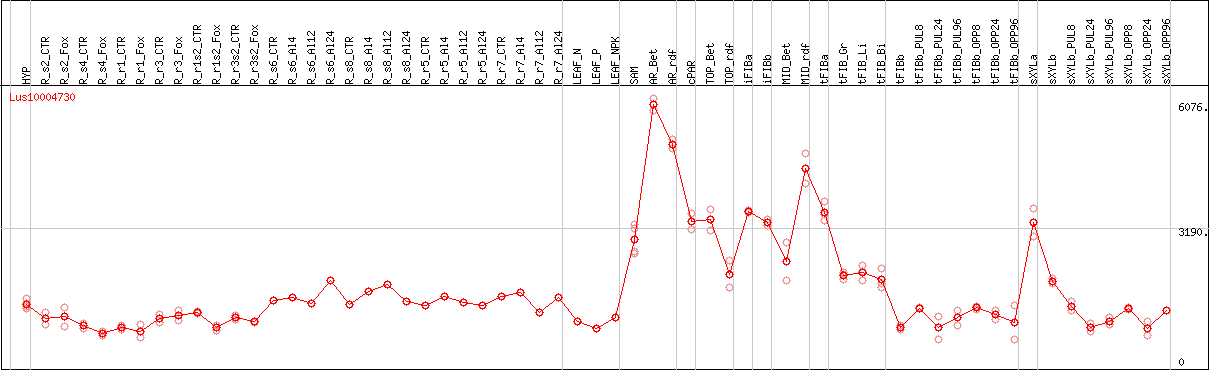
Coexpressed genes
Lus10004730 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
