External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G55280 187 / 7e-62
RPL23A2, RPL23AB
RIBOSOMAL PROTEIN L23A2, ribosomal protein L23AB (.1.2.3)
AT2G39460 181 / 2e-59
RPL23A1, RPL23AA, ATRPL23A
RIBOSOMAL PROTEIN L23A1, ARABIDOPSIS THALIANA RIBOSOMAL PROTEIN L23A, ribosomal protein L23AA (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005441
216 / 6e-73
AT3G55280 208 / 8e-70
RIBOSOMAL PROTEIN L23A2, ribosomal protein L23AB (.1.2.3)
Lus10023438
210 / 6e-71
AT3G55280 206 / 4e-69
RIBOSOMAL PROTEIN L23A2, ribosomal protein L23AB (.1.2.3)
Lus10030325
210 / 7e-71
AT3G55280 204 / 1e-68
RIBOSOMAL PROTEIN L23A2, ribosomal protein L23AB (.1.2.3)
Lus10040316
211 / 1e-70
AT3G55280 202 / 2e-67
RIBOSOMAL PROTEIN L23A2, ribosomal protein L23AB (.1.2.3)
Lus10005883
208 / 3e-70
AT3G55280 205 / 7e-69
RIBOSOMAL PROTEIN L23A2, ribosomal protein L23AB (.1.2.3)
Lus10003290
207 / 1e-69
AT3G55280 207 / 8e-70
RIBOSOMAL PROTEIN L23A2, ribosomal protein L23AB (.1.2.3)
Lus10040865
212 / 1e-65
AT5G59750 714 / 0.0
DHBP synthase RibB-like alpha/beta domain;GTP cyclohydrolase II (.1.2)
Lus10024430
89 / 3e-23
AT3G55280 98 / 8e-27
RIBOSOMAL PROTEIN L23A2, ribosomal protein L23AB (.1.2.3)
Lus10025302
89 / 3e-23
AT3G55280 98 / 8e-27
RIBOSOMAL PROTEIN L23A2, ribosomal protein L23AB (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.009G027000
204 / 2e-68
AT3G55280 199 / 2e-66
RIBOSOMAL PROTEIN L23A2, ribosomal protein L23AB (.1.2.3)
Potri.008G050200
202 / 1e-67
AT3G55280 187 / 8e-62
RIBOSOMAL PROTEIN L23A2, ribosomal protein L23AB (.1.2.3)
Potri.010G210300
202 / 2e-67
AT3G55280 195 / 7e-65
RIBOSOMAL PROTEIN L23A2, ribosomal protein L23AB (.1.2.3)
Potri.016G079800
201 / 2e-67
AT3G55280 187 / 5e-62
RIBOSOMAL PROTEIN L23A2, ribosomal protein L23AB (.1.2.3)
Potri.006G213300
199 / 1e-66
AT3G55280 204 / 1e-68
RIBOSOMAL PROTEIN L23A2, ribosomal protein L23AB (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00276
Ribosomal_L23
Ribosomal protein L23
PF03939
Ribosomal_L23eN
Ribosomal protein L23, N-terminal domain
Representative CDS sequence
>Lus10004942 pacid=23174815 polypeptide=Lus10004942 locus=Lus10004942.g ID=Lus10004942.BGIv1.0 annot-version=v1.0
ATGGCACCTCCGAAAGATAGCAAGGCGAAGCCAGATGCCAAGGCACAGGCAGCCAAGGCCGCAAAGGCAGTGAAGTCAGGCCCTACTTTCAAGAAGTCCG
CCAAGAAGATCAGGACCAAGGTCACCTTCCACCGACCGAGGACACTGAAGAAGGAGCGAAATCCCAAGTACCCCAGAATCAGTGCACCTCCGAGGAACAA
GCTTGACCATTACCAGATTCTGAAGTTCCCTTTGACCACCGAGTCTGCCATGAAGAAGATCGAGGACAACAACACTCTGGTGTTCATCGTTGACATCCGT
GCCGACAAGAAGAAGATCAAGGATGCCGTCAAGAAGATGTATGACATCCAAGCAAAGAAAGTCAACACTCTCATCAGGCCGGATGGTACCAAGAAGGCGT
ACGTTAGGTTGACACCGGACTACGATGCATTGGATGTCGCCAACAAAATCGGCATCATCTAA
AA sequence
>Lus10004942 pacid=23174815 polypeptide=Lus10004942 locus=Lus10004942.g ID=Lus10004942.BGIv1.0 annot-version=v1.0
MAPPKDSKAKPDAKAQAAKAAKAVKSGPTFKKSAKKIRTKVTFHRPRTLKKERNPKYPRISAPPRNKLDHYQILKFPLTTESAMKKIEDNNTLVFIVDIR
ADKKKIKDAVKKMYDIQAKKVNTLIRPDGTKKAYVRLTPDYDALDVANKIGII
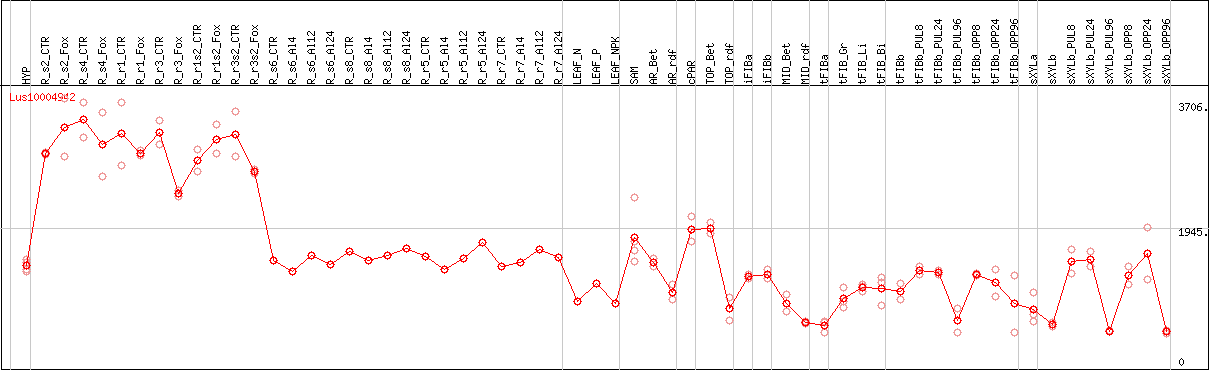
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10004942 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.