External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G07420 190 / 1e-61
SMO2-1, ATSMO1, ATSMO2-2, SMO2-2
Arabidopsis thaliana sterol 4-alpha-methyl oxidase 1, sterol 4-alpha-methyl-oxidase 2-1 (.1.2)
AT2G29390 187 / 4e-61
ATSMO2-2, SMO2-2, ATSMO2, SMO2-1
Arabidopsis thaliana sterol 4-alpha-methyl-oxidase 2-2, sterol 4-alpha-methyl-oxidase 2-2 (.1.2.3.4.5)
AT4G22756 110 / 9e-30
ATSMO1-2, ATSMO1, SMO1-2
sterol C4-methyl oxidase 1-2 (.1)
AT4G22753 108 / 6e-29
ATSMO1-3, ATSMO1, SMO1-3
sterol 4-alpha methyl oxidase 1-3 (.1.2)
AT4G12110 107 / 1e-28
ATSMO1-1, SMO1-1
sterol-4alpha-methyl oxidase 1-1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10001556
234 / 2e-78
AT1G07420 458 / 3e-165
Arabidopsis thaliana sterol 4-alpha-methyl oxidase 1, sterol 4-alpha-methyl-oxidase 2-1 (.1.2)
Lus10016492
198 / 5e-64
AT1G07420 463 / 3e-167
Arabidopsis thaliana sterol 4-alpha-methyl oxidase 1, sterol 4-alpha-methyl-oxidase 2-1 (.1.2)
Lus10040739
197 / 5e-64
AT2G29390 454 / 5e-164
Arabidopsis thaliana sterol 4-alpha-methyl-oxidase 2-2, sterol 4-alpha-methyl-oxidase 2-2 (.1.2.3.4.5)
Lus10004321
107 / 6e-29
AT4G12110 422 / 2e-150
sterol-4alpha-methyl oxidase 1-1 (.1)
Lus10032193
107 / 2e-28
AT4G12110 483 / 4e-174
sterol-4alpha-methyl oxidase 1-1 (.1)
Lus10028908
100 / 1e-25
AT4G22756 288 / 2e-97
sterol C4-methyl oxidase 1-2 (.1)
Lus10004322
91 / 4e-22
AT4G12110 357 / 2e-124
sterol-4alpha-methyl oxidase 1-1 (.1)
Lus10024555
89 / 2e-21
AT4G12110 442 / 1e-158
sterol-4alpha-methyl oxidase 1-1 (.1)
Lus10004324
88 / 2e-21
AT4G22756 345 / 3e-120
sterol C4-methyl oxidase 1-2 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G245300
192 / 7e-62
AT1G07420 476 / 2e-172
Arabidopsis thaliana sterol 4-alpha-methyl oxidase 1, sterol 4-alpha-methyl-oxidase 2-1 (.1.2)
Potri.009G037400
191 / 1e-61
AT1G07420 470 / 5e-170
Arabidopsis thaliana sterol 4-alpha-methyl oxidase 1, sterol 4-alpha-methyl-oxidase 2-1 (.1.2)
Potri.001G116500
107 / 1e-28
AT4G12110 439 / 1e-156
sterol-4alpha-methyl oxidase 1-1 (.1)
Potri.003G116000
103 / 3e-27
AT4G12110 448 / 3e-160
sterol-4alpha-methyl oxidase 1-1 (.1)
Potri.014G069500
100 / 7e-26
AT4G12110 403 / 2e-142
sterol-4alpha-methyl oxidase 1-1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04116
FA_hydroxylase
Fatty acid hydroxylase superfamily
Representative CDS sequence
>Lus10004988 pacid=23174850 polypeptide=Lus10004988 locus=Lus10004988.g ID=Lus10004988.BGIv1.0 annot-version=v1.0
ATGGATACCCTCTTCGAATCCGGATGGCAGGTAAACCAACCTCAACTCAACTCACTCTATTACTTTGGTGTTTGTTCTGTGCAAATTCCTGAGGGTTTGT
GTATTTTCAGGTATGCTACACCCTTTGGTCTGACATCTGAATATGCACACCCTGCTGAGATTCTGTTCCTCGGATTCGCCACGATCGCCGGCCCTGCAAT
CACCGGGCCGCATCTTTCGACGCTGTGGTCGTTGGTGGTGGTTCGGGTTCTCAAAACCGTTGACGTTCACTCAGGTTTCCATTTCCCCTGGAGCCTGTCG
AATTTCCTTCCTTTATATGGAGGTGCGGACTTCCACGACTATCACCATCGGCTTCTGTACACGAAATCCGGAAACTACACGTCCACCTTCACATACATGG
ACCGGATATTTGGCACTGATCAAGGTTACAGAAAGCTGAAGGCTTTGAAGAGTGGTGAAGACGTACACGTCAGCACCACCATAACCAGCAGCAGCAAGAT
TCTTTAG
AA sequence
>Lus10004988 pacid=23174850 polypeptide=Lus10004988 locus=Lus10004988.g ID=Lus10004988.BGIv1.0 annot-version=v1.0
MDTLFESGWQVNQPQLNSLYYFGVCSVQIPEGLCIFRYATPFGLTSEYAHPAEILFLGFATIAGPAITGPHLSTLWSLVVVRVLKTVDVHSGFHFPWSLS
NFLPLYGGADFHDYHHRLLYTKSGNYTSTFTYMDRIFGTDQGYRKLKALKSGEDVHVSTTITSSSKIL
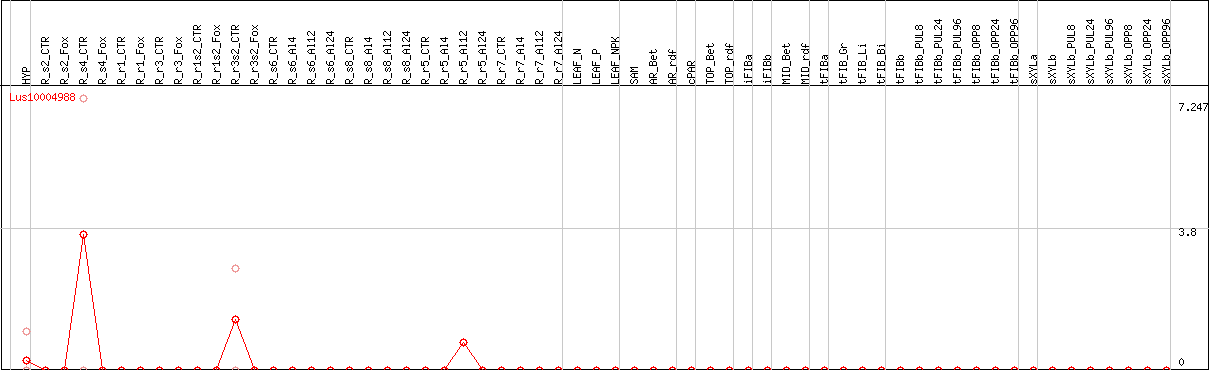
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10004988 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.