External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G02310 374 / 3e-132
MADS
AGL4, SEP2
SEPALLATA 2, AGAMOUS-like 4, K-box region and MADS-box transcription factor family protein (.1)
AT5G15800 360 / 9e-127
MADS
AGL2, SEP1
SEPALLATA1, AGAMOUS-like 2, K-box region and MADS-box transcription factor family protein (.1.2)
AT1G24260 278 / 1e-94
MADS
AGL9, SEP3
SEPALLATA3, AGAMOUS-like 9, K-box region and MADS-box transcription factor family protein (.1.2.3)
AT2G03710 260 / 2e-87
MADS
AGL3, SEP4
SEPALLATA 4, AGAMOUS-like 3, K-box region and MADS-box transcription factor family protein (.1.2.3)
AT2G45650 209 / 3e-67
MADS
AGL6
AGAMOUS-like 6 (.1)
AT1G69120 177 / 7e-55
MADS
AGL7, AP1
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
AT1G26310 176 / 3e-54
MADS
CAL1, AGL10, CAL
CAULIFLOWER, AGAMOUS-like 10, K-box region and MADS-box transcription factor family protein (.1)
AT5G60910 175 / 3e-54
MADS
FUL, AGL8
FRUITFULL, AGAMOUS-like 8 (.1.2)
AT3G61120 172 / 4e-53
MADS
AGL13
AGAMOUS-like 13 (.1)
AT4G09960 159 / 3e-48
MADS
AGL11, STK
SEEDSTICK, AGAMOUS-like 11, K-box region and MADS-box transcription factor family protein (.1.2.3.4)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10034371
495 / 4e-180
AT3G02310 364 / 3e-128
SEPALLATA 2, AGAMOUS-like 4, K-box region and MADS-box transcription factor family protein (.1)
Lus10003124
296 / 1e-101
AT1G24260 355 / 6e-125
SEPALLATA3, AGAMOUS-like 9, K-box region and MADS-box transcription factor family protein (.1.2.3)
Lus10015765
284 / 1e-96
AT1G24260 347 / 5e-122
SEPALLATA3, AGAMOUS-like 9, K-box region and MADS-box transcription factor family protein (.1.2.3)
Lus10037040
274 / 7e-93
AT1G24260 350 / 7e-123
SEPALLATA3, AGAMOUS-like 9, K-box region and MADS-box transcription factor family protein (.1.2.3)
Lus10034663
274 / 1e-92
AT5G15800 267 / 4e-90
SEPALLATA1, AGAMOUS-like 2, K-box region and MADS-box transcription factor family protein (.1.2)
Lus10003125
263 / 2e-89
AT1G24260 290 / 9e-101
SEPALLATA3, AGAMOUS-like 9, K-box region and MADS-box transcription factor family protein (.1.2.3)
Lus10004638
242 / 2e-80
AT3G02310 246 / 2e-82
SEPALLATA 2, AGAMOUS-like 4, K-box region and MADS-box transcription factor family protein (.1)
Lus10017870
223 / 4e-73
AT1G69120 212 / 3e-69
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Lus10026678
219 / 1e-71
AT5G15800 225 / 3e-74
SEPALLATA1, AGAMOUS-like 2, K-box region and MADS-box transcription factor family protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G115500
422 / 2e-151
AT3G02310 357 / 4e-126
SEPALLATA 2, AGAMOUS-like 4, K-box region and MADS-box transcription factor family protein (.1)
Potri.017G099700
416 / 5e-149
AT3G02310 372 / 8e-132
SEPALLATA 2, AGAMOUS-like 4, K-box region and MADS-box transcription factor family protein (.1)
Potri.003G169600
299 / 5e-103
AT1G24260 359 / 7e-127
SEPALLATA3, AGAMOUS-like 9, K-box region and MADS-box transcription factor family protein (.1.2.3)
Potri.001G058400
295 / 2e-101
AT1G24260 362 / 8e-128
SEPALLATA3, AGAMOUS-like 9, K-box region and MADS-box transcription factor family protein (.1.2.3)
Potri.008G098400
295 / 2e-101
AT5G15800 275 / 2e-93
SEPALLATA1, AGAMOUS-like 2, K-box region and MADS-box transcription factor family protein (.1.2)
Potri.014G074100
229 / 2e-75
AT2G45650 311 / 5e-108
AGAMOUS-like 6 (.1)
Potri.015G134800
206 / 5e-66
AT2G45650 209 / 2e-67
AGAMOUS-like 6 (.1)
Potri.012G132600
196 / 1e-62
AT2G45650 182 / 3e-57
AGAMOUS-like 6 (.1)
Potri.012G062300
182 / 5e-57
AT5G60910 288 / 5e-99
FRUITFULL, AGAMOUS-like 8 (.1.2)
Potri.004G115400
174 / 1e-53
AT5G60910 230 / 7e-76
FRUITFULL, AGAMOUS-like 8 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF01486
K-box
K-box region
PF00319
SRF-TF
SRF-type transcription factor (DNA-binding and dimerisation domain)
Representative CDS sequence
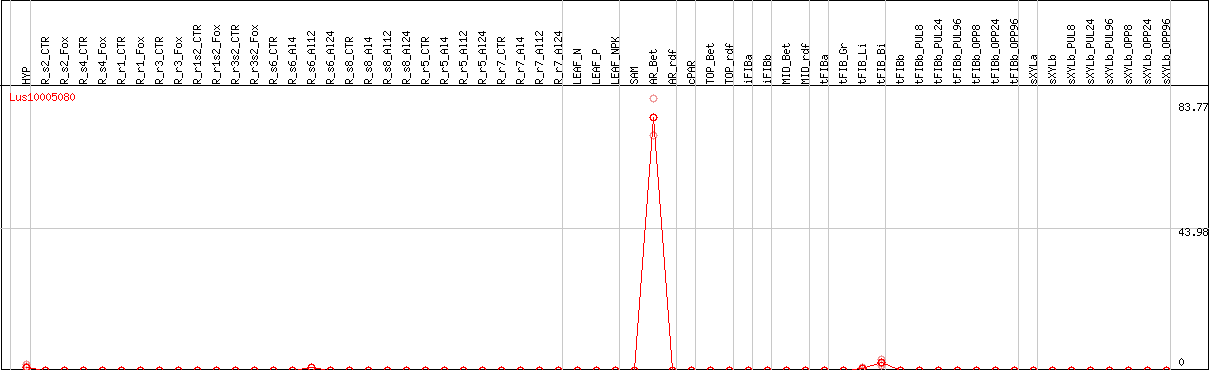
>Lus10005080 pacid=23171456 polypeptide=Lus10005080 locus=Lus10005080.g ID=Lus10005080.BGIv1.0 annot-version=v1.0
ATGGGAAGGGGGAGAGTGGAGCTGAAGAGGATAGAGAACAAGATAAACAGGCAGGTGACGTTTGCTAAGAGGAGGAATGGTTTGCTCAAGAAAGCTTATG
AGCTGTCCGTACTCTGTGATGCTGAGGTTGCTCTCATCATCTTCTCCAACCGTGGCAAGCTCTATGAGTTCTGCAGCACCAATAACATGCTCAAGACACT
GGAAAGGTATCAAAAGTGCAGCTATGGTGCAGTTGAAGTCAACAAACCTGCTAAAGAGCTCGAGAGCAGCTACCGTGAGTACTTGAAACTGAAGGGTAGA
TATGAAGCCCTACAACGAACCCAAAGGAACCTTCTGGGGGAGGATCTAGGTCCTCTGAACACAAAGGAACTGGAGCAGCTTGAGCGCTCTTTGGAAGGTT
CACTGAAGCAAGTCCGCTCCACTAAGACTCAGTACATGCTTGACCAGCTATCTGATCTTCAAAACAAGGAGCAGCTGCTGCTTGAAGCCAATAGAGCATT
GGCAATAAAGCTGGATGAAATCAGTTCAAGGAATAATCTCCGACAGTCGTGGGAAGGAGGGGAGCAGAATATGTCCTATGCTGATCACCACCAGCATCAG
GATCAGCATCAGCACCATAATCCACATGCTCAATCCCAAGGGTTGTTCCAGCATTTAGATTGCAACCCAACTTTGCAAATCGGGTATAATCCGGTAGTTT
CAGACCAGCAGATGACAGAAGCTACAACACATGGTCATCAACAGCAAGTGAATGGGTTTATTCCTGGATGGATGCTCTAA
AA sequence
>Lus10005080 pacid=23171456 polypeptide=Lus10005080 locus=Lus10005080.g ID=Lus10005080.BGIv1.0 annot-version=v1.0
MGRGRVELKRIENKINRQVTFAKRRNGLLKKAYELSVLCDAEVALIIFSNRGKLYEFCSTNNMLKTLERYQKCSYGAVEVNKPAKELESSYREYLKLKGR
YEALQRTQRNLLGEDLGPLNTKELEQLERSLEGSLKQVRSTKTQYMLDQLSDLQNKEQLLLEANRALAIKLDEISSRNNLRQSWEGGEQNMSYADHHQHQ
DQHQHHNPHAQSQGLFQHLDCNPTLQIGYNPVVSDQQMTEATTHGHQQQVNGFIPGWML