External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G69120 225 / 3e-73
MADS
AGL7, AP1
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
AT1G26310 221 / 8e-72
MADS
CAL1, AGL10, CAL
CAULIFLOWER, AGAMOUS-like 10, K-box region and MADS-box transcription factor family protein (.1)
AT5G60910 218 / 1e-70
MADS
FUL, AGL8
FRUITFULL, AGAMOUS-like 8 (.1.2)
AT3G30260 194 / 2e-61
MADS
AGL79
AGAMOUS-like 79 (.1)
AT5G15800 159 / 2e-47
MADS
AGL2, SEP1
SEPALLATA1, AGAMOUS-like 2, K-box region and MADS-box transcription factor family protein (.1.2)
AT1G24260 159 / 2e-47
MADS
AGL9, SEP3
SEPALLATA3, AGAMOUS-like 9, K-box region and MADS-box transcription factor family protein (.1.2.3)
AT2G45650 158 / 4e-47
MADS
AGL6
AGAMOUS-like 6 (.1)
AT2G03710 154 / 1e-46
MADS
AGL3, SEP4
SEPALLATA 4, AGAMOUS-like 3, K-box region and MADS-box transcription factor family protein (.1.2.3)
AT3G02310 154 / 1e-45
MADS
AGL4, SEP2
SEPALLATA 2, AGAMOUS-like 4, K-box region and MADS-box transcription factor family protein (.1)
AT5G62165 150 / 1e-44
MADS
FYF, AGL42
FOREVER YOUNG FLOWER, AGAMOUS-like 42 (.1.2.3.4.5)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10034370
365 / 6e-129
AT1G69120 224 / 1e-73
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Lus10021140
246 / 4e-81
AT5G60910 291 / 1e-99
FRUITFULL, AGAMOUS-like 8 (.1.2)
Lus10007983
245 / 4e-81
AT5G60910 298 / 3e-102
FRUITFULL, AGAMOUS-like 8 (.1.2)
Lus10034662
244 / 1e-80
AT5G60910 284 / 3e-97
FRUITFULL, AGAMOUS-like 8 (.1.2)
Lus10017871
236 / 1e-78
AT1G69120 264 / 2e-90
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Lus10026679
219 / 4e-71
AT1G69120 338 / 1e-118
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Lus10004637
218 / 5e-71
AT1G69120 318 / 2e-111
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Lus10015765
164 / 1e-49
AT1G24260 347 / 5e-122
SEPALLATA3, AGAMOUS-like 9, K-box region and MADS-box transcription factor family protein (.1.2.3)
Lus10003125
159 / 1e-48
AT1G24260 290 / 9e-101
SEPALLATA3, AGAMOUS-like 9, K-box region and MADS-box transcription factor family protein (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G115400
245 / 4e-81
AT5G60910 230 / 7e-76
FRUITFULL, AGAMOUS-like 8 (.1.2)
Potri.017G099800
238 / 2e-78
AT5G60910 255 / 7e-86
FRUITFULL, AGAMOUS-like 8 (.1.2)
Potri.012G062300
230 / 2e-75
AT5G60910 288 / 5e-99
FRUITFULL, AGAMOUS-like 8 (.1.2)
Potri.010G154100
227 / 3e-74
AT1G69120 324 / 7e-113
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Potri.008G098500
221 / 4e-72
AT1G69120 336 / 9e-118
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Potri.008G098400
166 / 2e-50
AT5G15800 275 / 2e-93
SEPALLATA1, AGAMOUS-like 2, K-box region and MADS-box transcription factor family protein (.1.2)
Potri.014G074100
164 / 1e-49
AT2G45650 311 / 5e-108
AGAMOUS-like 6 (.1)
Potri.004G115500
160 / 3e-48
AT3G02310 357 / 4e-126
SEPALLATA 2, AGAMOUS-like 4, K-box region and MADS-box transcription factor family protein (.1)
Potri.012G132600
159 / 1e-47
AT2G45650 182 / 3e-57
AGAMOUS-like 6 (.1)
Potri.001G058400
158 / 3e-47
AT1G24260 362 / 8e-128
SEPALLATA3, AGAMOUS-like 9, K-box region and MADS-box transcription factor family protein (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF01486
K-box
K-box region
PF00319
SRF-TF
SRF-type transcription factor (DNA-binding and dimerisation domain)
Representative CDS sequence
>Lus10005081 pacid=23171473 polypeptide=Lus10005081 locus=Lus10005081.g ID=Lus10005081.BGIv1.0 annot-version=v1.0
ATGGGGAGAGGAAGGGTAGAGATGAGGAGGATAGAGAACAAGATAAGCAGGCAAGTGACCTTCTCCAAGAGGAGAAGTGGTCTTTTGAAAAAAGCTCATG
AGATCTCGGTTCTTTGTGATGCTCAGCTAGCCCTCATTGTTTTCTCCACTAAGGGCAAGCTCTTTGAGTACTCCTCTGATGATTCCAGCATGGACATGAT
ACTGGAGAGATACGAGAGGTACTCGTATGCTGAAAGAAACCATGAGCTTCCCATTACCGATCCTCAACAACACCCGGGAAGTAGCTGGACTATGGAATGT
CCAAAGCTCATGTCTAGGATTGAAGTGTTGCAGAGAAACCTCAGGAACTTGGTAGGAGAAGAAATTGATCCGATGAGTCTAAGGGAGCTTCAGCAGCTGG
AGCAGCAAATTGATACTGCTCTTAAGAAAGTCAGATCAAGAAAGAACCAACTCTACCATGAATCCATCTCTAATCTCCAGAAAAAGGTGCAAGAACGTTC
CTTGAACAAACAAAACGGCGTGCTAGCCAAAAAGGTGAAACAAAATGAGAAGGAAGTGGCAACAACTGGGAATGAAGAGCAGCAGCAGCAGCAGCAGAAC
AATGAATATGAGAACGAGCCTCGTCGACTTGAGTCGCAACCAATGCTTCAACTTCCTCCTCTGACTATTGGGTGGGCTCTGTTGTGCAGTACATATGGGT
TCCAAGCTAGAGGGCAAATGATGAATGCGAATGAGAACATTGAAGAAGAAGAAGAAGCTGTTGAAAGACAGGATCCGCCAACCAGAGTCGTCACTAAAAC
TACTACCCATTTTCCAGCCTGGATGCTTCCGTATCTCCGTGATCAATAG
AA sequence
>Lus10005081 pacid=23171473 polypeptide=Lus10005081 locus=Lus10005081.g ID=Lus10005081.BGIv1.0 annot-version=v1.0
MGRGRVEMRRIENKISRQVTFSKRRSGLLKKAHEISVLCDAQLALIVFSTKGKLFEYSSDDSSMDMILERYERYSYAERNHELPITDPQQHPGSSWTMEC
PKLMSRIEVLQRNLRNLVGEEIDPMSLRELQQLEQQIDTALKKVRSRKNQLYHESISNLQKKVQERSLNKQNGVLAKKVKQNEKEVATTGNEEQQQQQQN
NEYENEPRRLESQPMLQLPPLTIGWALLCSTYGFQARGQMMNANENIEEEEEAVERQDPPTRVVTKTTTHFPAWMLPYLRDQ
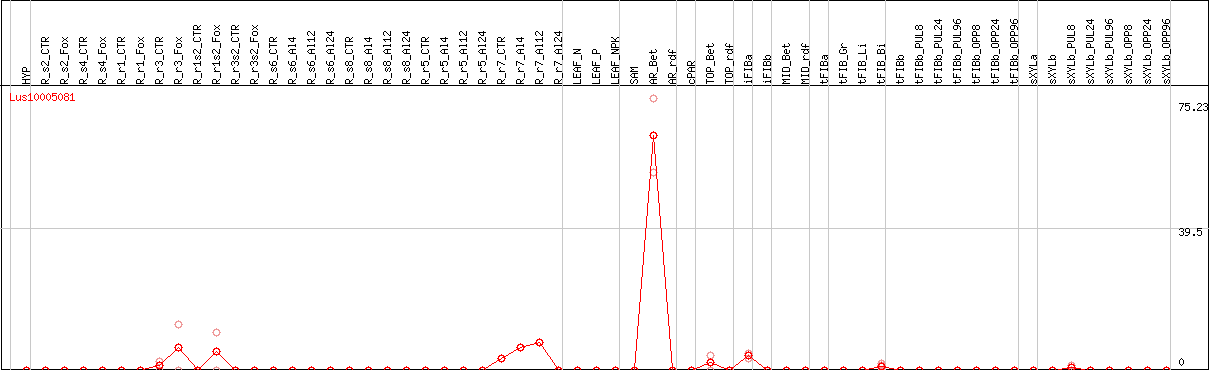
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10005081 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.