External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G10880 328 / 3e-108
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT5G16170 320 / 6e-108
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT5G11730 314 / 7e-106
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT1G10280 314 / 2e-105
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT1G51770 312 / 4e-105
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1.2)
AT3G21310 310 / 2e-104
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT5G25970 306 / 5e-103
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT1G68390 286 / 1e-94
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT1G73810 271 / 7e-89
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT1G68380 263 / 8e-86
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005504
314 / 1e-105
AT5G11730 499 / 1e-177
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10005505
314 / 1e-105
AT5G11730 499 / 1e-177
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10007457
314 / 1e-103
AT5G11730 489 / 1e-170
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10035828
306 / 4e-102
AT1G68390 401 / 7e-138
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10036610
305 / 8e-102
AT1G68390 404 / 3e-139
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10002989
302 / 6e-101
AT1G10280 525 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10036774
301 / 6e-101
AT5G16170 350 / 1e-118
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10024468
301 / 1e-100
AT1G10280 524 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10009738
295 / 3e-98
AT3G21310 511 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G121900
410 / 2e-143
AT5G11730 409 / 1e-141
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.015G045500
354 / 2e-121
AT5G16170 394 / 2e-135
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.004G097700
348 / 3e-119
AT5G16170 452 / 6e-159
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.006G233400
325 / 3e-110
AT5G11730 557 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.018G059100
312 / 2e-105
AT5G11730 548 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.015G045400
311 / 1e-104
AT1G68390 390 / 2e-134
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.010G121800
310 / 2e-104
AT1G68390 399 / 1e-137
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.001G195900
307 / 2e-103
AT3G21310 504 / 8e-180
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.003G001700
304 / 8e-102
AT1G10280 543 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.004G228100
296 / 8e-99
AT1G10280 512 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0110
GT-A
PF02485
Branch
Core-2/I-Branching enzyme
Representative CDS sequence
>Lus10005108 pacid=23171478 polypeptide=Lus10005108 locus=Lus10005108.g ID=Lus10005108.BGIv1.0 annot-version=v1.0
ATGAGCGACGATGAGCTATTTCAGCGAGCATCCATGGATCCACACATCTCCAACAACTATCCTTTTATTCCTACACCAAAACTTGCCTTCATGTTCTTAA
TCAAGGATGGAAACTTCCCTTATCTCGCTCCCTTATGGGAGTTGTTCTTCAAAGGTCACAACCAATCTTTTTATTCCATCTATGTCCATGTTTCCCGTCT
TCATGATCCTACCAACTTATCCACATCACAACCACCAATATCCTCTGTCTTCTACAACCGGATTATCCCGAGTAAGCCGGTCCGATGGGGGACATCGACC
ATGGTCGATGCAGAAAGGAGGCTACTCGCCAACGCCCTCCTCGACTTCTCCAACCAACGGTTTGTCCTCCTCTCCGAGTCTTGCATCCCTTTATTCAATT
TCACCACCATCTACAACTACCTAGTTTTCAGTTTCTCCAACACGAGCTTCTTGGGCTCGATTGATGATCCCCGACCCATGGGCCGAGGCAGATACAACAA
GCGTATGCAGCCAAACATCACGCTATCAGACTGGCGCAAAGGTAGCCAATGGTTCGTAGTTCATCACAAGGTAGCCATTCTGATGGTCTCTGATGTTAAA
TACTACCATATTTTCGAACAACATTGTAAGCCTCCGTGTTACATGGATGAACACTATTTCCCGACCCTTGTGACCAAACTTTTGCTCGAGATGAACTCGA
ACCATAGCATCACTTGGGTTGATTGGTCCAAAGGTGGGTCACATCCTACCATGTTTTTGAGGAAGGATGTGAGCCTAGGGTTCTTGATGAGGATCAGAAA
CATGTCCGATTGTGCTTATAGTGATCAAAACCATGGAACAATTTGTTTCCTCTTTGCTAGAAAGTTTCACCCTGGTACTTTAGAGCCTTTGTTGAGGGTA
GCTCCATCTTTGTTTGGTTTAAACACTTAG
AA sequence
>Lus10005108 pacid=23171478 polypeptide=Lus10005108 locus=Lus10005108.g ID=Lus10005108.BGIv1.0 annot-version=v1.0
MSDDELFQRASMDPHISNNYPFIPTPKLAFMFLIKDGNFPYLAPLWELFFKGHNQSFYSIYVHVSRLHDPTNLSTSQPPISSVFYNRIIPSKPVRWGTST
MVDAERRLLANALLDFSNQRFVLLSESCIPLFNFTTIYNYLVFSFSNTSFLGSIDDPRPMGRGRYNKRMQPNITLSDWRKGSQWFVVHHKVAILMVSDVK
YYHIFEQHCKPPCYMDEHYFPTLVTKLLLEMNSNHSITWVDWSKGGSHPTMFLRKDVSLGFLMRIRNMSDCAYSDQNHGTICFLFARKFHPGTLEPLLRV
APSLFGLNT
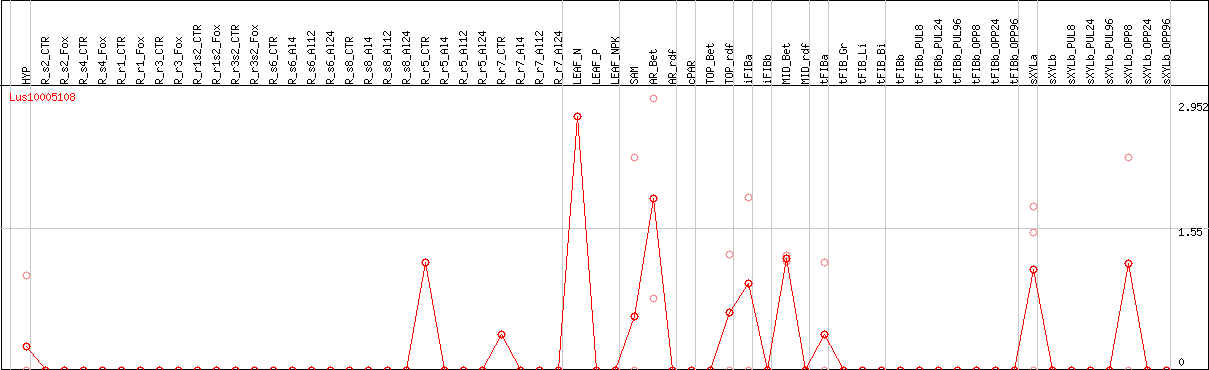
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10005108 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.