External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G41540 48 / 2e-07
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT1G27170 48 / 2e-07
transmembrane receptors;ATP binding (.1.2)
AT5G17680 47 / 3e-07
disease resistance protein (TIR-NBS-LRR class), putative (.1)
AT4G12010 47 / 3e-07
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT1G65850 47 / 4e-07
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
AT5G41550 47 / 6e-07
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G45220 46 / 6e-07
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT1G09665 45 / 6e-07
Toll-Interleukin-Resistance (TIR) domain family protein (.1)
AT4G16990 46 / 1e-06
RLM3
RESISTANCE TO LEPTOSPHAERIA MACULANS 3, disease resistance protein (TIR-NBS class), putative
AT4G09430 45 / 1e-06
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026845
109 / 2e-30
AT5G36930 145 / 8e-39
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10041606
103 / 8e-28
AT5G36930 140 / 5e-37
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10027920
91 / 6e-23
AT5G36930 139 / 3e-36
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10012063
86 / 1e-21
AT5G36930 124 / 5e-34
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10041605
81 / 4e-20
AT1G72940 93 / 3e-22
Toll-Interleukin-Resistance (TIR) domain-containing protein (.1)
Lus10004482
59 / 2e-11
AT1G56540 111 / 2e-27
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Lus10029919
58 / 3e-11
AT5G36930 105 / 5e-26
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10029921
54 / 2e-09
AT5G36930 105 / 7e-26
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10019142
51 / 2e-08
AT1G72890 140 / 1e-36
Disease resistance protein (TIR-NBS class) (.1), Disease resistance protein (TIR-NBS class) (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G097050
54 / 4e-10
AT5G36930 158 / 3e-45
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G019710
55 / 7e-10
AT5G36930 445 / 2e-144
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G069200
55 / 7e-10
AT5G17680 590 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.005G206400
54 / 1e-09
AT5G36930 587 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.T126506
54 / 2e-09
AT3G14470 575 / 0.0
NB-ARC domain-containing disease resistance protein (.1)
Potri.019G003285
54 / 2e-09
AT5G36930 640 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G003942
53 / 3e-09
AT5G36930 591 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.005G004233
52 / 3e-09
AT1G27170 153 / 8e-42
transmembrane receptors;ATP binding (.1.2)
Potri.013G098100
52 / 3e-09
AT5G36930 159 / 1e-45
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.T126306
52 / 5e-09
AT3G14470 585 / 0.0
NB-ARC domain-containing disease resistance protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0173
STIR
PF01582
TIR
TIR domain
Representative CDS sequence
>Lus10005174 pacid=23163366 polypeptide=Lus10005174 locus=Lus10005174.g ID=Lus10005174.BGIv1.0 annot-version=v1.0
ATGTCAATCCTGGTTCTCTCGAGGGGATACGCCTCCTCGAAGTGGTGTTTGGACGAGCTGGTGAAGATCATGGAGGCCGCCAATGACAAGGCTCGTCTCC
ACCAGCCTTTCCCTTTTTTCTACAAAGTGTCCGATCAGTCATCGGGGTGCGACAAGGAAGACTTCGATATGCACAAGAGAAGGTTCAGCTACACGCAAAT
TGGTCGAATGCAGCAGGAATCGACCGGCGACAAACGCACCTGTTACCAAACCTGCAACGCCGCCGAGTTCTTGTCCGCGGTCTGCCATCGTCCCCGAATC
GCGGGTGTCTTCCGCATGTTGATCGGATAA
AA sequence
>Lus10005174 pacid=23163366 polypeptide=Lus10005174 locus=Lus10005174.g ID=Lus10005174.BGIv1.0 annot-version=v1.0
MSILVLSRGYASSKWCLDELVKIMEAANDKARLHQPFPFFYKVSDQSSGCDKEDFDMHKRRFSYTQIGRMQQESTGDKRTCYQTCNAAEFLSAVCHRPRI
AGVFRMLIG
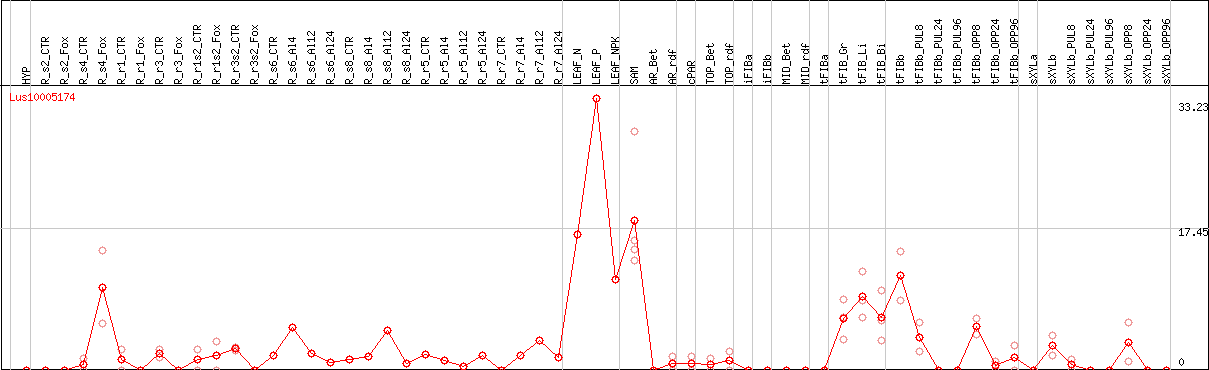
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10005174 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.