External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G09960 303 / 9e-105
MADS
AGL11, STK
SEEDSTICK, AGAMOUS-like 11, K-box region and MADS-box transcription factor family protein (.1.2.3.4)
AT3G58780 260 / 3e-87
MADS
AGL1, SHP1
SHATTERPROOF 1, AGAMOUS-like 1, K-box region and MADS-box transcription factor family protein (.1.2.3)
AT4G18960 259 / 4e-87
MADS
AG
AGAMOUS, K-box region and MADS-box transcription factor family protein (.1.2)
AT2G42830 256 / 5e-86
MADS
AGL5, SHP2
SHATTERPROOF 2, AGAMOUS-like 5, K-box region and MADS-box transcription factor family protein (.1.2)
AT4G22950 149 / 3e-44
MADS
AGL19
AGAMOUS-like 19 (.1)
AT5G62165 144 / 2e-42
MADS
FYF, AGL42
FOREVER YOUNG FLOWER, AGAMOUS-like 42 (.1.2.3.4.5)
AT2G45650 144 / 7e-42
MADS
AGL6
AGAMOUS-like 6 (.1)
AT2G45660 142 / 7e-42
MADS
ATSOC1, SOC1, AGL20
SUPPRESSOR OF OVEREXPRESSION OF CO 1, AGAMOUS-like 20 (.1)
AT3G57230 142 / 2e-41
MADS
AGL16
AGAMOUS-like 16 (.1.2)
AT2G03710 137 / 4e-40
MADS
AGL3, SEP4
SEPALLATA 4, AGAMOUS-like 3, K-box region and MADS-box transcription factor family protein (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10008264
415 / 6e-148
AT4G09960 328 / 3e-114
SEEDSTICK, AGAMOUS-like 11, K-box region and MADS-box transcription factor family protein (.1.2.3.4)
Lus10009481
298 / 1e-101
AT4G09960 274 / 1e-92
SEEDSTICK, AGAMOUS-like 11, K-box region and MADS-box transcription factor family protein (.1.2.3.4)
Lus10004638
139 / 2e-40
AT3G02310 246 / 2e-82
SEPALLATA 2, AGAMOUS-like 4, K-box region and MADS-box transcription factor family protein (.1)
Lus10006715
140 / 3e-40
AT4G22950 223 / 1e-73
AGAMOUS-like 19 (.1)
Lus10034663
139 / 1e-39
AT5G15800 267 / 4e-90
SEPALLATA1, AGAMOUS-like 2, K-box region and MADS-box transcription factor family protein (.1.2)
Lus10015765
137 / 2e-39
AT1G24260 347 / 5e-122
SEPALLATA3, AGAMOUS-like 9, K-box region and MADS-box transcription factor family protein (.1.2.3)
Lus10026678
136 / 3e-39
AT5G15800 225 / 3e-74
SEPALLATA1, AGAMOUS-like 2, K-box region and MADS-box transcription factor family protein (.1.2)
Lus10031665
133 / 6e-39
AT5G62165 179 / 8e-58
FOREVER YOUNG FLOWER, AGAMOUS-like 42 (.1.2.3.4.5)
Lus10037040
136 / 1e-38
AT1G24260 350 / 7e-123
SEPALLATA3, AGAMOUS-like 9, K-box region and MADS-box transcription factor family protein (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF01486
K-box
K-box region
PF00319
SRF-TF
SRF-type transcription factor (DNA-binding and dimerisation domain)
Representative CDS sequence
>Lus10005302 pacid=23168817 polypeptide=Lus10005302 locus=Lus10005302.g ID=Lus10005302.BGIv1.0 annot-version=v1.0
ATGGATCTCTCCTACTCAATTCCTCAGGAACAAGAAGTGGCAGAAGATCAAGAGGAGGTCATAGTCAACAACAACAGCAGCAGCAGCAGCATGGGGAGGG
GAAAGATCGAGATCAAGAGGATCGAAAACACCACGAACCGACAGGTGACATTCTGCAAGCGAAGGAACGGACTTTTGAAGAAAGCCTATGAGCTTTCCGT
GCTCTGTGATGCTGAGGTTGCTCTCATCGTCTTCTCCACCCGCGGCCGCCTCTACGAGTACTCCAACAACAACATCAAGGCCACCATAGAGAAATACAAG
AAGGCATGTTCTGACAGTACCCACACTCCCTCCGTTACTGAAATCAACGCCCAGTATTATCAGCAGGAGTCAGCTAAGCTGCGACAACAGATCCAAATGC
TGCAGAATTCCAACAGGCACCTCATGGGGGAATCTTTGAGCTCACTCTCTATAAAGGAGCTGAAGCAGCTGGAGAACAGACTTGACCGAGGAGTCAGCAG
AATCAGGGCCAAGAAGCATGAGCTTCTGCTAGCTGAAATCGAATATATGCAGAAAAGGGAGATTGAGCTGGAGAATGAAAGTGTGTGTCTTCGTACCAAG
ATAGCGGAGGTGGAGAGGGTGAATGAGGAAGCGAACATGGTGAGTGGAGCGGAGCTGAATGCAATCCACGCACTGGCTGCGTCTCGCAACTTCTTTGCTC
CAAATGAAATGATGGATCATCCTCCTCCGACGTCGGCGAAAGAACATGACGCATCCCCCCCCCCCGCCGTCGGCGACAAGCACCCCTTTCTCTCCCGGCA
GTAG
AA sequence
>Lus10005302 pacid=23168817 polypeptide=Lus10005302 locus=Lus10005302.g ID=Lus10005302.BGIv1.0 annot-version=v1.0
MDLSYSIPQEQEVAEDQEEVIVNNNSSSSSMGRGKIEIKRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALIVFSTRGRLYEYSNNNIKATIEKYK
KACSDSTHTPSVTEINAQYYQQESAKLRQQIQMLQNSNRHLMGESLSSLSIKELKQLENRLDRGVSRIRAKKHELLLAEIEYMQKREIELENESVCLRTK
IAEVERVNEEANMVSGAELNAIHALAASRNFFAPNEMMDHPPPTSAKEHDASPPPAVGDKHPFLSRQ
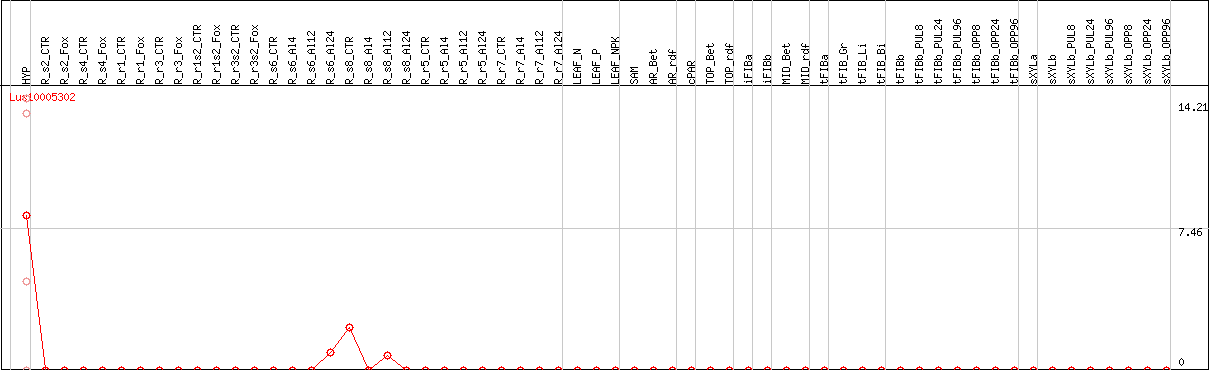
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10005302 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.