External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G01390 92 / 8e-23
UDP-Glycosyltransferase superfamily protein (.1)
AT4G01070 89 / 7e-22
UGT72B1, GT72B1
UDP-GLUCOSE-DEPENDENT GLUCOSYLTRANSFERASE 72 B1, UDP-Glycosyltransferase superfamily protein (.1.2)
AT1G01420 81 / 1e-18
UGT72B3
UDP-glucosyl transferase 72B3 (.1)
AT2G18570 66 / 2e-13
UDP-Glycosyltransferase superfamily protein (.1)
AT5G66690 66 / 2e-13
UGT72E2
UDP-Glycosyltransferase superfamily protein (.1)
AT2G18560 64 / 1e-12
UDP-Glycosyltransferase superfamily protein (.1)
AT5G26310 61 / 1e-11
UGT72E3
UDP-Glycosyltransferase superfamily protein (.1)
AT3G50740 58 / 9e-11
UGT72E1
UDP-glucosyl transferase 72E1 (.1)
AT3G16520 56 / 8e-10
UGT88A1
UDP-glucosyl transferase 88A1 (.1.2.3)
AT3G21760 48 / 4e-07
HYR1
HYPOSTATIN RESISTANCE 1, UDP-Glycosyltransferase superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10039588
250 / 1e-82
AT4G01070 419 / 2e-143
UDP-GLUCOSE-DEPENDENT GLUCOSYLTRANSFERASE 72 B1, UDP-Glycosyltransferase superfamily protein (.1.2)
Lus10029452
117 / 1e-31
AT4G01070 582 / 0.0
UDP-GLUCOSE-DEPENDENT GLUCOSYLTRANSFERASE 72 B1, UDP-Glycosyltransferase superfamily protein (.1.2)
Lus10005950
114 / 8e-31
AT4G01070 573 / 0.0
UDP-GLUCOSE-DEPENDENT GLUCOSYLTRANSFERASE 72 B1, UDP-Glycosyltransferase superfamily protein (.1.2)
Lus10029453
94 / 4e-23
AT4G01070 515 / 0.0
UDP-GLUCOSE-DEPENDENT GLUCOSYLTRANSFERASE 72 B1, UDP-Glycosyltransferase superfamily protein (.1.2)
Lus10005951
92 / 1e-22
AT4G01070 510 / 5e-179
UDP-GLUCOSE-DEPENDENT GLUCOSYLTRANSFERASE 72 B1, UDP-Glycosyltransferase superfamily protein (.1.2)
Lus10024037
88 / 6e-21
AT2G18570 412 / 3e-140
UDP-Glycosyltransferase superfamily protein (.1)
Lus10041713
80 / 2e-18
AT2G18570 415 / 3e-141
UDP-Glycosyltransferase superfamily protein (.1)
Lus10003944
72 / 2e-15
AT4G01070 400 / 1e-135
UDP-GLUCOSE-DEPENDENT GLUCOSYLTRANSFERASE 72 B1, UDP-Glycosyltransferase superfamily protein (.1.2)
Lus10001906
69 / 3e-14
AT4G01070 431 / 1e-147
UDP-GLUCOSE-DEPENDENT GLUCOSYLTRANSFERASE 72 B1, UDP-Glycosyltransferase superfamily protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G168600
102 / 1e-26
AT4G01070 595 / 0.0
UDP-GLUCOSE-DEPENDENT GLUCOSYLTRANSFERASE 72 B1, UDP-Glycosyltransferase superfamily protein (.1.2)
Potri.014G096100
102 / 2e-26
AT4G01070 610 / 0.0
UDP-GLUCOSE-DEPENDENT GLUCOSYLTRANSFERASE 72 B1, UDP-Glycosyltransferase superfamily protein (.1.2)
Potri.014G096000
96 / 6e-24
AT4G01070 544 / 0.0
UDP-GLUCOSE-DEPENDENT GLUCOSYLTRANSFERASE 72 B1, UDP-Glycosyltransferase superfamily protein (.1.2)
Potri.014G041900
87 / 9e-21
AT4G01070 295 / 2e-95
UDP-GLUCOSE-DEPENDENT GLUCOSYLTRANSFERASE 72 B1, UDP-Glycosyltransferase superfamily protein (.1.2)
Potri.007G030500
86 / 3e-20
AT2G18570 494 / 8e-173
UDP-Glycosyltransferase superfamily protein (.1)
Potri.014G041800
80 / 3e-18
AT4G01070 390 / 9e-132
UDP-GLUCOSE-DEPENDENT GLUCOSYLTRANSFERASE 72 B1, UDP-Glycosyltransferase superfamily protein (.1.2)
Potri.003G138200
78 / 1e-17
AT4G01070 539 / 0.0
UDP-GLUCOSE-DEPENDENT GLUCOSYLTRANSFERASE 72 B1, UDP-Glycosyltransferase superfamily protein (.1.2)
Potri.007G030400
77 / 4e-17
AT3G50740 566 / 0.0
UDP-glucosyl transferase 72E1 (.1)
Potri.007G030300
63 / 3e-12
AT3G50740 496 / 1e-173
UDP-glucosyl transferase 72E1 (.1)
Potri.007G029800
63 / 3e-12
AT3G50740 478 / 3e-166
UDP-glucosyl transferase 72E1 (.1)
PFAM info
Representative CDS sequence
>Lus10005335 pacid=23154621 polypeptide=Lus10005335 locus=Lus10005335.g ID=Lus10005335.BGIv1.0 annot-version=v1.0
ATGCCCCATCTCCGCTACGCGATCAGGTCGCTGTCGGAGAAATTTCCTCTATCGTCCTTGATCGCCGACATATTCGGCACCGACGCATTCGACGTGGCCA
GGGAGTTCAAATTGGAGTCGTATTTTTTCGTGACGTTGAACGTTTTGACTCTGGCGCTATGCAACTATATGTCGAAGCTGGATGCCGAGGTTCAAGGGGA
CTACCATCAACTGACCGAACCGATTCGGCTGCCAGGGTTCCGGTTCGTGTTTCCGGTTGAGGACCTGCACCCTTCGATCCTAGACAGGAACATCGATGCT
TACCCGATGCTGCTTCGTCACTCGAGCCGGCAGCGTTTGGCTGACGGGTTTATAGTGAACAGCTTCATGGAGGTGGAAGTAGAGATCATCGAAGCTTTGA
GAGGGTAG
AA sequence
>Lus10005335 pacid=23154621 polypeptide=Lus10005335 locus=Lus10005335.g ID=Lus10005335.BGIv1.0 annot-version=v1.0
MPHLRYAIRSLSEKFPLSSLIADIFGTDAFDVAREFKLESYFFVTLNVLTLALCNYMSKLDAEVQGDYHQLTEPIRLPGFRFVFPVEDLHPSILDRNIDA
YPMLLRHSSRQRLADGFIVNSFMEVEVEIIEALRG
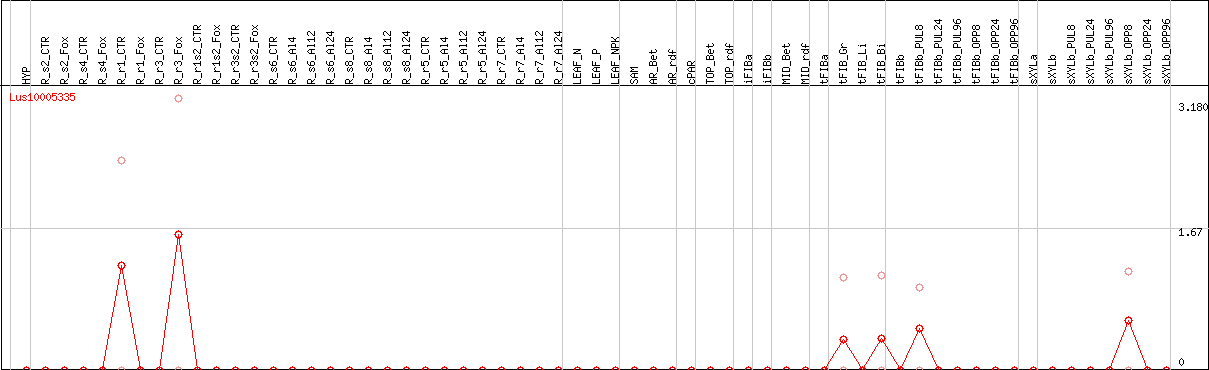
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10005335 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.