External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G17615 127 / 2e-35
Disease resistance protein (TIR-NBS class) (.1)
AT1G72920 117 / 2e-32
Toll-Interleukin-Resistance (TIR) domain family protein (.1)
AT1G72940 115 / 5e-31
Toll-Interleukin-Resistance (TIR) domain-containing protein (.1)
AT1G72900 113 / 4e-30
Toll-Interleukin-Resistance (TIR) domain-containing protein (.1)
AT5G36930 115 / 6e-30
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
AT1G27170 115 / 6e-30
transmembrane receptors;ATP binding (.1.2)
AT1G72950 112 / 9e-30
Disease resistance protein (TIR-NBS class) (.1)
AT1G72930 105 / 3e-29
TIR
toll/interleukin-1 receptor-like (.1.2)
AT1G72910 108 / 3e-28
Toll-Interleukin-Resistance (TIR) domain-containing protein (.1)
AT1G72890 107 / 1e-27
Disease resistance protein (TIR-NBS class) (.1), Disease resistance protein (TIR-NBS class) (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005373
295 / 5e-94
AT5G36930 404 / 8e-121
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10020533
246 / 8e-76
AT1G27170 405 / 1e-122
transmembrane receptors;ATP binding (.1.2)
Lus10008415
222 / 7e-75
AT5G36930 137 / 5e-38
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10020524
237 / 7e-73
AT1G27170 395 / 9e-119
transmembrane receptors;ATP binding (.1.2)
Lus10020536
236 / 2e-72
AT1G27170 392 / 8e-118
transmembrane receptors;ATP binding (.1.2)
Lus10001040
216 / 3e-71
AT5G36930 167 / 2e-47
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10020534
233 / 4e-71
AT5G36930 414 / 1e-124
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10020537
233 / 6e-71
AT1G27170 404 / 6e-121
transmembrane receptors;ATP binding (.1.2)
Lus10008027
212 / 5e-69
AT5G36930 197 / 5e-57
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.019G019710
150 / 4e-43
AT5G36930 445 / 2e-144
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G017082
151 / 1e-42
AT5G36930 638 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.T002200
150 / 3e-42
AT5G36930 617 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.003G014200
150 / 5e-42
AT5G36930 673 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.T002568
146 / 1e-41
AT5G36930 439 / 2e-142
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.T002400
148 / 2e-41
AT5G36930 658 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.T002428
145 / 2e-41
AT5G36930 387 / 2e-122
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G052000
147 / 4e-41
AT5G36930 641 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.T001933
147 / 4e-41
AT5G36930 617 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G016425
146 / 9e-41
AT5G36930 650 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0173
STIR
PF01582
TIR
TIR domain
Representative CDS sequence
>Lus10005374 pacid=23148511 polypeptide=Lus10005374 locus=Lus10005374.g ID=Lus10005374.BGIv1.0 annot-version=v1.0
ATGATGAAATCCGAATATGAGAGCTCCTCCTCCGCGGTGGACCCGTCCATCCTCCCCCGTACACCGGGAGAGTACGAGGTTTTCCTGAGCTTCAGAGGGG
GAGATGTCCGCAGAACCTTTGCGGACCATCTCTACAACTACCTTACGGATTTGAAAATCCGCACATTTCGAGATGAGGAAGAGCTCCCGAACGGGGAGTT
CATCGCCCAGGCTCTAACCAAAGCTATTACCGAATCCAAGGTCTACGTCCCGATCTTCTCGCGGAACTATGCTGGAAGCAAATGGTGCCTTCAGGAGCTG
GCTAAGATGGTGGAGTGCTGGAAGGCCGGGAAGGGAGGAAAAGGACAGCTGATTTTCCTCCCCGTTTTCTATATGATTGATCCGAGAAGCGTGAGGCATT
CCACGTCTGAGCCTTACAAGGCGGCGTTCAAAGAACATCGCCGGAAGCATGACGCTCGAACCGTGCCGGCGTGGAAGAAAGCACTCCAAGTGGTTGGTAA
AATGAAAGGATGGCACATCACTGAGTTAGATCATGATTCGAGTAGTTAA
AA sequence
>Lus10005374 pacid=23148511 polypeptide=Lus10005374 locus=Lus10005374.g ID=Lus10005374.BGIv1.0 annot-version=v1.0
MMKSEYESSSSAVDPSILPRTPGEYEVFLSFRGGDVRRTFADHLYNYLTDLKIRTFRDEEELPNGEFIAQALTKAITESKVYVPIFSRNYAGSKWCLQEL
AKMVECWKAGKGGKGQLIFLPVFYMIDPRSVRHSTSEPYKAAFKEHRRKHDARTVPAWKKALQVVGKMKGWHITELDHDSSS
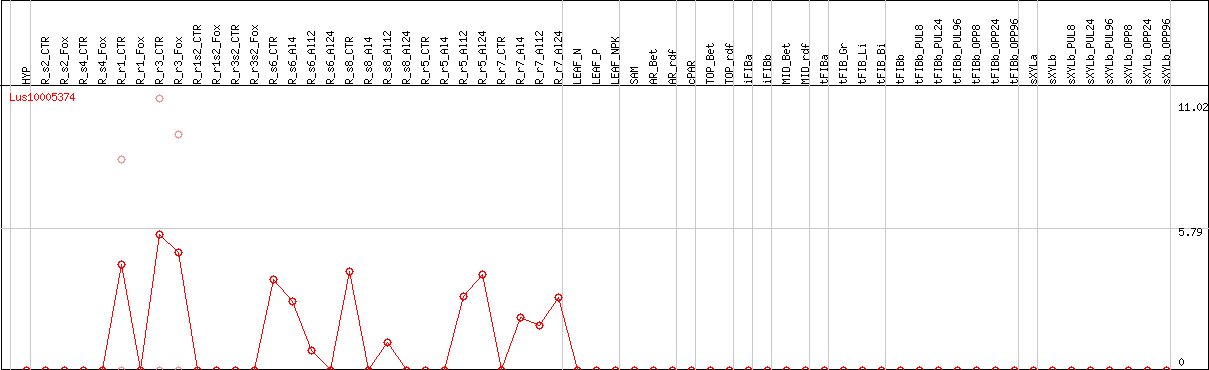
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10005374 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.