Lus10005385 [FLAX]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Poplar homologues |
|
|||||||||||||||
| PFAM info |
| |||||||||||||||
|
Representative CDS sequence |
>Lus10005385 pacid=23155609 polypeptide=Lus10005385 locus=Lus10005385.g ID=Lus10005385.BGIv1.0 annot-version=v1.0
ATGCCGAAGAACGTGTACAGGAACAGCTCACTCGCATGGAAGCCCTCTCCAGTTGTAGCCCTAGCTACCAGTGCCGATGACTCCCAAGTTGCCGCCGCTA
GGGAGGATGGCTCCATGGAGATTTGGCTTGTCTCGCCTGGCTCAGTGGGGTGGCACTGCCAGCTGACAATTCATGGCGATTCTAGTTCGAGGGTTTCGTC
GCTCGTGTGGTGTCGCACTGGCTCAAAGGGATTGCCTTGTGGTAGATTGTTCTCATCAAGCATTGATGGCTCTGTATCAGAATGGGACCTCTTTCACTTG
GAGCAGAAGATTGTTCTGGAGTCTATTGGCTTCTCAATTTGGCAGATGTCTGTTGCACCTCAATGTGCTTCACTGATTCACCCGGAGGTCAAATCTCTGG
ACTTGGGAAATGGCTTCTTGAATCACAGTCATAATGACGACCACGATGATGAATCAAGTGAAAGTGATAATGACTCAAGTTCTGAGGAGCTTCACAATCC
AGTTTCCGCAGATACACGCTTGGCTATTGCATGTGATGATGGTTGTGTGCGTCTATACACTATTCCTGAGTCAGATGAGTTGATTTACAATAGAACATTG
TCTAGGATAAGCCATAAAGGCGATATCAAGCTCATAGCTTTGGTCGTCAATACTCTTCATCATCCTCGAATGAGAAGAGTTGCTGAAGCAGTGATTTGCC
GATTTCGCAATCGGCTTAAAGAAAGTTTCGATTTCGCAATAGGCAGAGGTGAAGTAGTCATCTTGGTTCGGTGGTCAGCGTTGGCAGCGGCAGGAAGGAG
TGAGGACCGGCGGTGGCAGGCTCGGCGTCGATCGTCTGCACAGAGAGGATATGGAAGGAGGATGACGGCCAGCTGCAAGTTTGGCGTCGGAGGGAGGAGT
GAGGACCGGTTGCAAGCTCGACGTCGGTGGGAAGCTCGGCAGTGGCAGAGTCTGTTGACGGTAGCTTCAATCTCGATCAAATATCTGTATTTGGGAAATG
GCTTCTTGAATACCAGTCATAATGACGACCACGATGATGAATCAAGTGAAAGTGATAATGACTCAAGTTCTGAGGAGCTTCACAATCCAGTTTCCGCAGA
TACACGCTTGGCTATTGCATGTGATGATGGTTGTGTGCGTCTATACACTATTCCTGAGTCAGATGAGTTGATTTACAATAGAACATTGTCTAGGGTCAGT
GGACGCACTTTGAGTGTGACATGGAGTTCGGATGCTACTAGGATTTTCTCAGGCAGCAGTGATGGTTTTGTTAGATGCTGGGATGCTTCACTTGGCCATG
AGATATATCGGATAACAGTTGGGCTTGGTGGTCTGGGTGGTGGAACTGAAATTCGTGTGTGGTCTCTACTTGCCCTGAGATGTGGGACACTCGTTAGTGC
AGACAGTACTGGAACCGTCATTCTCTATAAGCTTTCTGGTGAGATAGGATCTAGTGATGATACATCTACATCCACGGTGATGAGGAGGTGGGTGTATGTG
GGTTATGTCAGAGCTCATACACATGATATTAGAGCCTTGACAATGGCTGTACCAATCTGCCGTGAAGATGCACTACCTGAGGAAAAGGTGAAGAAAGACC
GGAAGAGGAGGCGTCATGTTGAATTTAGCTATCACAAGTGGGCGCATTTAGGAGTTCCCATGCTTATATCTGCCAGTGATGACACAAAGTTGTTCGCATA
TTCTGCGAAGGAGTTCTCTAAGTTCTCGCCACATGATATTTGTCCTGCACCTCAGAGAATGCCAATCCACTTGGTCCCTAATACAATCTCCAACAAGAAC
CCTTTACTTCTTGTCCAGGCATCTAGTACTTTGGATATTCTTCGTGTCCGCACAGAGGATGGAAACATCTCAGATAGTGTTTCTGGTACATCAAAGGGCC
TTGCAAAGACAGTTTTGAGGGCTCAAATATTATCTAAAAAGAAGATCATATGCAGCTGCATTAGCAATTCTGGAATGCTACTTGCTTACTCGGATCACAA
GAAGTCCAATTTATTTGATTTGAAGATGCATGTTGATCCTAAAATTCCATGGAAAGTTACCAAAAGGGAAGCTAGCGAAGAGGAAGCTTGCAAGGAGTTT
CCATATGCACATTGTATGGCCTTTAGTTCGGATTCTTCAAGGCTAATATTGGCAGGCCATAACAGAAGGATATATATTGTGGATGTGGATAGCTTGAAAA
TAGTGCATACCTTCACACCGCAGCGTCACGATTACAATGAAGAATCACCCAGTGAGCCGCCTGTGACAAAACTGTTTACTAGCAATGATGGGCAGTGGTT
AGCTGCTATTAATTGCTTTGGTGATATATATGTGTTCGACATGGAGACACAAAGGCAGCATTGGTTCATCTCAAGACTAGATGGTGCCTCAGTTACTGCT
GGAGGTTTCCCCCCACAAAATAACAATGTGCTTGTAGTCACTACCTCCACGAACCAAGTCTATGCATTTGACGTGGAAGCAAAACAGTTAGGGGAATGGT
CACAACGGAATTCATCTTTTTTGCCGAAGACGTACCAGGAGTTTCCGGGAGAAGTTATTGGTCTCTCGTTTCTGCCTTTATCAAACTCTACATCTGCCAT
CATTTATAGTTCCAGGGCAATGTGTCTGATCGATTTTGGGATGCCAGTGGAAAGAGAGGAAGATAGTGGTTCACGATTAGTAAGATTAAAAAGCTCTGCA
AACAACAAGGCTAGTCTGAAACGCACAAGGAAAGAGTATGAAGCGAAGATTGAAAATAGGAAGAACTTCGAGTTTGTTCCTTTCAATAATCCAGTTTTAT
TTGTGGGACACCTTTCAGAGAACGGCATATTGATCTTAGACAAGCCATGGTTGAAGACATATATTTGGCACTTAAAAGGAAAAAAGAACGAAAATGGAGA
TATCAGCGAAGGAGGGTGTCTGGAGCAGAGAGTTCAAGAAAAGATGTTTTGGTATTTGCATGGCCTCCTCAGTGCTTGGTGTGGCATGGGATCCTCGATA
CTTCTGCTTGGTCGTGCAGGCAAGGTTTGGTTGGTGATCGACTGGATGAGGAAAAAAGAATGTTGTGGTATCGACTATTTATTGTATCCCTAG
|
|||||||||||||||
|
AA sequence
|
>Lus10005385 pacid=23155609 polypeptide=Lus10005385 locus=Lus10005385.g ID=Lus10005385.BGIv1.0 annot-version=v1.0
MPKNVYRNSSLAWKPSPVVALATSADDSQVAAAREDGSMEIWLVSPGSVGWHCQLTIHGDSSSRVSSLVWCRTGSKGLPCGRLFSSSIDGSVSEWDLFHL
EQKIVLESIGFSIWQMSVAPQCASLIHPEVKSLDLGNGFLNHSHNDDHDDESSESDNDSSSEELHNPVSADTRLAIACDDGCVRLYTIPESDELIYNRTL
SRISHKGDIKLIALVVNTLHHPRMRRVAEAVICRFRNRLKESFDFAIGRGEVVILVRWSALAAAGRSEDRRWQARRRSSAQRGYGRRMTASCKFGVGGRS
EDRLQARRRWEARQWQSLLTVASISIKYLYLGNGFLNTSHNDDHDDESSESDNDSSSEELHNPVSADTRLAIACDDGCVRLYTIPESDELIYNRTLSRVS
GRTLSVTWSSDATRIFSGSSDGFVRCWDASLGHEIYRITVGLGGLGGGTEIRVWSLLALRCGTLVSADSTGTVILYKLSGEIGSSDDTSTSTVMRRWVYV
GYVRAHTHDIRALTMAVPICREDALPEEKVKKDRKRRRHVEFSYHKWAHLGVPMLISASDDTKLFAYSAKEFSKFSPHDICPAPQRMPIHLVPNTISNKN
PLLLVQASSTLDILRVRTEDGNISDSVSGTSKGLAKTVLRAQILSKKKIICSCISNSGMLLAYSDHKKSNLFDLKMHVDPKIPWKVTKREASEEEACKEF
PYAHCMAFSSDSSRLILAGHNRRIYIVDVDSLKIVHTFTPQRHDYNEESPSEPPVTKLFTSNDGQWLAAINCFGDIYVFDMETQRQHWFISRLDGASVTA
GGFPPQNNNVLVVTTSTNQVYAFDVEAKQLGEWSQRNSSFLPKTYQEFPGEVIGLSFLPLSNSTSAIIYSSRAMCLIDFGMPVEREEDSGSRLVRLKSSA
NNKASLKRTRKEYEAKIENRKNFEFVPFNNPVLFVGHLSENGILILDKPWLKTYIWHLKGKKNENGDISEGGCLEQRVQEKMFWYLHGLLSAWCGMGSSI
LLLGRAGKVWLVIDWMRKKECCGIDYLLYP
|
DESeq2's median of ratios [FLAX]
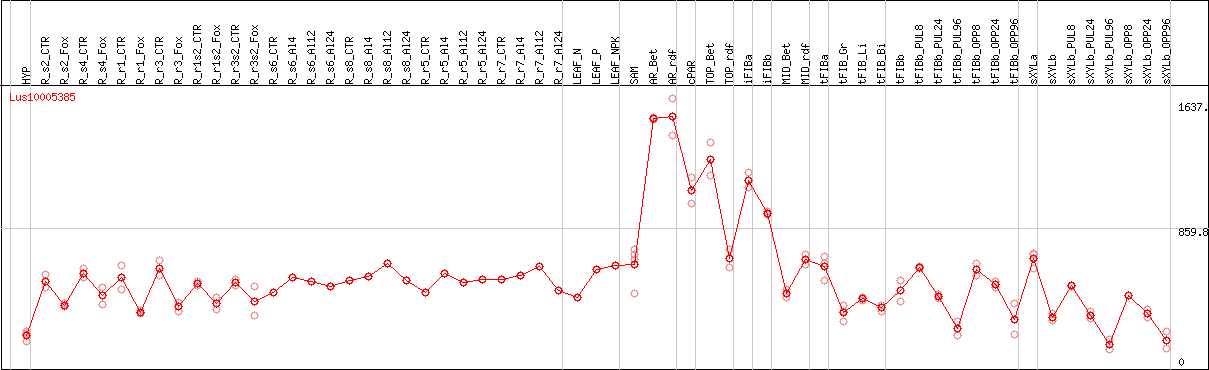
Coexpressed genes
Lus10005385 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
