External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G18380 115 / 9e-32
SHH2
SAWADEE homeodomain homolog 2, sequence-specific DNA binding transcription factors;sequence-specific DNA binding (.1.2.3)
AT1G15215 104 / 2e-28
SHH1, DTF1
SAWADEE homeodomain homolog 1, DNA-binding transcription factor 1, unknown protein
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017944
124 / 2e-35
AT3G18380 418 / 7e-147
SAWADEE homeodomain homolog 2, sequence-specific DNA binding transcription factors;sequence-specific DNA binding (.1.2.3)
Lus10013684
124 / 2e-35
AT3G18380 416 / 4e-146
SAWADEE homeodomain homolog 2, sequence-specific DNA binding transcription factors;sequence-specific DNA binding (.1.2.3)
Lus10025826
115 / 4e-32
AT1G15215 236 / 1e-77
SAWADEE homeodomain homolog 1, DNA-binding transcription factor 1, unknown protein
Lus10038276
114 / 6e-32
AT1G15215 266 / 3e-89
SAWADEE homeodomain homolog 1, DNA-binding transcription factor 1, unknown protein
Lus10014151
104 / 1e-27
AT1G15215 224 / 5e-72
SAWADEE homeodomain homolog 1, DNA-binding transcription factor 1, unknown protein
Lus10037564
68 / 8e-15
AT1G15215 130 / 2e-37
SAWADEE homeodomain homolog 1, DNA-binding transcription factor 1, unknown protein
Lus10006038
41 / 2e-05
ND 40 / 1e-04
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G046200
152 / 3e-47
AT1G15215 195 / 4e-62
SAWADEE homeodomain homolog 1, DNA-binding transcription factor 1, unknown protein
Potri.001G189200
124 / 2e-36
AT1G15215 259 / 3e-87
SAWADEE homeodomain homolog 1, DNA-binding transcription factor 1, unknown protein
Potri.001G373300
120 / 9e-34
AT3G18380 369 / 7e-127
SAWADEE homeodomain homolog 2, sequence-specific DNA binding transcription factors;sequence-specific DNA binding (.1.2.3)
Potri.011G097500
119 / 1e-33
AT3G18380 366 / 1e-126
SAWADEE homeodomain homolog 2, sequence-specific DNA binding transcription factors;sequence-specific DNA binding (.1.2.3)
PFAM info
Representative CDS sequence
>Lus10005480 pacid=23182573 polypeptide=Lus10005480 locus=Lus10005480.g ID=Lus10005480.BGIv1.0 annot-version=v1.0
ATGATGGCTCGGCTCCGAACAAGAGAGAAACTCGTCTTCTCTGGCTTCACCAAGGCGGAGGAAGTTCATGTCAGATTTGAAGGATTTGGTGCGGAGGAAA
ATGAATGGGTAAATGTAAGATATGGAGTGAGACAACGGTCTATCCCATTTGATCATTCAGAGTGTCACAAAGTGAAGGAGGGAGATCTAGTATGCTGCTT
CCAGGAGAGGAAGGATTATGCAATGTATTACGATGCTCATGTGATAGAGATTCAAAGGAAAACACATGACATAAGGGGTTGCAGGTGCATTTTCAAGATA
CTCTATGATCATGATAACATAGAGATGTCAGAAGCCGGGGAGAGCTTTCGGTATGAATGGCCGATCCAGCAAACTGTAGCCTAA
AA sequence
>Lus10005480 pacid=23182573 polypeptide=Lus10005480 locus=Lus10005480.g ID=Lus10005480.BGIv1.0 annot-version=v1.0
MMARLRTREKLVFSGFTKAEEVHVRFEGFGAEENEWVNVRYGVRQRSIPFDHSECHKVKEGDLVCCFQERKDYAMYYDAHVIEIQRKTHDIRGCRCIFKI
LYDHDNIEMSEAGESFRYEWPIQQTVA
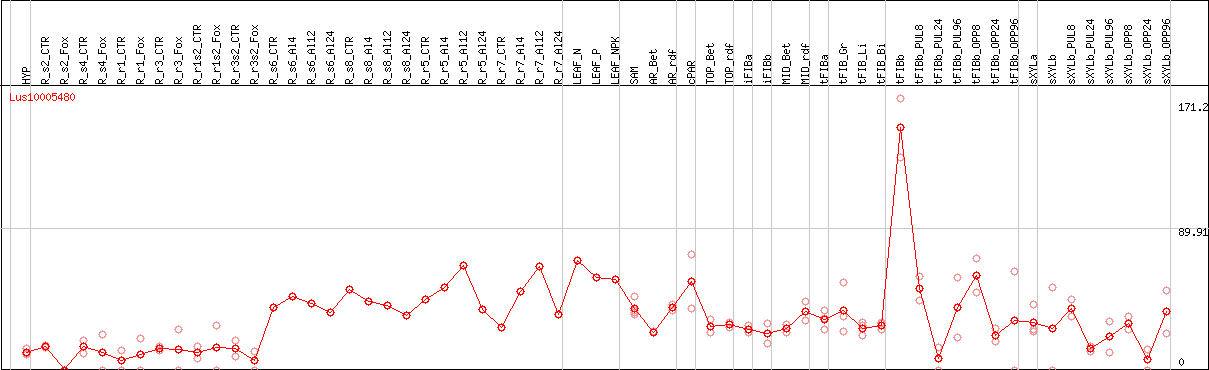
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10005480 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.