External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G13410 157 / 3e-45
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G06150 160 / 1e-44
bHLH
bHLH089, EMB1444
EMBRYO DEFECTIVE 1444, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
AT4G18840 153 / 3e-43
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
AT1G62260 153 / 1e-42
MEF9
mitochondrial editing factor 9 (.1)
AT1G09410 151 / 8e-42
pentatricopeptide (PPR) repeat-containing protein (.1)
AT3G29230 150 / 1e-41
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G44880 149 / 1e-41
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
AT1G74630 147 / 1e-40
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G15930 145 / 8e-40
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT4G02750 145 / 9e-40
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10027455
379 / 2e-130
AT3G29230 324 / 4e-102
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10039974
168 / 1e-48
AT1G06150 612 / 0.0
EMBRYO DEFECTIVE 1444, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Lus10008825
168 / 2e-48
AT1G06150 614 / 0.0
EMBRYO DEFECTIVE 1444, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Lus10003422
164 / 5e-47
AT2G44880 679 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Lus10041097
159 / 5e-45
AT5G66520 367 / 2e-119
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10014113
157 / 3e-44
AT3G29230 767 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10031423
157 / 5e-44
AT5G66520 538 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10040680
157 / 7e-44
AT2G29760 926 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10002288
155 / 1e-43
AT1G31430 689 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G140400
298 / 1e-98
AT3G29230 398 / 2e-132
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.014G046200
174 / 1e-50
AT2G44880 676 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Potri.017G086100
173 / 1e-50
AT5G15300 675 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.008G121400
166 / 7e-48
AT1G13410 561 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.002G258766
165 / 1e-47
AT1G06150 705 / 0.0
EMBRYO DEFECTIVE 1444, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Potri.003G087100
159 / 4e-45
AT3G29230 368 / 1e-119
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.002G072500
158 / 6e-45
AT3G21470 565 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Potri.005G035900
157 / 2e-44
AT1G74630 884 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.006G265200
156 / 2e-44
AT3G29230 364 / 9e-120
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.009G139200
155 / 1e-43
AT1G62260 749 / 0.0
mitochondrial editing factor 9 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10005741 pacid=23169839 polypeptide=Lus10005741 locus=Lus10005741.g ID=Lus10005741.BGIv1.0 annot-version=v1.0
ATGCCTGAAAGACTTGTGGAAGTTTGTAACCGGCTGATAACCGAGTGCGTAAACTCTGGAGATGTCAGGTCTGCTAGAGAGTTGTTTGATGAAATGAAAG
AGAGAGACGTCATTTCCTGGAACTCTCTGATCTCAGGTTATTCAAAAGCTGGAGAAGTAGCCATTGCAAGGGAACTCTTCGACTCTATGCCGGAAAAGAA
TGTCATTTCCTGGACTTCCATGATTGGTGCATACTCTGATTCAGGGGATCTCAAAACAACGATACAGGTGTTCAAAGAAATGCCTCTAAGGAATGTGGTA
TCGTGGAACTCGATGATCTCAAGCTACAACAACCATAACAAGTTCGCAGAAGCGTTGGATCTTTTCCTTCGGATGCAGTCAGAAGGATTCTTGCCTAATG
GTTACACCTTTTTATCTGTTCTACTTGCTTGTTCTAAGCTAGGCAACGTGGAGTTTGGCAAATACGTACATTACCTGATACGAGATTGGTTGCAACTGCA
AGTCATGGTAGGGACTGCCGTAGTCGAAATGTCATTAGCCATTAATGGAAGAGCAGAAGATGCCGTCAAGCTTTTCTGGTCGATGCAGAAGAACACGGAA
TTGAAGCCTAACGATTTCAGTTTCACAAGCATGTTGTTCGTTTGTAGTCACGGAGGTTTGGTCGAAGAAGGCCGTGGGATCTTTAGTAGTATGGACGTGT
ACTTCAATGTTATCGAAGATCGAACATTATGGATGTATGGTTGA
AA sequence
>Lus10005741 pacid=23169839 polypeptide=Lus10005741 locus=Lus10005741.g ID=Lus10005741.BGIv1.0 annot-version=v1.0
MPERLVEVCNRLITECVNSGDVRSARELFDEMKERDVISWNSLISGYSKAGEVAIARELFDSMPEKNVISWTSMIGAYSDSGDLKTTIQVFKEMPLRNVV
SWNSMISSYNNHNKFAEALDLFLRMQSEGFLPNGYTFLSVLLACSKLGNVEFGKYVHYLIRDWLQLQVMVGTAVVEMSLAINGRAEDAVKLFWSMQKNTE
LKPNDFSFTSMLFVCSHGGLVEEGRGIFSSMDVYFNVIEDRTLWMYG
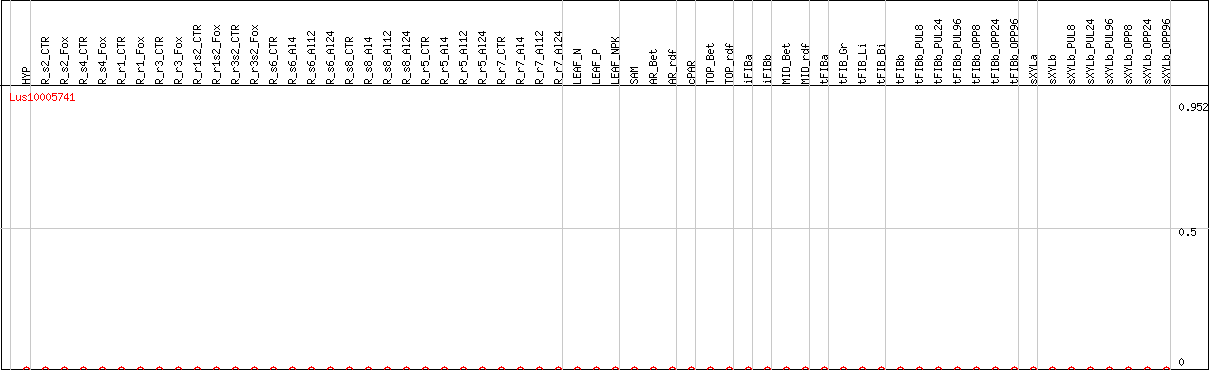
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10005741 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.