External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G23730 226 / 4e-72
EFO2, RUP2
REPRESSOR OF UV-B PHOTOMORPHOGENESIS 2, EARLY FLOWERING BY OVEREXPRESSION 2, Transducin/WD40 repeat-like superfamily protein (.1)
AT5G52250 198 / 4e-61
EFO1, RUP1
REPRESSOR OF UV-B PHOTOMORPHOGENESIS 1, EARLY FLOWERING BY OVEREXPRESSION 1, Transducin/WD40 repeat-like superfamily protein (.1)
AT1G53090 137 / 3e-36
SPA4
SPA1-related 4 (.1.2)
AT3G15354 130 / 1e-33
SPA3
SPA1-related 3 (.1)
AT2G32950 115 / 1e-28
FUS1, EMB168, DET340, ATCOP1, COP1
FUSCA 1, EMBRYO DEFECTIVE 168, DEETIOLATED MUTANT 340, ARABIDOPSIS THALIANA CONSTITUTIVE PHOTOMORPHOGENIC 1, Transducin/WD40 repeat-like superfamily protein (.1)
AT4G11110 102 / 3e-24
SPA2
SPA1-related 2 (.1)
AT2G46340 102 / 4e-24
SPA1
SUPPRESSOR OF PHYA-105 1, SPA (suppressor of phyA-105) protein family (.1)
AT1G15440 50 / 1e-06
ATPWP2
\(PERIODIC TRYPTOPHAN PROTEIN 2, periodic tryptophan protein 2 (.1.2)
AT4G02730 49 / 1e-06
AtWDR5b
human WDR5 \(WD40 repeat\) homolog b, Transducin/WD40 repeat-like superfamily protein (.1)
AT3G21540 48 / 5e-06
transducin family protein / WD-40 repeat family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10027454
371 / 9e-131
AT5G23730 248 / 2e-81
REPRESSOR OF UV-B PHOTOMORPHOGENESIS 2, EARLY FLOWERING BY OVEREXPRESSION 2, Transducin/WD40 repeat-like superfamily protein (.1)
Lus10043323
128 / 7e-33
AT3G15354 906 / 0.0
SPA1-related 3 (.1)
Lus10019475
125 / 4e-32
AT3G15354 992 / 0.0
SPA1-related 3 (.1)
Lus10029990
119 / 7e-30
AT2G32950 1088 / 0.0
FUSCA 1, EMBRYO DEFECTIVE 168, DEETIOLATED MUTANT 340, ARABIDOPSIS THALIANA CONSTITUTIVE PHOTOMORPHOGENIC 1, Transducin/WD40 repeat-like superfamily protein (.1)
Lus10035332
119 / 7e-30
AT2G32950 1088 / 0.0
FUSCA 1, EMBRYO DEFECTIVE 168, DEETIOLATED MUTANT 340, ARABIDOPSIS THALIANA CONSTITUTIVE PHOTOMORPHOGENIC 1, Transducin/WD40 repeat-like superfamily protein (.1)
Lus10032358
96 / 8e-22
AT4G11110 784 / 0.0
SPA1-related 2 (.1)
Lus10033942
88 / 3e-19
AT4G11110 766 / 0.0
SPA1-related 2 (.1)
Lus10012475
49 / 3e-06
AT3G21540 1094 / 0.0
transducin family protein / WD-40 repeat family protein (.1)
Lus10023903
48 / 6e-06
AT3G21540 1223 / 0.0
transducin family protein / WD-40 repeat family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.015G143200
234 / 2e-75
AT5G23730 395 / 2e-137
REPRESSOR OF UV-B PHOTOMORPHOGENESIS 2, EARLY FLOWERING BY OVEREXPRESSION 2, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.012G140200
229 / 8e-73
AT5G23730 427 / 3e-149
REPRESSOR OF UV-B PHOTOMORPHOGENESIS 2, EARLY FLOWERING BY OVEREXPRESSION 2, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.011G119500
131 / 4e-34
AT3G15354 982 / 0.0
SPA1-related 3 (.1)
Potri.001G399600
125 / 6e-32
AT3G15354 1003 / 0.0
SPA1-related 3 (.1)
Potri.014G159300
118 / 9e-30
AT2G32950 1091 / 0.0
FUSCA 1, EMBRYO DEFECTIVE 168, DEETIOLATED MUTANT 340, ARABIDOPSIS THALIANA CONSTITUTIVE PHOTOMORPHOGENIC 1, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.014G095300
112 / 2e-27
AT2G46340 846 / 0.0
SUPPRESSOR OF PHYA-105 1, SPA (suppressor of phyA-105) protein family (.1)
Potri.004G002700
104 / 5e-25
AT2G32950 832 / 0.0
FUSCA 1, EMBRYO DEFECTIVE 168, DEETIOLATED MUTANT 340, ARABIDOPSIS THALIANA CONSTITUTIVE PHOTOMORPHOGENIC 1, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.011G021700
102 / 1e-24
AT2G32950 686 / 0.0
FUSCA 1, EMBRYO DEFECTIVE 168, DEETIOLATED MUTANT 340, ARABIDOPSIS THALIANA CONSTITUTIVE PHOTOMORPHOGENIC 1, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.003G137100
102 / 4e-24
AT4G11110 1016 / 0.0
SPA1-related 2 (.1)
Potri.001G094500
99 / 7e-23
AT4G11110 1016 / 0.0
SPA1-related 2 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0186
Beta_propeller
PF00400
WD40
WD domain, G-beta repeat
Representative CDS sequence
>Lus10005742 pacid=23169844 polypeptide=Lus10005742 locus=Lus10005742.g ID=Lus10005742.BGIv1.0 annot-version=v1.0
ATGAATAACCCCAACCCAGAACCTGAACAAGAGCGAGCCAGATGCGAGTGGGACTTCACCCTTCGAACCACCGTCTCCTCCGCTGCCGCCTCATCCTCCG
ACGCCCTAGGCGTCATCGAATGCGACCCCTCCGGCCGCCTCATCGCCACCGGCGGCAACGCCAGAAANNNNNNNNNNNNNNNNNNCGAGCGGGATGAGCA
CGGCGGCCGACGAGTATGGAGCGTCGACTACGGTTGGTTCGGCGGCGGCGCGACGGCGGTTCTCTGCGCCTCGGGTTCCGACGACGGGACTACCCACGTG
TGGGACCCGCGATGCGAGGTAGGGGAATGCGTGGCTGTAATACGGCCAAGCGGGTCCGGATCTGCTGTTTGCTGCGTGGAGTTCAATCGTCCTCGTGGGA
CGACGCTAGCCGTAGGATGCGCTGACCGGAAGGTGTACGTGTACGACTGCCGTATGCTGGTCCACCCTCTCCTTACCATGGGCGGGCACACCAAAACGGT
AACGTACGTCAAGTTCCTCGACGAAACGACTCTGGTCTCCGCGTCTATAGACGGGAGCTTGAAGCTGTGGGATCTGGCTAACCATTCGGTCGAACCGGTT
CGGACATACCGCGGACATTCCAACGAGGTTAAAGTGGCGTTGGTATATGATCAGCGGTGGGGCGAACCGATTTGGGTGCATGGGTTCGAACCGGCCGAAC
CAAACACCGACGGGTTTGTTAGCAGTGTGTGTTGGAAGCAAGGTGTGGACGACAATGTCGACGGTGGACAATGCACGTTCGTGGCCGGCGGATCGGACGG
TACGTTGCGAGTTTTCGAAGGGATAAAGAAGAAAACGGTGGCGCCGGCTTCTTCCGTCGCCGACGAACACATGAGATAA
AA sequence
>Lus10005742 pacid=23169844 polypeptide=Lus10005742 locus=Lus10005742.g ID=Lus10005742.BGIv1.0 annot-version=v1.0
MNNPNPEPEQERARCEWDFTLRTTVSSAAASSSDALGVIECDPSGRLIATGGNARXXXXXXXERDEHGGRRVWSVDYGWFGGGATAVLCASGSDDGTTHV
WDPRCEVGECVAVIRPSGSGSAVCCVEFNRPRGTTLAVGCADRKVYVYDCRMLVHPLLTMGGHTKTVTYVKFLDETTLVSASIDGSLKLWDLANHSVEPV
RTYRGHSNEVKVALVYDQRWGEPIWVHGFEPAEPNTDGFVSSVCWKQGVDDNVDGGQCTFVAGGSDGTLRVFEGIKKKTVAPASSVADEHMR
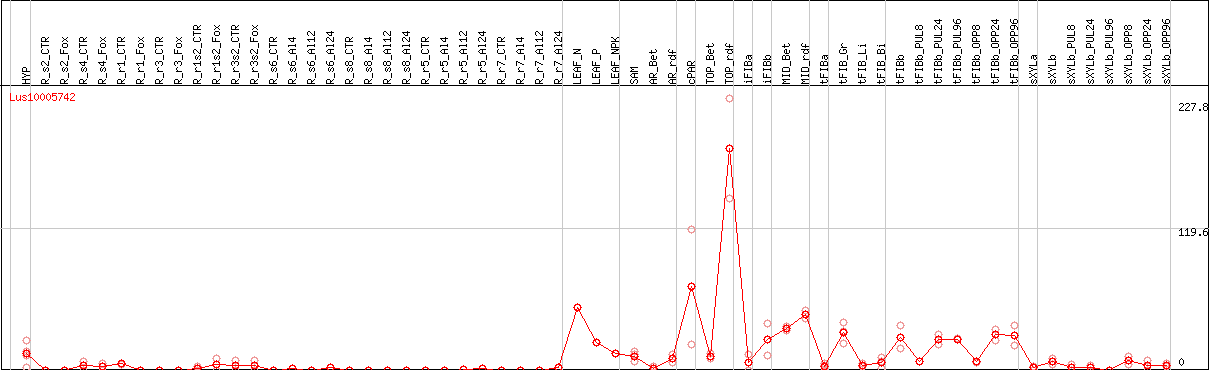
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10005742 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.