External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G48890 270 / 5e-92
MSBP2, ATMP2, ATMAPR3
MEMBRANE STEROID BINDING PROTEIN 2, ARABIDOPSIS THALIANA MEMBRANE-ASSOCIATED PROGESTERONE BINDING PROTEIN 3, membrane-associated progesterone binding protein 3 (.1)
AT5G52240 253 / 2e-85
ATMAPR5, ATMP1, MSBP1
ARABIDOPSIS THALIANA MEMBRANE-ASSOCIATED PROGESTERONE BINDING PROTEIN 5, membrane steroid binding protein 1 (.1.2)
AT2G24940 105 / 5e-29
ATMAPR2
membrane-associated progesterone binding protein 2 (.1)
AT4G14965 78 / 3e-17
ATMAPR4
membrane-associated progesterone binding protein 4 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10027449
461 / 1e-167
AT3G48890 270 / 3e-92
MEMBRANE STEROID BINDING PROTEIN 2, ARABIDOPSIS THALIANA MEMBRANE-ASSOCIATED PROGESTERONE BINDING PROTEIN 3, membrane-associated progesterone binding protein 3 (.1)
Lus10014988
340 / 1e-119
AT3G48890 260 / 2e-88
MEMBRANE STEROID BINDING PROTEIN 2, ARABIDOPSIS THALIANA MEMBRANE-ASSOCIATED PROGESTERONE BINDING PROTEIN 3, membrane-associated progesterone binding protein 3 (.1)
Lus10038868
338 / 4e-119
AT3G48890 265 / 2e-90
MEMBRANE STEROID BINDING PROTEIN 2, ARABIDOPSIS THALIANA MEMBRANE-ASSOCIATED PROGESTERONE BINDING PROTEIN 3, membrane-associated progesterone binding protein 3 (.1)
Lus10038356
191 / 1e-61
AT3G48890 216 / 2e-71
MEMBRANE STEROID BINDING PROTEIN 2, ARABIDOPSIS THALIANA MEMBRANE-ASSOCIATED PROGESTERONE BINDING PROTEIN 3, membrane-associated progesterone binding protein 3 (.1)
Lus10002241
105 / 7e-29
AT2G24940 172 / 1e-57
membrane-associated progesterone binding protein 2 (.1)
Lus10029567
110 / 5e-28
AT5G47740 187 / 4e-57
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Lus10018026
74 / 2e-15
AT4G14965 361 / 3e-127
membrane-associated progesterone binding protein 4 (.1)
Lus10034799
69 / 3e-15
AT2G24940 87 / 3e-24
membrane-associated progesterone binding protein 2 (.1)
Lus10042022
63 / 2e-11
AT4G14965 357 / 5e-126
membrane-associated progesterone binding protein 4 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G137800
317 / 5e-111
AT3G48890 259 / 3e-88
MEMBRANE STEROID BINDING PROTEIN 2, ARABIDOPSIS THALIANA MEMBRANE-ASSOCIATED PROGESTERONE BINDING PROTEIN 3, membrane-associated progesterone binding protein 3 (.1)
Potri.015G139800
312 / 7e-109
AT3G48890 277 / 3e-95
MEMBRANE STEROID BINDING PROTEIN 2, ARABIDOPSIS THALIANA MEMBRANE-ASSOCIATED PROGESTERONE BINDING PROTEIN 3, membrane-associated progesterone binding protein 3 (.1)
Potri.015G125100
184 / 2e-58
AT3G48890 201 / 1e-65
MEMBRANE STEROID BINDING PROTEIN 2, ARABIDOPSIS THALIANA MEMBRANE-ASSOCIATED PROGESTERONE BINDING PROTEIN 3, membrane-associated progesterone binding protein 3 (.1)
Potri.018G016900
108 / 3e-30
AT2G24940 171 / 6e-57
membrane-associated progesterone binding protein 2 (.1)
Potri.006G266200
107 / 7e-30
AT2G24940 160 / 1e-52
membrane-associated progesterone binding protein 2 (.1)
Potri.016G022700
73 / 3e-15
AT4G14965 345 / 3e-121
membrane-associated progesterone binding protein 4 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00173
Cyt-b5
Cytochrome b5-like Heme/Steroid binding domain
Representative CDS sequence
>Lus10005746 pacid=23169816 polypeptide=Lus10005746 locus=Lus10005746.g ID=Lus10005746.BGIv1.0 annot-version=v1.0
ATGGCGCTTCAGCTATGGGAGACGCTGCAGCAAGCTATAACGGCGTACACGGGGCTAACGCCGGCGACTTTCTTCACGGTCCTTGCTTTTGGGTTGGCCG
TTTACTATCTGCTTTCTGGGATGTTCGGGTCAACTAATCCTCCGCCTAGACCCAGGAGCTTCGAGGAAGACGTGCAGCCGCTGCCGCCGCCTGTGCAGCT
TGGGGAGATCGGCGAGGATGAGCTGAAGCAGTACGATGGGACTGATCCTAAGAAGCCTCTGCTTATGGCTATCAAGGGTCAGATCTACGATGTCTCCCAG
AGCAGGATGTTTTACGGTCCCGGTGGGCCATACGCGTTATTTGCCGGGAAGGATGCCAGCAGAGCCCTGGCGAAGATGTCGTTCGAAGAGAAGGATCTGA
CAGGAGACATCTCAGGACTCGGTTCGTTTGAGATGGAAGCTCTGCAGGACTGGGAATACAAGTTCATGAGTAAGTATGTCAAGGTCGGTACCATCAAGAA
GGAAGTACCAGTCGTCGATGATGGTTCATCCAGCGAGCAGCCTGCTGCCGAAGGCACTGCTGATGCTACTGAGAACGGCGATGTGAAGCCTGCAAGCACT
GAAGCTGACGACAACAAGGCAGCAGCTGAAGACGGTCCATCCTCGACAGGTGTTGAGCATCCGGTAGCTCCTCCATCAGGTGGTGAATCAAAAGAAGAGT
GA
AA sequence
>Lus10005746 pacid=23169816 polypeptide=Lus10005746 locus=Lus10005746.g ID=Lus10005746.BGIv1.0 annot-version=v1.0
MALQLWETLQQAITAYTGLTPATFFTVLAFGLAVYYLLSGMFGSTNPPPRPRSFEEDVQPLPPPVQLGEIGEDELKQYDGTDPKKPLLMAIKGQIYDVSQ
SRMFYGPGGPYALFAGKDASRALAKMSFEEKDLTGDISGLGSFEMEALQDWEYKFMSKYVKVGTIKKEVPVVDDGSSSEQPAAEGTADATENGDVKPAST
EADDNKAAAEDGPSSTGVEHPVAPPSGGESKEE
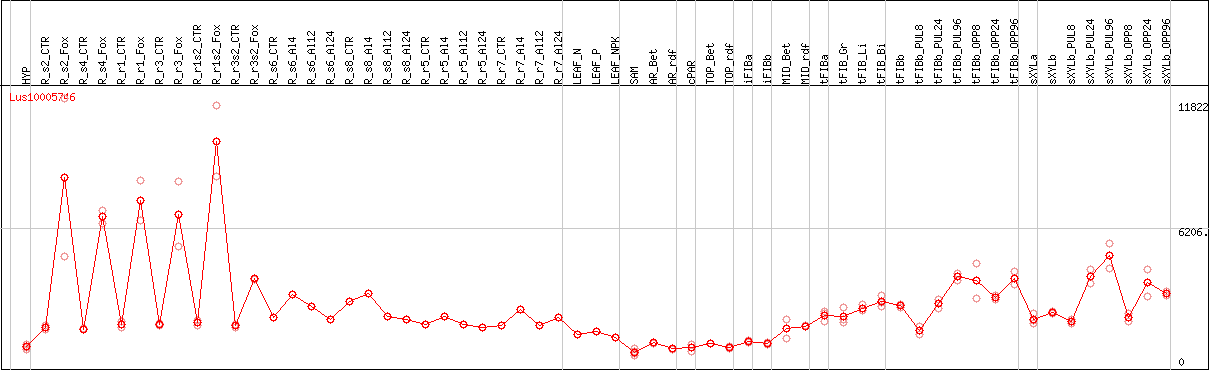
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10005746 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.