External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G00730 155 / 3e-44
HD
AHDP, ANL2
ANTHOCYANINLESS 2, ARABIDOPSIS THALIANA HOMEODOMAIN PROTEIN, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
AT3G61150 155 / 1e-43
HD
HD-GL2-1, HDG1
HOMEODOMAIN-GLABRA2 1, homeodomain GLABROUS 1 (.1)
AT4G04890 139 / 4e-38
HD
PDF2
protodermal factor 2 (.1)
AT4G21750 134 / 2e-36
HD
ATML1
MERISTEM LAYER 1, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
AT5G52170 134 / 2e-36
HD
HDG7
homeodomain GLABROUS 7 (.1)
AT1G73360 131 / 3e-35
HD
ATHDG11, HDG11, EDT1
ENHANCED DROUGHT TOLERANCE 1, homeodomain GLABROUS 11 (.1)
AT1G05230 128 / 5e-34
HD
HDG2
homeodomain GLABROUS 2 (.1.2.3.4)
AT1G17920 119 / 8e-31
HD
HDG12
homeodomain GLABROUS 12 (.1)
AT5G46880 116 / 7e-30
HD
HDG5, HB-7
HOMEODOMAIN GLABROUS 5, homeobox-7 (.1)
AT2G32370 101 / 9e-25
HD
HDG3
homeodomain GLABROUS 3 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10027437
215 / 2e-65
AT3G61150 715 / 0.0
HOMEODOMAIN-GLABRA2 1, homeodomain GLABROUS 1 (.1)
Lus10005759
206 / 4e-62
AT3G61150 706 / 0.0
HOMEODOMAIN-GLABRA2 1, homeodomain GLABROUS 1 (.1)
Lus10036567
164 / 9e-47
AT4G00730 979 / 0.0
ANTHOCYANINLESS 2, ARABIDOPSIS THALIANA HOMEODOMAIN PROTEIN, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
Lus10041362
147 / 2e-41
AT4G00730 379 / 2e-122
ANTHOCYANINLESS 2, ARABIDOPSIS THALIANA HOMEODOMAIN PROTEIN, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
Lus10018360
140 / 2e-41
AT1G05230 299 / 4e-97
homeodomain GLABROUS 2 (.1.2.3.4)
Lus10038877
139 / 9e-40
AT4G00730 261 / 6e-83
ANTHOCYANINLESS 2, ARABIDOPSIS THALIANA HOMEODOMAIN PROTEIN, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
Lus10007650
141 / 1e-38
AT4G04890 1121 / 0.0
protodermal factor 2 (.1)
Lus10027175
141 / 1e-38
AT1G05230 1138 / 0.0
homeodomain GLABROUS 2 (.1.2.3.4)
Lus10039667
140 / 4e-38
AT1G05230 1142 / 0.0
homeodomain GLABROUS 2 (.1.2.3.4)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.015G141800
174 / 1e-50
AT4G00730 829 / 0.0
ANTHOCYANINLESS 2, ARABIDOPSIS THALIANA HOMEODOMAIN PROTEIN, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
Potri.012G139300
167 / 6e-48
AT4G00730 838 / 0.0
ANTHOCYANINLESS 2, ARABIDOPSIS THALIANA HOMEODOMAIN PROTEIN, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
Potri.014G075200
166 / 2e-47
AT4G00730 1045 / 0.0
ANTHOCYANINLESS 2, ARABIDOPSIS THALIANA HOMEODOMAIN PROTEIN, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
Potri.002G154700
161 / 1e-45
AT4G00730 1057 / 0.0
ANTHOCYANINLESS 2, ARABIDOPSIS THALIANA HOMEODOMAIN PROTEIN, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
Potri.004G020400
152 / 9e-43
AT4G04890 1185 / 0.0
protodermal factor 2 (.1)
Potri.014G152000
144 / 7e-40
AT1G05230 1179 / 0.0
homeodomain GLABROUS 2 (.1.2.3.4)
Potri.002G230200
144 / 1e-39
AT1G05230 1159 / 0.0
homeodomain GLABROUS 2 (.1.2.3.4)
Potri.011G025000
142 / 5e-39
AT4G21750 1164 / 0.0
MERISTEM LAYER 1, Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein (.1.2)
Potri.015G034100
138 / 1e-37
AT1G73360 927 / 0.0
ENHANCED DROUGHT TOLERANCE 1, homeodomain GLABROUS 11 (.1)
Potri.012G038500
130 / 8e-35
AT1G73360 920 / 0.0
ENHANCED DROUGHT TOLERANCE 1, homeodomain GLABROUS 11 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0123
HTH
PF00046
Homeodomain
Homeodomain
Representative CDS sequence
>Lus10005757 pacid=23169814 polypeptide=Lus10005757 locus=Lus10005757.g ID=Lus10005757.BGIv1.0 annot-version=v1.0
ATGGAGGGCCAATCTTTTGCTGAAGAACAACAACATCCGAGAAAGAAAAATTACAAGCGCCATTCTCCCTCCCAAATCCTTGAGCTTGAGAAAGCTTTCA
AAGAGTGCTCTCGTCCAGATAAAGAGCAAAGGCTAAAGCTTGGATTGGACACCAGTCAGGTTAAGTTTTGGTTCCAGAACCACCGCACCCAATTAAAGAC
TCAAATGGAGAGGTATGAGACAGTGCTAATGAGTGAGGCCATTGAAAAGATCAAAGTTGAAAATGTTATGTTGAGCCAGTATTTGCAGAACCCACTCTGC
CACAACTGTTGTGGTCCCGAGGTATTCGAACTCTTCTCGTTTGGGAAACAGCAGCTGCGAATCGAGAATGCTCGTCTCAAGGAAGAAATCGTTCGTGTTT
GTTCCTTGGTCAAGAAGTACCTTGGCCGGCCTCTTTTGTCCTTTGTTCTTGCTTCCCCATATGCTCCTTCCAATTATGGTAAATATGGACCAACTACTTC
AAATGTTGATATTGGGATTGACTTCTCTCATCCTTCCATTAAAGATGTTAACCGAAAAATGGTGGTACCGCCAACAAGTGGTGAAGAAGTGTCGAGTTTC
GACAAGTCCATGTATGTTGATTTCTATGAATGA
AA sequence
>Lus10005757 pacid=23169814 polypeptide=Lus10005757 locus=Lus10005757.g ID=Lus10005757.BGIv1.0 annot-version=v1.0
MEGQSFAEEQQHPRKKNYKRHSPSQILELEKAFKECSRPDKEQRLKLGLDTSQVKFWFQNHRTQLKTQMERYETVLMSEAIEKIKVENVMLSQYLQNPLC
HNCCGPEVFELFSFGKQQLRIENARLKEEIVRVCSLVKKYLGRPLLSFVLASPYAPSNYGKYGPTTSNVDIGIDFSHPSIKDVNRKMVVPPTSGEEVSSF
DKSMYVDFYE
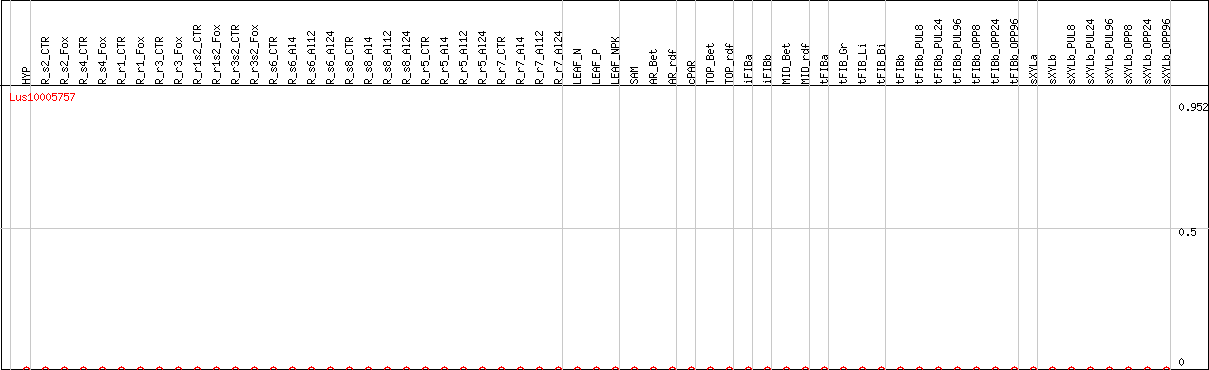
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10005757 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.