External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G52790 144 / 2e-41
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G52800 140 / 7e-40
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G03070 128 / 3e-35
AOP1.1, AOP1
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G80320 126 / 1e-34
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G52820 118 / 1e-31
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G15540 117 / 4e-31
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G28030 103 / 4e-26
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G52810 86 / 1e-19
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT2G36690 61 / 1e-10
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G03050 55 / 2e-08
AOP3
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005773
457 / 4e-164
AT1G52790 205 / 3e-64
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10005776
375 / 7e-132
AT1G52790 198 / 2e-61
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10021025
241 / 3e-79
AT1G52800 226 / 2e-72
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10027419
225 / 2e-75
AT1G52800 89 / 4e-22
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10005775
159 / 5e-50
AT1G52800 70 / 1e-15
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10023024
117 / 3e-31
AT1G52820 371 / 2e-129
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10016659
116 / 8e-31
AT1G52820 445 / 1e-158
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10013132
96 / 6e-23
AT1G52800 222 / 8e-71
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10005516
95 / 2e-22
AT1G52820 278 / 7e-92
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G176500
164 / 3e-49
AT1G52800 338 / 1e-116
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G176000
163 / 1e-48
AT1G52790 460 / 2e-164
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G176200
135 / 6e-38
AT1G52820 496 / 8e-179
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G176100
132 / 5e-37
AT1G52800 286 / 5e-96
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.018G033400
121 / 1e-32
AT1G52820 269 / 4e-89
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G175900
117 / 5e-31
AT1G52820 328 / 2e-112
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.006G248000
117 / 6e-31
AT1G52820 280 / 6e-93
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G175800
112 / 4e-29
AT1G52820 338 / 2e-116
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.017G049000
65 / 6e-12
AT2G36690 209 / 5e-65
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.017G048900
59 / 7e-10
AT2G36690 250 / 5e-80
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0029
Cupin
PF03171
2OG-FeII_Oxy
2OG-Fe(II) oxygenase superfamily
Representative CDS sequence
>Lus10005774 pacid=23169827 polypeptide=Lus10005774 locus=Lus10005774.g ID=Lus10005774.BGIv1.0 annot-version=v1.0
ATGGCAGCTCCCAGCAGTATTCCTGTTATATATTTCTCGGGAGTGAAGCCGGGAACCAATTCATGGGTTTCTGCTCGGACAACCGTGAGAAGGGCGATGG
AAGAGGTCGGGTGTTTCAAGGCCATATACACAGGTGCCGGCGAGATTTCGATTTTCGAGACGGTAAAAGAGCTGTTCGATCTTCCAGATGAGATAAAAAA
CAAGAACACCAGCACTAAGCCCTTTTTCGGAGGTTCCGTGCTGAGTCCTCTCAACAGTCATACTCACGAATCACTAGGAATCGATAACCCGACAAGCCTC
GATACCACTCAAGGCTTCACCAATCTTATGTGGCCGGATTCTGGAAATGATGCCTTCTGCCAGTCGAGTCTTTCCATGTCGAAGCTAATGGTGGAACTAT
TCGACATGATATTGAAGATGGTGATGGAAAGCTACGAGGTTGTCGAATTAGGGTTTGGACCTGAGTCGATGAATCACCTAATTCGATACTCTAAGTTCCC
ATTGCCGTCGTCTAATCAAACCTCCGGTACTGATCTGAGCGTCAATCCTCATAAAGATTTTACCTTTCTGTCCATACTTCATCAGAACCAAGTCAACGGC
TTGCAAATCCAACCACCGATCGATCCACAACGTTGGGTTGACGTTGACCTTACTACGCCGTCGTCCTTCTTGGTTTTCGCGGGGGATGCACTCATGGGTA
GTCACATACATGCTTATTTTGATTACCCAAAGTTTAAAACTAGTTCTATTTTGTAA
AA sequence
>Lus10005774 pacid=23169827 polypeptide=Lus10005774 locus=Lus10005774.g ID=Lus10005774.BGIv1.0 annot-version=v1.0
MAAPSSIPVIYFSGVKPGTNSWVSARTTVRRAMEEVGCFKAIYTGAGEISIFETVKELFDLPDEIKNKNTSTKPFFGGSVLSPLNSHTHESLGIDNPTSL
DTTQGFTNLMWPDSGNDAFCQSSLSMSKLMVELFDMILKMVMESYEVVELGFGPESMNHLIRYSKFPLPSSNQTSGTDLSVNPHKDFTFLSILHQNQVNG
LQIQPPIDPQRWVDVDLTTPSSFLVFAGDALMGSHIHAYFDYPKFKTSSIL
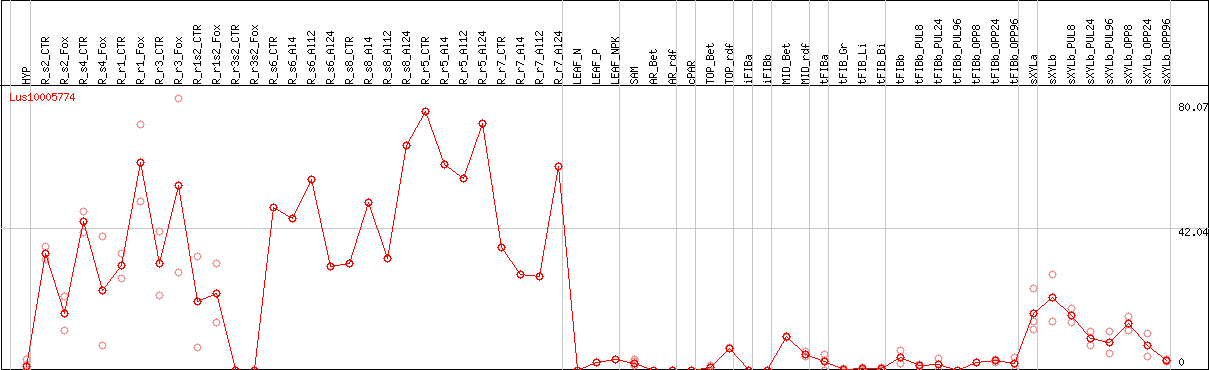
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10005774 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.