External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G07740 258 / 1e-84
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G09900 119 / 4e-31
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
AT1G05670 118 / 1e-30
Pentatricopeptide repeat (PPR-like) superfamily protein (.1), Pentatricopeptide repeat (PPR-like) superfamily protein (.2)
AT1G63070 118 / 1e-30
pentatricopeptide (PPR) repeat-containing protein (.1)
AT5G39710 117 / 3e-30
EMB2745
EMBRYO DEFECTIVE 2745, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G64320 116 / 6e-30
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G62680 115 / 1e-29
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT4G01400 114 / 1e-29
unknown protein
AT1G12775 114 / 4e-29
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G12300 113 / 8e-29
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10040869
386 / 1e-134
AT1G07740 491 / 5e-172
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10013283
121 / 1e-31
AT1G05670 785 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1), Pentatricopeptide repeat (PPR-like) superfamily protein (.2)
Lus10039056
120 / 3e-31
AT5G64320 832 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10027916
116 / 8e-30
AT5G39710 1038 / 0.0
EMBRYO DEFECTIVE 2745, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10021074
112 / 2e-29
AT1G62930 273 / 5e-86
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10012068
115 / 3e-29
AT5G39710 1028 / 0.0
EMBRYO DEFECTIVE 2745, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10027914
113 / 9e-29
AT5G39710 1025 / 0.0
EMBRYO DEFECTIVE 2745, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10005303
113 / 1e-28
AT1G30290 1037 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10002675
112 / 2e-28
AT4G01400 570 / 0.0
unknown protein
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G204100
295 / 3e-99
AT1G07740 502 / 2e-176
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.001G020501
168 / 3e-53
AT1G07740 218 / 4e-70
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G014500
134 / 3e-36
AT1G09820 696 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Potri.013G034200
122 / 3e-32
AT1G12700 511 / 6e-174
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.013G032600
119 / 5e-31
AT1G12700 493 / 2e-166
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.005G046200
117 / 3e-30
AT1G12700 501 / 5e-170
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.001G075900
117 / 4e-30
AT4G20090 806 / 0.0
embryo defective 1025, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.005G050400
115 / 7e-30
AT1G12700 504 / 3e-171
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.014G090400
115 / 1e-29
AT5G28460 637 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.002G109500
114 / 3e-29
AT1G09900 855 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10005876 pacid=23176947 polypeptide=Lus10005876 locus=Lus10005876.g ID=Lus10005876.BGIv1.0 annot-version=v1.0
ATGATTTCGAAAGGGAAGCGACCGAACGAGGTGACTTATGCGTTGTTAATGGAAGGATTATGCTCCATGGGAGAGCATAAGGAGGCTAAGAAGTTGATGT
TCGACATGGAATATCGAGGGTGCAAGCCGAAAGTTTTGAACTTTGGGATCTTGATGAGCGATCTAGGGAAGCGAGGCAAGATCGAGGAAGCGAAGGCTTT
GCTTGTCGAAATGAAGAAGAAGAGGAAGTTGAAGCCGGATGTTGTAACTTATAATATACTGGTGAATTGTTTGTGTAAAGAAGGGAGGGTTTCGGAGGCG
TATAAGGTGTTGCTGGAGATGGAGATGGGAGGGTGCGAGGCGAATGCAGTGACGTATAGGATGCTGGTTGATGGATTTTGCAGGGAAGGGGATTTCGAAG
GTGGGTTGAGAGTGTTGACTGCGATGTTGAGTAGTAAGCATTTCCCGAGAACTGAGACGTTTGTGTGTATGGTTGGGGGATTGCTTGAATCTGGGAATGT
TGAAGGAGCTGGGTTTGTGTTGGAGGAGATGCGTAAGAGGAAATTGGCATTGGGTTGGGATGGCTGGGAGGATCTGGTGAAGGGTTCGTCTGACAGTGAC
GAAAAATTGAGGGAACTTGTGAATCGAGTGATTGAGGGATACAAATGA
AA sequence
>Lus10005876 pacid=23176947 polypeptide=Lus10005876 locus=Lus10005876.g ID=Lus10005876.BGIv1.0 annot-version=v1.0
MISKGKRPNEVTYALLMEGLCSMGEHKEAKKLMFDMEYRGCKPKVLNFGILMSDLGKRGKIEEAKALLVEMKKKRKLKPDVVTYNILVNCLCKEGRVSEA
YKVLLEMEMGGCEANAVTYRMLVDGFCREGDFEGGLRVLTAMLSSKHFPRTETFVCMVGGLLESGNVEGAGFVLEEMRKRKLALGWDGWEDLVKGSSDSD
EKLRELVNRVIEGYK
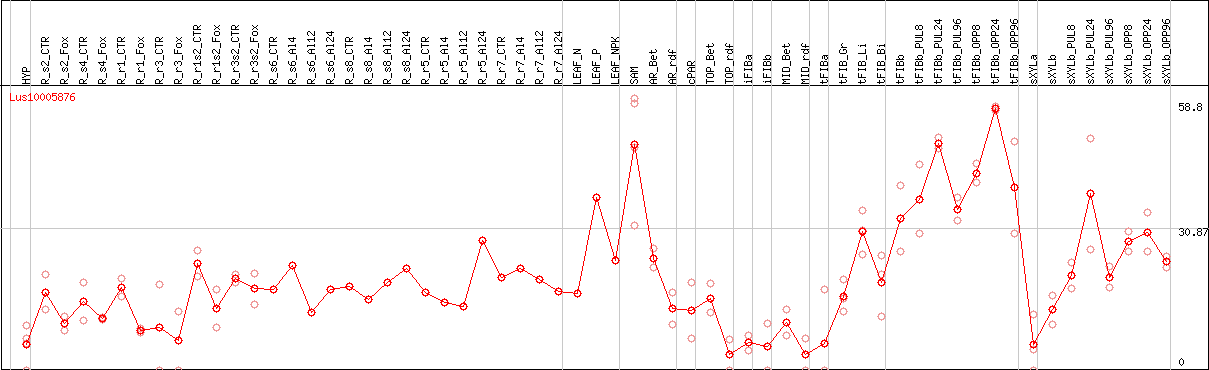
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10005876 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.