External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G46440 513 / 0
UXS5
UDP-XYL synthase 5 (.1.2)
AT2G28760 508 / 0
UXS6
UDP-XYL synthase 6 (.1.2.3)
AT5G59290 506 / 0
ATUXS3, UXS3
UDP-glucuronic acid decarboxylase 3 (.1.2)
AT2G47650 363 / 7e-125
UXS4
UDP-xylose synthase 4 (.1.2)
AT3G62830 362 / 3e-124
ATUXS2, UXS2, AUD1
UDP-GLUCURONIC ACID DECARBOXYLASE 2, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT3G53520 353 / 5e-121
ATUXS1, UXS1
UDP-glucuronic acid decarboxylase 1 (.1.2.3.4)
AT2G28755 86 / 2e-21
UDP-D-glucuronate carboxy-lyase-related (.1)
AT5G28840 88 / 2e-19
GME
"GDP-D-mannose 3',5'-epimerase", GDP-D-mannose 3',5'-epimerase (.1.2)
AT1G53500 88 / 2e-19
ATRHM2, ATMUM4, RHM2, MUM4
ARABIDOPSIS THALIANA RHAMNOSE BIOSYNTHESIS 2, ARABIDOPSIS THALIANA MUCILAGE-MODIFIED 4, NAD-dependent epimerase/dehydratase family protein (.1)
AT3G14790 87 / 7e-19
ATRHM3, RHM3
ARABIDOPSIS THALIANA RHAMNOSE BIOSYNTHESIS 3, rhamnose biosynthesis 3 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10040847
565 / 0
AT3G46440 635 / 0.0
UDP-XYL synthase 5 (.1.2)
Lus10005450
526 / 0
AT3G46440 635 / 0.0
UDP-XYL synthase 5 (.1.2)
Lus10005155
508 / 0
AT2G28760 639 / 0.0
UDP-XYL synthase 6 (.1.2.3)
Lus10001707
508 / 0
AT2G28760 636 / 0.0
UDP-XYL synthase 6 (.1.2.3)
Lus10001705
378 / 5e-133
AT2G28760 501 / 0.0
UDP-XYL synthase 6 (.1.2.3)
Lus10003605
372 / 5e-128
AT2G47650 693 / 0.0
UDP-xylose synthase 4 (.1.2)
Lus10030368
369 / 1e-126
AT3G62830 723 / 0.0
UDP-GLUCURONIC ACID DECARBOXYLASE 2, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10006510
363 / 7e-125
AT3G62830 637 / 0.0
UDP-GLUCURONIC ACID DECARBOXYLASE 2, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Lus10024436
362 / 4e-124
AT3G53520 736 / 0.0
UDP-glucuronic acid decarboxylase 1 (.1.2.3.4)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G207200
521 / 0
AT2G28760 625 / 0.0
UDP-XYL synthase 6 (.1.2.3)
Potri.008G053100
520 / 0
AT2G28760 623 / 0.0
UDP-XYL synthase 6 (.1.2.3)
Potri.001G237200
519 / 0
AT3G46440 637 / 0.0
UDP-XYL synthase 5 (.1.2)
Potri.002G204400
363 / 7e-125
AT3G62830 723 / 0.0
UDP-GLUCURONIC ACID DECARBOXYLASE 2, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.014G129200
363 / 1e-124
AT3G62830 720 / 0.0
UDP-GLUCURONIC ACID DECARBOXYLASE 2, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.016G080500
359 / 3e-123
AT3G53520 736 / 0.0
UDP-glucuronic acid decarboxylase 1 (.1.2.3.4)
Potri.006G214000
356 / 4e-122
AT3G53520 709 / 0.0
UDP-glucuronic acid decarboxylase 1 (.1.2.3.4)
Potri.005G053000
93 / 2e-21
AT5G28840 720 / 0.0
"GDP-D-mannose 3',5'-epimerase", GDP-D-mannose 3',5'-epimerase (.1.2)
Potri.013G040600
93 / 3e-21
AT5G28840 723 / 0.0
"GDP-D-mannose 3',5'-epimerase", GDP-D-mannose 3',5'-epimerase (.1.2)
Potri.006G272700
92 / 1e-20
AT1G78570 1078 / 0.0
REPRESSOR OF LRX1 1, ARABIDOPSIS THALIANA RHAMNOSE BIOSYNTHESIS 1, rhamnose biosynthesis 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF01370
Epimerase
NAD dependent epimerase/dehydratase family
Representative CDS sequence
>Lus10005900 pacid=23176921 polypeptide=Lus10005900 locus=Lus10005900.g ID=Lus10005900.BGIv1.0 annot-version=v1.0
ATGGCGACCGATTCAGCTAACGGGCAGCACCAATCAACCACCAAGCCGCCACCACAGCCGTCCCCATTGCGTTCTTCCAAGTTTTGCCAGTCGAATATGA
GGATTTTGGTCACTGGAGGAGCCGGATTCATTGGTTCCCACTTGGTGGATCGGCTGATGGAAAATGAGAAGAATGAGGTAATTGTTGCTGATAACTATTT
CACCGGATCGAAAGAGAACCTCAGGAAGTGGATTGGCCACCCAAGATTCGAGCTCATCCGTCATGATGTGACAGAGCCATTGCTGGTTGAGGTGGACCAG
ATTTACCATCTTGCATGTCCTGCTTCTCCCATTTTCTACAAGTACAATCCTGTAAAGAATGTGATAGGGGTTCGAAGCTGCTACGATGAGGGGAAACGTG
TGGCTGAGACATTGATGTTTGATTATCACAGGCAGCATGGCATTGAAATCCGAATCGCGAGGATCTTCAACACATATGGACCGCGCATGAACATTGACGA
TGGGCGTGTTGTCAGCAACTTCATAGCTCAAGCACTTCGCGGTGAGCCATTGACTGTACAGAAGCCTGGAACTCAAACCCGTAGCTTCTGCTATGTATCT
GACATGGTTGATGGGCTGATCAGGCTGATGGGGGGAGAGAACACTGGACCGATCAACATCGGGAACCCAGGGGAGTTCACCATGACTGAACTTGCCGAGA
CAGTGAAGGAGCTGATAAACCCGGAGGTGGAGATAAAGAGTGTGGAGAACACACCGGATGATCCGAGGCAGAGGAAGCCTGACATCACAAAGGCCAAGGA
ATTGCTGGGTTGGGAGCCTAAGGTCAAGTTGAAGGACGGACTTCCCCTCATGGAAGAAGATTTCCGTAGGAGGCTTGGAGTGTCCAAGAAGTGA
AA sequence
>Lus10005900 pacid=23176921 polypeptide=Lus10005900 locus=Lus10005900.g ID=Lus10005900.BGIv1.0 annot-version=v1.0
MATDSANGQHQSTTKPPPQPSPLRSSKFCQSNMRILVTGGAGFIGSHLVDRLMENEKNEVIVADNYFTGSKENLRKWIGHPRFELIRHDVTEPLLVEVDQ
IYHLACPASPIFYKYNPVKNVIGVRSCYDEGKRVAETLMFDYHRQHGIEIRIARIFNTYGPRMNIDDGRVVSNFIAQALRGEPLTVQKPGTQTRSFCYVS
DMVDGLIRLMGGENTGPINIGNPGEFTMTELAETVKELINPEVEIKSVENTPDDPRQRKPDITKAKELLGWEPKVKLKDGLPLMEEDFRRRLGVSKK
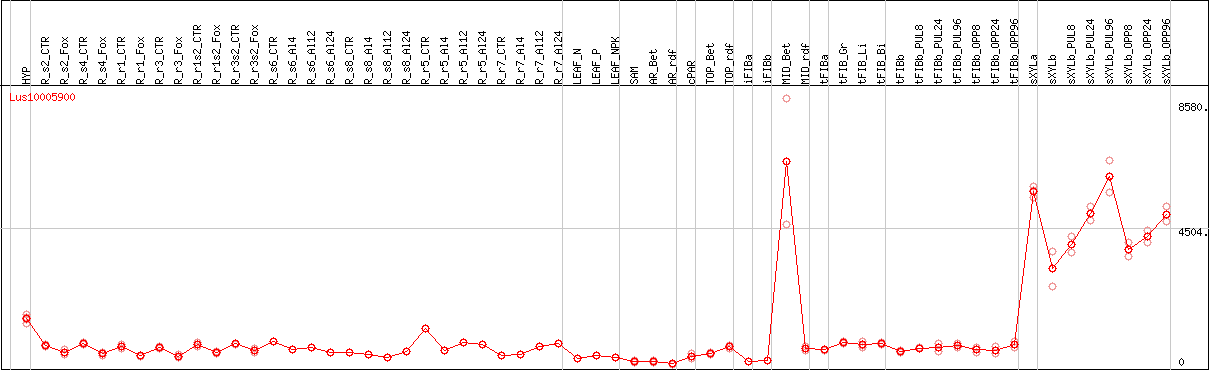
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10005900 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.