External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G13180 91 / 3e-24
NAC
VNDIP2, ANAC083, VNI2
VND-interacting 2, NAC domain containing protein 83 (.1)
AT2G33480 88 / 4e-23
NAC
ANAC041
NAC domain containing protein 41 (.1.2)
AT1G61110 67 / 3e-15
NAC
ANAC025
NAC domain containing protein 25 (.1)
AT3G04070 65 / 3e-14
NAC
ANAC047
NAC domain containing protein 47 (.1.2)
AT1G69490 62 / 2e-13
NAC
NAP, ANAC029, ATNAP
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
AT3G15510 61 / 9e-13
NAC
ATNAC2, ANAC056, NARS1
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
AT1G77450 58 / 4e-12
NAC
ANAC032
NAC domain containing protein 32 (.1)
AT1G52880 57 / 2e-11
NAC
ATNAM, NAM, ANAC018, NARS2
NAC-REGULATED SEED MORPHOLOGY 2, Arabidopsis NAC domain containing protein 18, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT1G65910 56 / 3e-11
NAC
ANAC028
NAC domain containing protein 28 (.1)
AT1G54330 56 / 4e-11
NAC
ANAC020
NAC domain containing protein 20 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002581
100 / 3e-28
AT5G13180 273 / 6e-93
VND-interacting 2, NAC domain containing protein 83 (.1)
Lus10001809
100 / 5e-28
AT5G13180 271 / 6e-92
VND-interacting 2, NAC domain containing protein 83 (.1)
Lus10026496
66 / 2e-14
AT1G61110 244 / 6e-79
NAC domain containing protein 25 (.1)
Lus10032004
65 / 2e-14
AT5G14000 113 / 2e-30
NAC domain containing protein 84 (.1)
Lus10036773
65 / 2e-14
AT1G69490 318 / 1e-109
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
Lus10037156
65 / 2e-14
AT1G69490 318 / 2e-109
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
Lus10011215
65 / 3e-14
AT1G61110 305 / 1e-102
NAC domain containing protein 25 (.1)
Lus10023179
64 / 4e-14
AT1G61110 248 / 2e-80
NAC domain containing protein 25 (.1)
Lus10043095
64 / 4e-14
AT3G15510 351 / 1e-119
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G061200
94 / 2e-25
AT5G13180 310 / 6e-107
VND-interacting 2, NAC domain containing protein 83 (.1)
Potri.003G166500
91 / 3e-24
AT5G13180 310 / 4e-107
VND-interacting 2, NAC domain containing protein 83 (.1)
Potri.015G102100
86 / 2e-22
AT5G13180 232 / 8e-77
VND-interacting 2, NAC domain containing protein 83 (.1)
Potri.012G103500
81 / 9e-21
AT5G13180 229 / 2e-75
VND-interacting 2, NAC domain containing protein 83 (.1)
Potri.001G325100
75 / 2e-18
AT5G13180 163 / 2e-49
VND-interacting 2, NAC domain containing protein 83 (.1)
Potri.017G063300
72 / 2e-17
AT5G13180 150 / 9e-45
VND-interacting 2, NAC domain containing protein 83 (.1)
Potri.011G046700
67 / 5e-15
AT1G61110 331 / 8e-113
NAC domain containing protein 25 (.1)
Potri.008G089000
66 / 6e-15
AT1G69490 331 / 6e-115
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
Potri.010G166200
64 / 4e-14
AT1G69490 305 / 1e-104
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
Potri.001G404400
64 / 5e-14
AT3G15510 345 / 1e-117
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02365
NAM
No apical meristem (NAM) protein
Representative CDS sequence
>Lus10005917 pacid=23140127 polypeptide=Lus10005917 locus=Lus10005917.g ID=Lus10005917.BGIv1.0 annot-version=v1.0
ATGGAGAAGATGAGTTGTGTCAAGAAAGGAGTGTTGAGACTGCCACCTAGCTTCAGATTCCACCCAACCAACGAGGAGCTCGTGGTTCAATACCTCAAAC
GGAAAGTCTGTTCCTTTCCTTTGCCCGCTTCAATCATTCCTGAAGTCAATATCAGCAAGTCTGATCCTTGA
AA sequence
>Lus10005917 pacid=23140127 polypeptide=Lus10005917 locus=Lus10005917.g ID=Lus10005917.BGIv1.0 annot-version=v1.0
MEKMSCVKKGVLRLPPSFRFHPTNEELVVQYLKRKVCSFPLPASIIPEVNISKSDP
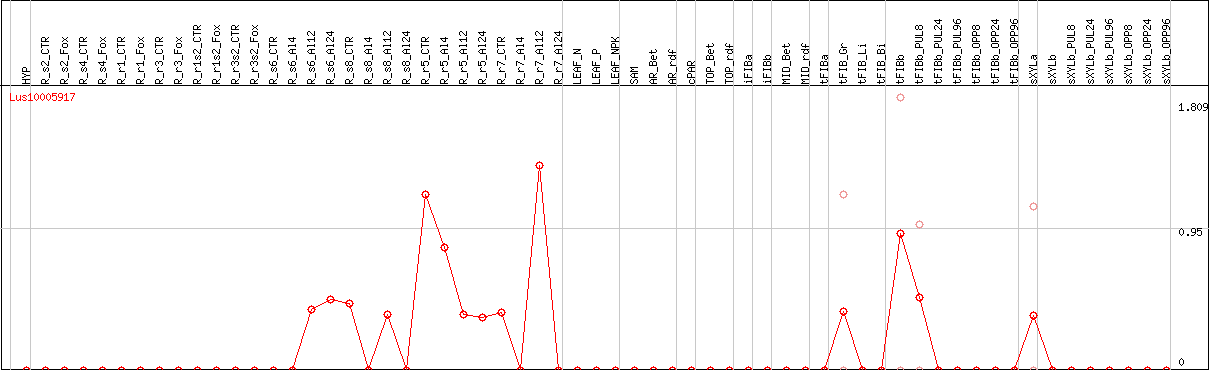
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10005917 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.