External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G59750 75 / 4e-17
ARF
ARF1
auxin response factor 1 (.1.2.3.4)
AT3G61830 72 / 5e-16
ARF
ARF18
auxin response factor 18 (.1)
AT5G62000 69 / 4e-15
ARF
ORE14, HSS, ARF1-BP, ARF2
ORESARA 14, HLS1 SUPPRESSOR, ARF1-BINDING PROTEIN, auxin response factor 2 (.1.2.3.4)
AT2G46530 64 / 5e-13
ARF
ARF11
auxin response factor 11 (.1.2.3)
AT1G34170 62 / 2e-12
ARF
ARF13
AUXIN RESPONSE FACTOR 13 (.1.2.3)
AT4G23980 59 / 1e-11
ARF
ARF9
auxin response factor 9 (.1.2)
AT1G35540 55 / 5e-10
ARF
ARF14
auxin response factor 14 (.1)
AT1G34410 53 / 2e-09
ARF
ARF21
auxin response factor 21 (.1)
AT1G34310 53 / 2e-09
ARF
ARF12
auxin response factor 12 (.1)
AT1G43950 52 / 3e-09
ARF
ARF23
auxin response factor 23 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10030240
130 / 3e-37
AT2G46530 381 / 1e-126
auxin response factor 11 (.1.2.3)
Lus10041340
111 / 4e-30
AT3G61830 370 / 2e-122
auxin response factor 18 (.1)
Lus10024455
82 / 1e-19
AT1G59750 905 / 0.0
auxin response factor 1 (.1.2.3.4)
Lus10007440
80 / 9e-19
AT1G59750 917 / 0.0
auxin response factor 1 (.1.2.3.4)
Lus10032541
74 / 9e-17
AT5G62000 1095 / 0.0
ORESARA 14, HLS1 SUPPRESSOR, ARF1-BINDING PROTEIN, auxin response factor 2 (.1.2.3.4)
Lus10043200
73 / 3e-16
AT5G62000 1091 / 0.0
ORESARA 14, HLS1 SUPPRESSOR, ARF1-BINDING PROTEIN, auxin response factor 2 (.1.2.3.4)
Lus10023058
69 / 4e-15
AT4G23980 743 / 0.0
auxin response factor 9 (.1.2)
Lus10005930
62 / 2e-12
AT5G62000 361 / 2e-112
ORESARA 14, HLS1 SUPPRESSOR, ARF1-BINDING PROTEIN, auxin response factor 2 (.1.2.3.4)
Lus10040906
61 / 3e-12
AT5G62000 367 / 2e-114
ORESARA 14, HLS1 SUPPRESSOR, ARF1-BINDING PROTEIN, auxin response factor 2 (.1.2.3.4)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G228800
80 / 9e-19
AT1G59750 906 / 0.0
auxin response factor 1 (.1.2.3.4)
Potri.003G001000
76 / 3e-17
AT1G59750 898 / 0.0
auxin response factor 1 (.1.2.3.4)
Potri.001G066200
73 / 2e-16
AT5G62000 765 / 0.0
ORESARA 14, HLS1 SUPPRESSOR, ARF1-BINDING PROTEIN, auxin response factor 2 (.1.2.3.4)
Potri.012G106100
72 / 4e-16
AT5G62000 1035 / 0.0
ORESARA 14, HLS1 SUPPRESSOR, ARF1-BINDING PROTEIN, auxin response factor 2 (.1.2.3.4)
Potri.003G142100
69 / 4e-15
AT4G23980 706 / 0.0
auxin response factor 9 (.1.2)
Potri.014G100100
68 / 1e-14
AT4G23980 694 / 0.0
auxin response factor 9 (.1.2)
Potri.002G172800
68 / 1e-14
AT4G23980 662 / 0.0
auxin response factor 9 (.1.2)
Potri.015G105300
68 / 1e-14
AT5G62000 1065 / 0.0
ORESARA 14, HLS1 SUPPRESSOR, ARF1-BINDING PROTEIN, auxin response factor 2 (.1.2.3.4)
Potri.003G163600
68 / 2e-14
AT5G62000 691 / 0.0
ORESARA 14, HLS1 SUPPRESSOR, ARF1-BINDING PROTEIN, auxin response factor 2 (.1.2.3.4)
Potri.001G088600
66 / 4e-14
AT4G23980 710 / 0.0
auxin response factor 9 (.1.2)
PFAM info
Representative CDS sequence
>Lus10005970 pacid=23175076 polypeptide=Lus10005970 locus=Lus10005970.g ID=Lus10005970.BGIv1.0 annot-version=v1.0
ATGGCTTCTGCTAATGTTCCAACTGGTTCATCTTCCTCAAGGGTCAATAAGAATGATGCTCTGCTTGATCAGCTCTGGAACGTATGTGCAAGAACAACAT
ATGGTATCCCGAAAGAAGAAGAAATGGTTTACTATTTCCTCCAGGGACACATGGAACAACTGCAAGGGACGATGAACGTGCAACATGACCACCAAATCAC
TGTCTACGACTTCCCTTGCAAAATCTTTTGCCAAATTATCAACGTCACGAGAATGGTCGGGATTTACCTAACTGTTGTGTCACAAAGCGACAATCTATGG
TAG
AA sequence
>Lus10005970 pacid=23175076 polypeptide=Lus10005970 locus=Lus10005970.g ID=Lus10005970.BGIv1.0 annot-version=v1.0
MASANVPTGSSSSRVNKNDALLDQLWNVCARTTYGIPKEEEMVYYFLQGHMEQLQGTMNVQHDHQITVYDFPCKIFCQIINVTRMVGIYLTVVSQSDNLW
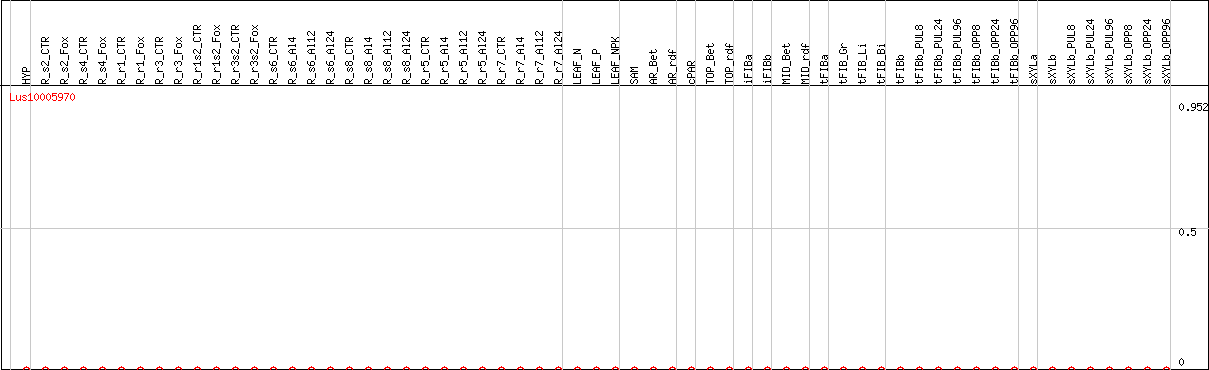
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10005970 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.