External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G71960 355 / 3e-118
ATABCG25, ABCG25
Arabidopsis thaliana ATP-binding cassette G25, ATP-binding casette G25, ATP-binding casette family G25 (.1)
AT1G31770 261 / 4e-82
ABCG14
ATP-binding cassette G14, ATP-binding cassette 14 (.1)
AT5G06530 234 / 7e-71
AtABCG22, ABCG22
Arabidopsis thaliana ATP-binding cassette G22, ATP-binding cassette G22, ABC-2 type transporter family protein (.1.2.3)
AT3G25620 226 / 1e-68
ABCG21
ATP-binding cassette G21, ABC-2 type transporter family protein (.1.2)
AT3G52310 214 / 3e-63
ABCG27
ATP-binding cassette G27, ABC-2 type transporter family protein (.1)
AT4G27420 208 / 4e-62
ABCG9
ATP-binding cassette G9, ABC-2 type transporter family protein (.1)
AT3G13220 133 / 2e-34
MSR02, AtABCG26, ABCG26, WBC27
ATP-binding cassette G26, ABC-2 type transporter family protein (.1)
AT2G01320 82 / 4e-17
ABCG7
ATP-binding cassette G7, ABC-2 type transporter family protein (.1.2.3.4)
AT2G13610 74 / 2e-14
ABCG5
ATP-binding cassette G5, ABC-2 type transporter family protein (.1)
AT3G53480 65 / 3e-11
PIS1, ABCG37, PDR9, ATPDR9
polar auxin transport inhibitor sensitive 1, ATP-binding cassette G37, pleiotropic drug resistance 9 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10028704
549 / 0
AT1G71960 818 / 0.0
Arabidopsis thaliana ATP-binding cassette G25, ATP-binding casette G25, ATP-binding casette family G25 (.1)
Lus10035692
265 / 2e-83
AT1G31770 1001 / 0.0
ATP-binding cassette G14, ATP-binding cassette 14 (.1)
Lus10033314
258 / 1e-81
AT1G31770 941 / 0.0
ATP-binding cassette G14, ATP-binding cassette 14 (.1)
Lus10037283
258 / 1e-80
AT1G31770 984 / 0.0
ATP-binding cassette G14, ATP-binding cassette 14 (.1)
Lus10034775
253 / 1e-78
AT1G31770 1028 / 0.0
ATP-binding cassette G14, ATP-binding cassette 14 (.1)
Lus10034387
249 / 4e-77
AT3G25620 791 / 0.0
ATP-binding cassette G21, ABC-2 type transporter family protein (.1.2)
Lus10015994
229 / 5e-70
AT5G06530 1033 / 0.0
Arabidopsis thaliana ATP-binding cassette G22, ATP-binding cassette G22, ABC-2 type transporter family protein (.1.2.3)
Lus10029635
216 / 7e-65
AT4G27420 698 / 0.0
ATP-binding cassette G9, ABC-2 type transporter family protein (.1)
Lus10025597
213 / 7e-64
AT4G27420 707 / 0.0
ATP-binding cassette G9, ABC-2 type transporter family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G111900
412 / 3e-140
AT1G71960 795 / 0.0
Arabidopsis thaliana ATP-binding cassette G25, ATP-binding casette G25, ATP-binding casette family G25 (.1)
Potri.019G083000
397 / 1e-134
AT1G71960 782 / 0.0
Arabidopsis thaliana ATP-binding cassette G25, ATP-binding casette G25, ATP-binding casette family G25 (.1)
Potri.004G236500
255 / 9e-80
AT1G31770 970 / 0.0
ATP-binding cassette G14, ATP-binding cassette 14 (.1)
Potri.010G131900
249 / 4e-77
AT3G25620 852 / 0.0
ATP-binding cassette G21, ABC-2 type transporter family protein (.1.2)
Potri.003G046800
234 / 2e-71
AT1G31770 945 / 0.0
ATP-binding cassette G14, ATP-binding cassette 14 (.1)
Potri.001G404000
229 / 3e-70
AT4G27420 690 / 0.0
ATP-binding cassette G9, ABC-2 type transporter family protein (.1)
Potri.016G065900
225 / 1e-67
AT5G06530 1139 / 0.0
Arabidopsis thaliana ATP-binding cassette G22, ATP-binding cassette G22, ABC-2 type transporter family protein (.1.2.3)
Potri.011G123200
207 / 7e-62
AT4G27420 737 / 0.0
ATP-binding cassette G9, ABC-2 type transporter family protein (.1)
Potri.006G199500
204 / 7e-60
AT5G06530 1013 / 0.0
Arabidopsis thaliana ATP-binding cassette G22, ATP-binding cassette G22, ABC-2 type transporter family protein (.1.2.3)
Potri.001G370600
156 / 1e-42
AT3G13220 1012 / 0.0
ATP-binding cassette G26, ABC-2 type transporter family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0181
ABC-2
PF01061
ABC2_membrane
ABC-2 type transporter
Representative CDS sequence
>Lus10006046 pacid=23147752 polypeptide=Lus10006046 locus=Lus10006046.g ID=Lus10006046.BGIv1.0 annot-version=v1.0
ATGGCATCCTATAATGCAATTCTGGCTCCAAAGGTGAAATCAGCTTGCTTGGAAGCCACGGCAAAGGAGTTGTCAAGTGTTACCAATGGTGGCATCAACA
ACAAAAATTCGTCATCGAAATCGCATCTGAGCTTCCGAGGAGAGCACAGAGTTAGTCTCACGACTTGGTTCCACCAATTCACCATTCTCCTATGCCGTAG
CTTGAAGGAGAGAAGGCACGAGTCTTTCAACACACTCCGCGTGTTCCAGGTCCTGGCGGGGGCTCTGTTGGCCGGTTTGATGTGGTGGCACTCGGATTTC
CGAGACATCCAGGACCGGCTCGGCTTGTTATTCTTCATATCGATCTTCTGGGGCGTCCTGCCTTCGTTCAACGCGGTCTTCGCCTTCCCTCAGGAGCGTG
CCATATTCATGAAGGAAAGGGCGTCCGGAATGTACACGTTGTCATCCTACTTCATGGCTCGAATCGTGGGGGATATGCCGATGGAGCTAGCCCTTCCGGC
TATCTTCATCGCCATGGCCTACTGGATGACCGGAATGAAGCCCGATTTGGGTGCGTTTCTCTTGACTCTTCTCATCCTACTAAGCTACGTTCTTGTGTCG
CAAGGCCTAGGTCTAGCCCTTGGAGCGGCCATCATGGACGCCAAGCAAGCATCCACGTTGGCCACTGTTACGATGCTCGCATTCGTGCTGACGGGAGGGT
TCTACGTGCACAAGGTCCCGACATGTATGGTGTGGATGAAGTACGTCTCCACCACTTTCTACACTTACAGGTTGCTCATCAACGTCCAGTACGGGGACGG
GGAGCGGATCGCAAAGGCGTTGGGCTGCTCGGGGCGAAACGGGAGTAGCGGGGGCGATGGAGGACCGAACTGCAAGTTTGTCAGGGAAGATGTGCAAGGA
CAAATCAGCCCTTGGTTTAGTGTTGGGGTGATGGTGTTCATGTTTCTGGGGTATAGAGTGCTGGCTTACTTGGCTTTGAGGAGGATCAGAACTTGA
AA sequence
>Lus10006046 pacid=23147752 polypeptide=Lus10006046 locus=Lus10006046.g ID=Lus10006046.BGIv1.0 annot-version=v1.0
MASYNAILAPKVKSACLEATAKELSSVTNGGINNKNSSSKSHLSFRGEHRVSLTTWFHQFTILLCRSLKERRHESFNTLRVFQVLAGALLAGLMWWHSDF
RDIQDRLGLLFFISIFWGVLPSFNAVFAFPQERAIFMKERASGMYTLSSYFMARIVGDMPMELALPAIFIAMAYWMTGMKPDLGAFLLTLLILLSYVLVS
QGLGLALGAAIMDAKQASTLATVTMLAFVLTGGFYVHKVPTCMVWMKYVSTTFYTYRLLINVQYGDGERIAKALGCSGRNGSSGGDGGPNCKFVREDVQG
QISPWFSVGVMVFMFLGYRVLAYLALRRIRT
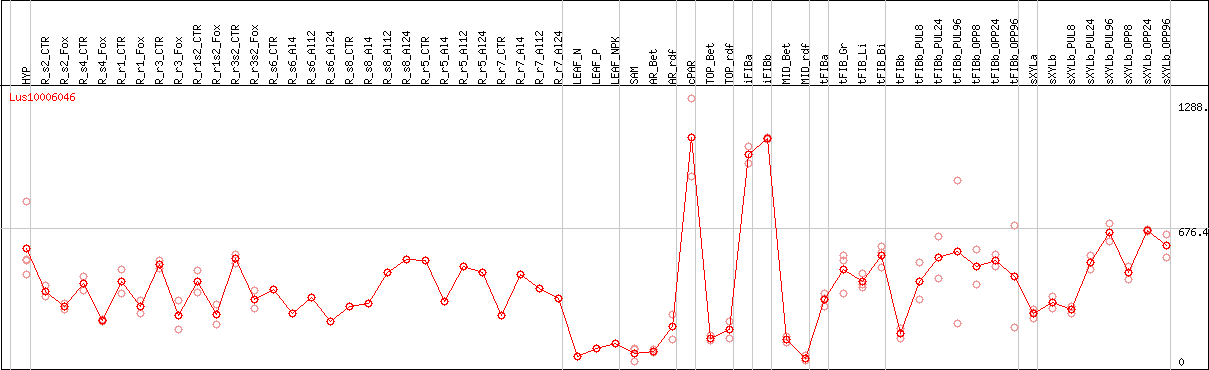
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10006046 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.