External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G62990 169 / 1e-49
EMB1692
embryo defective 1692, Ubiquitin carboxyl-terminal hydrolase family protein (.1)
AT4G01037 96 / 2e-22
AtWTF1
what's this factor?, Ubiquitin carboxyl-terminal hydrolase family protein (.1)
AT2G31290 75 / 2e-15
Ubiquitin carboxyl-terminal hydrolase family protein (.1.2)
AT5G21970 74 / 5e-15
Ubiquitin carboxyl-terminal hydrolase family protein (.1)
AT1G06440 68 / 6e-13
Ubiquitin carboxyl-terminal hydrolase family protein (.1)
AT5G45790 66 / 4e-12
Ubiquitin carboxyl-terminal hydrolase family protein (.1.2)
AT4G33495 64 / 1e-11
RPD1
ROOT PRIMORDIUM DEFECTIVE 1, Ubiquitin carboxyl-terminal hydrolase family protein (.1)
AT3G58520 60 / 3e-10
Ubiquitin carboxyl-terminal hydrolase family protein (.1.2)
AT1G79120 51 / 2e-07
Ubiquitin carboxyl-terminal hydrolase family protein (.1)
AT2G39120 51 / 2e-07
WTF9
what's this factor 9, Ubiquitin carboxyl-terminal hydrolase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10010567
340 / 8e-116
AT5G62990 509 / 2e-178
embryo defective 1692, Ubiquitin carboxyl-terminal hydrolase family protein (.1)
Lus10029446
93 / 2e-21
AT4G01037 588 / 0.0
what's this factor?, Ubiquitin carboxyl-terminal hydrolase family protein (.1)
Lus10005945
93 / 2e-21
AT4G01037 582 / 0.0
what's this factor?, Ubiquitin carboxyl-terminal hydrolase family protein (.1)
Lus10016529
73 / 1e-14
AT5G21970 561 / 0.0
Ubiquitin carboxyl-terminal hydrolase family protein (.1)
Lus10003062
64 / 2e-11
AT2G31290 558 / 0.0
Ubiquitin carboxyl-terminal hydrolase family protein (.1.2)
Lus10034086
63 / 3e-11
AT2G31290 559 / 0.0
Ubiquitin carboxyl-terminal hydrolase family protein (.1.2)
Lus10005755
63 / 4e-11
AT4G33495 608 / 0.0
ROOT PRIMORDIUM DEFECTIVE 1, Ubiquitin carboxyl-terminal hydrolase family protein (.1)
Lus10027440
62 / 5e-11
AT4G33495 607 / 0.0
ROOT PRIMORDIUM DEFECTIVE 1, Ubiquitin carboxyl-terminal hydrolase family protein (.1)
Lus10041673
62 / 7e-11
AT1G06440 495 / 2e-175
Ubiquitin carboxyl-terminal hydrolase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G233900
196 / 8e-60
AT5G62990 498 / 6e-174
embryo defective 1692, Ubiquitin carboxyl-terminal hydrolase family protein (.1)
Potri.014G096800
99 / 2e-23
AT4G01037 588 / 0.0
what's this factor?, Ubiquitin carboxyl-terminal hydrolase family protein (.1)
Potri.002G169300
96 / 2e-22
AT4G01037 586 / 0.0
what's this factor?, Ubiquitin carboxyl-terminal hydrolase family protein (.1)
Potri.006G218100
79 / 1e-16
AT5G21970 566 / 0.0
Ubiquitin carboxyl-terminal hydrolase family protein (.1)
Potri.015G141700
69 / 2e-13
AT4G33495 607 / 0.0
ROOT PRIMORDIUM DEFECTIVE 1, Ubiquitin carboxyl-terminal hydrolase family protein (.1)
Potri.002G245100
69 / 3e-13
AT1G06440 544 / 0.0
Ubiquitin carboxyl-terminal hydrolase family protein (.1)
Potri.005G191100
68 / 4e-13
AT2G31290 575 / 0.0
Ubiquitin carboxyl-terminal hydrolase family protein (.1.2)
Potri.016G030300
67 / 1e-12
AT5G45790 466 / 8e-163
Ubiquitin carboxyl-terminal hydrolase family protein (.1.2)
Potri.002G244900
66 / 2e-12
AT1G06440 503 / 7e-180
Ubiquitin carboxyl-terminal hydrolase family protein (.1)
Potri.002G220000
54 / 2e-08
AT3G63090 499 / 7e-177
Ubiquitin carboxyl-terminal hydrolase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF11955
PORR
Plant organelle RNA recognition domain
Representative CDS sequence
>Lus10006104 pacid=23174770 polypeptide=Lus10006104 locus=Lus10006104.g ID=Lus10006104.BGIv1.0 annot-version=v1.0
ATGCTGGGTCATTACAGGATTCATAGATCCAGGTCGTGCTCTGTTGCAGTGACTCATGATGGCTGCTGCAACCGCAAAGGGCTGAATCTTAAGAGGCATC
ACGAGGGTTTCCTGATCAAGTGTTCGGAGCTTCCGGATGTTTGTCGTTATAACACCACGATAGAGGAGTTGAGTAAAGAATCATTGGAAGCTGAGAAGAG
GGCTTGCGCTCTGGTGAGGGAGGTGCTGGGATTGACAGCTGAGAAGAGAACTTTGATAGATCATTTGACGCATTTCAGGAAAGAATTCGGTCTGCCTAAC
AAGTTGAAGAATATGATTCTACGCCACCCGGAGATGTTCTACGTTAGCGTGAAAGGTGTAAGGAATTCTGTTTTCCTTGTCGAGGGTTTCAATGACAAAG
GTAAACTCTATGTGAAAGATGATACGTCTGTTATCAAAGATAAGTTGATGCATTTGGTGAGGGAATCGAAAAGAATGAGACAAGAGAGGCGCAAGGCTGC
GATGAACAGAGGTGTTGATGGAGTTGTTCCTAATGGTATTGGCAGTGATGATGTATCTCATGATGTCGATGATGAGTATGATGATGGTTTCGAGAATATC
TTTGAGGATTCGGACCTTGAAGATTTTGATGTCGATTCCATTGAAGACGTTGAGTTTTGGGGAAATAGAGAGAACACAGAATTTTGGGCTGCTGATGGTT
CTTTTGTTCCTCAAGAGGGGGTAGTACCACCTCCAGAACCATGGTAG
AA sequence
>Lus10006104 pacid=23174770 polypeptide=Lus10006104 locus=Lus10006104.g ID=Lus10006104.BGIv1.0 annot-version=v1.0
MLGHYRIHRSRSCSVAVTHDGCCNRKGLNLKRHHEGFLIKCSELPDVCRYNTTIEELSKESLEAEKRACALVREVLGLTAEKRTLIDHLTHFRKEFGLPN
KLKNMILRHPEMFYVSVKGVRNSVFLVEGFNDKGKLYVKDDTSVIKDKLMHLVRESKRMRQERRKAAMNRGVDGVVPNGIGSDDVSHDVDDEYDDGFENI
FEDSDLEDFDVDSIEDVEFWGNRENTEFWAADGSFVPQEGVVPPPEPW
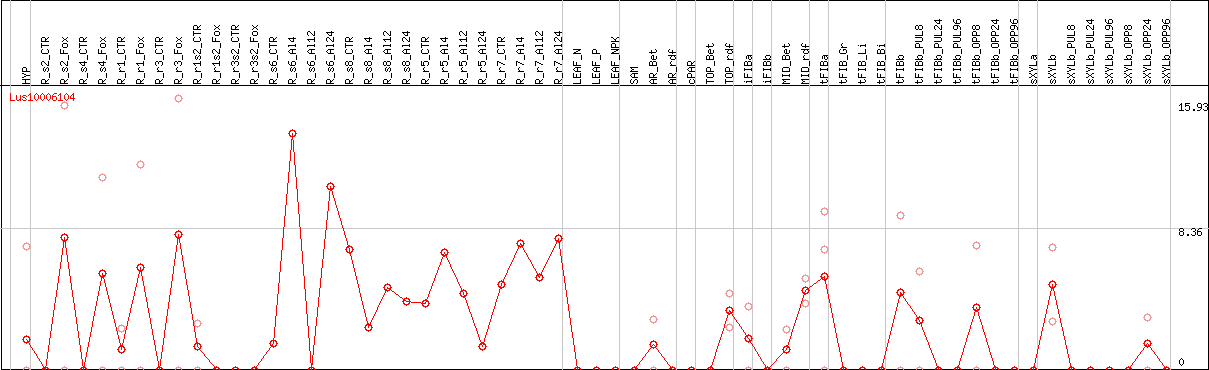
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10006104 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.