External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G32090 199 / 4e-67
Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
AT2G28420 64 / 1e-13
GLYI8
glyoxylase I 8, Lactoylglutathione lyase / glyoxalase I family protein (.1)
AT1G80160 50 / 2e-08
GLYI7
glyoxylase I 7, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
AT1G15380 49 / 5e-08
GLYI4
glyoxylase I 4, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10010552
282 / 9e-100
AT2G32090 197 / 3e-66
Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Lus10039579
58 / 6e-11
AT2G28420 248 / 2e-84
glyoxylase I 8, Lactoylglutathione lyase / glyoxalase I family protein (.1)
Lus10005324
55 / 8e-10
AT2G28420 245 / 2e-83
glyoxylase I 8, Lactoylglutathione lyase / glyoxalase I family protein (.1)
Lus10030070
47 / 7e-07
AT1G80160 247 / 7e-85
glyoxylase I 7, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Lus10014480
46 / 8e-07
AT1G80160 244 / 1e-83
glyoxylase I 7, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Lus10025971
44 / 5e-06
AT1G15380 165 / 3e-52
glyoxylase I 4, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Lus10014269
44 / 1e-05
AT1G15380 166 / 2e-52
glyoxylase I 4, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G087000
217 / 5e-74
AT2G32090 189 / 3e-63
Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Potri.008G153602
156 / 2e-49
AT2G32090 139 / 8e-43
Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Potri.009G055500
59 / 1e-11
AT2G28420 229 / 1e-77
glyoxylase I 8, Lactoylglutathione lyase / glyoxalase I family protein (.1)
Potri.001G171600
52 / 6e-09
AT1G80160 270 / 6e-94
glyoxylase I 7, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Potri.003G009000
51 / 2e-08
AT1G80160 190 / 8e-62
glyoxylase I 7, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Potri.003G062400
50 / 4e-08
AT1G80160 261 / 2e-90
glyoxylase I 7, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Potri.004G223300
50 / 4e-08
AT1G80160 187 / 1e-60
glyoxylase I 7, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Potri.007G015100
47 / 7e-07
AT1G15380 167 / 7e-53
glyoxylase I 4, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Potri.005G117000
45 / 3e-06
AT1G80160 166 / 2e-52
glyoxylase I 7, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
Potri.007G129200
41 / 4e-05
AT1G80160 225 / 1e-76
glyoxylase I 7, Lactoylglutathione lyase / glyoxalase I family protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0104
Glyoxalase
PF00903
Glyoxalase
Glyoxalase/Bleomycin resistance protein/Dioxygenase superfamily
Representative CDS sequence
>Lus10006122 pacid=23174766 polypeptide=Lus10006122 locus=Lus10006122.g ID=Lus10006122.BGIv1.0 annot-version=v1.0
ATGGCGGCAGCCTCGGCACCTATTCTCAGCCACATCGCCAGAGAATCCACAGACATTAGACGCCTGGCCAATTTCTACAAGGAGATCTTTGGGTTTGAGG
AGATCGAGACTCCCGATTTTGGATTCAACGTGATATGGCTGAATCTGGCGAATGCTTTCTCTTTCCACCTCATTGAGAGGAGCCCCGAGACCAAGCTTCC
AGAAGGACCTTACAGTGCTTCAGAGCCGATTCGCGATGTCACCCATCTCGCCAGAGGCCATCATATCTGTATCTCCGTCGCCAACTTCGACGCTTTTGTT
CAATCTCTAAAGGAGAAAGGGATAGAGACGTTCCAGAGGGCAGTTCCTGGTCGGTCAATTAGGCAAGTTTTCTTCTTTGATCCAGATGGCAATGGACTGG
AGGTGGCAAGTCGGGATGAGTAG
AA sequence
>Lus10006122 pacid=23174766 polypeptide=Lus10006122 locus=Lus10006122.g ID=Lus10006122.BGIv1.0 annot-version=v1.0
MAAASAPILSHIARESTDIRRLANFYKEIFGFEEIETPDFGFNVIWLNLANAFSFHLIERSPETKLPEGPYSASEPIRDVTHLARGHHICISVANFDAFV
QSLKEKGIETFQRAVPGRSIRQVFFFDPDGNGLEVASRDE
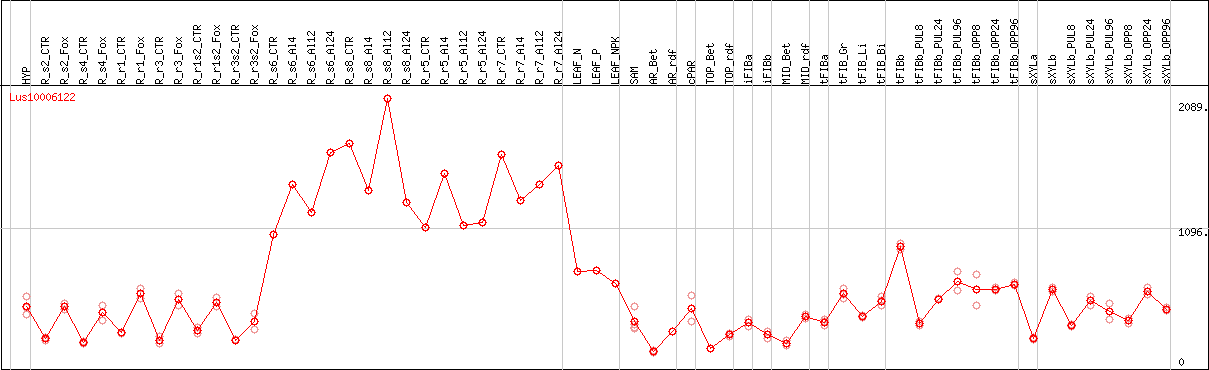
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10006122 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.