External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G77630 43 / 6e-05
LYM3
lysin-motif \(LysM\) domain protein 3, Peptidoglycan-binding LysM domain-containing protein (.1)
AT1G21880 42 / 0.0002
LYM1
lysm domain GPI-anchored protein 1 precursor (.1.2)
AT1G51940 40 / 0.0007
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10036847
318 / 1e-111
AT1G51940 42 / 1e-04
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Lus10004625
167 / 3e-48
AT1G51940 347 / 7e-111
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Lus10026689
162 / 1e-46
AT1G51940 348 / 4e-111
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Lus10027677
73 / 9e-15
AT1G51940 320 / 7e-101
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Lus10037586
52 / 9e-08
AT1G51940 807 / 0.0
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Lus10006841
52 / 1e-07
AT1G51940 812 / 0.0
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Lus10008586
47 / 3e-06
AT2G33580 594 / 0.0
Protein kinase superfamily protein (.1)
Lus10014441
44 / 5e-05
AT4G38380 501 / 2e-167
MATE efflux family protein (.1)
Lus10026372
42 / 0.0002
AT2G17120 249 / 3e-80
lysm domain GPI-anchored protein 2 precursor (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.015G081601
144 / 3e-40
AT1G51940 337 / 5e-107
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Potri.T125208
144 / 3e-40
AT1G51940 336 / 9e-107
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Potri.008G187500
117 / 2e-30
AT1G51940 362 / 7e-117
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Potri.006G252600
52 / 1e-07
AT1G51940 326 / 4e-103
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Potri.001G190200
50 / 4e-07
AT1G51940 824 / 0.0
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Potri.008G160501
46 / 5e-06
AT2G23770 66 / 3e-12
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Potri.007G032300
45 / 2e-05
AT2G33580 247 / 6e-73
Protein kinase superfamily protein (.1)
Potri.005G128200
45 / 2e-05
AT2G23770 514 / 4e-176
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Potri.007G032100
41 / 0.0003
AT2G23770 513 / 6e-175
protein kinase family protein / peptidoglycan-binding LysM domain-containing protein (.1)
Potri.008G160600
41 / 0.0004
AT2G33580 217 / 8e-62
Protein kinase superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0187
LysM
PF01476
LysM
LysM domain
Representative CDS sequence
>Lus10006200 pacid=23139734 polypeptide=Lus10006200 locus=Lus10006200.g ID=Lus10006200.BGIv1.0 annot-version=v1.0
ATGGCTTCACTTCAATTCCTTCTCATCCTCACCTTCATAATTCCCCTCTTCAACGCACAACCACCAACCATGTTCCCATTCCCATGCACCAAAAAGATCA
TCAACACATGCACATCAACCCTCTACCACATCACCACCACAAACTCCACCACCACCCTCCAAGAACTCGGCCACAATTACTCGGTCCAACCATCCCAAAT
CAAACCTGTTACCCACATCGAGAAGCAGGACTTCCTCATTACAGTCTCTTGCTCCTGCCAGACTGCTGCCGACCAGACGACCGCCTACTTCTACAACGTT
GACTACACGGTCAATATTGGGGACACACTCTTCGACGTGTCAACGAATGCCTACAGCGGGCAAGTGTGGGCCACGGGCGGCGAGCAGGATAGGTTCGTTG
CCGGGAACAATGTGACCATGAACTTGTTGTGCGGGTGTCTCGGGAGCACGGCGGAGAACGTTGTGACGTATACTGTTCAGGATGGGGACACTTTGTTCGG
GATTGCTAGAATGTTTGGTAGTGATTTGGGCGAGATTCATAAGCTTAATAACAAGTTTTTAGGGAATTCCAGTGAGTTTATTGATGTGGGTTGGGTTCTT
TATGTGCCGGTAGGAACTAATGGAACTTTGTTCTAG
AA sequence
>Lus10006200 pacid=23139734 polypeptide=Lus10006200 locus=Lus10006200.g ID=Lus10006200.BGIv1.0 annot-version=v1.0
MASLQFLLILTFIIPLFNAQPPTMFPFPCTKKIINTCTSTLYHITTTNSTTTLQELGHNYSVQPSQIKPVTHIEKQDFLITVSCSCQTAADQTTAYFYNV
DYTVNIGDTLFDVSTNAYSGQVWATGGEQDRFVAGNNVTMNLLCGCLGSTAENVVTYTVQDGDTLFGIARMFGSDLGEIHKLNNKFLGNSSEFIDVGWVL
YVPVGTNGTLF
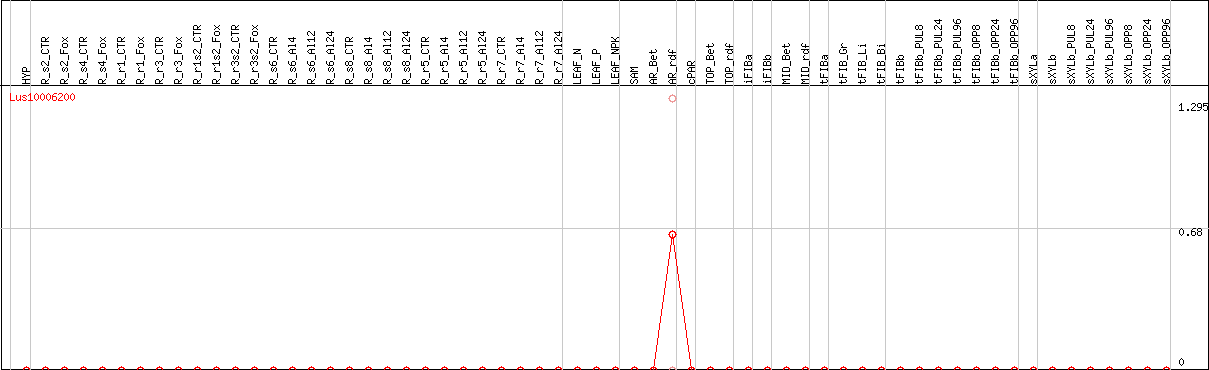
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10006200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.