External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G45490 429 / 4e-153
ATAUR3
ataurora3 (.1)
AT4G32830 334 / 7e-116
ATAUR1
ataurora1 (.1)
AT2G25880 334 / 7e-116
ATAUR2
ataurora2 (.1.2)
AT2G26980 184 / 3e-56
CIPK3, SnRK3.17
SNF1-RELATED PROTEIN KINASE 3.17, CBL-interacting protein kinase 3 (.1.2.3.4.5)
AT1G30270 184 / 9e-55
PKS17, ATCIPK23, SnRK3.23, LKS1, CIPK23
SNF1-RELATED PROTEIN KINASE 3.23, SOS2-like protein kinase 17, LOW-K+-SENSITIVE 1, CBL-interacting protein kinase 23 (.1)
AT5G21326 179 / 2e-53
Ca2+regulated serine-threonine protein kinase family protein (.1)
AT5G10930 179 / 4e-53
CIPK5, SnRK3.24
SNF1-RELATED PROTEIN KINASE 3.24, CBL-interacting protein kinase 5 (.1)
AT5G25110 179 / 6e-53
CIPK25, SnRK3.25
SNF1-RELATED PROTEIN KINASE 3.25, CBL-interacting protein kinase 25 (.1)
AT1G01140 175 / 1e-51
PKS6, CIPK9, SnRK3.12
SNF1-RELATED PROTEIN KINASE 3.12, PROTEIN KINASE 6, CBL-interacting protein kinase 9 (.1.2.3)
AT5G01810 171 / 4e-51
ATPK10, SnRK3.1, SIP2, PKS3, CIPK15
SNF1-RELATED PROTEIN KINASE 3.1, SOS3-INTERACTING PROTEIN 2, PROTEIN KINASE 10, CBL-interacting protein kinase 15 (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10020580
499 / 0
AT2G45490 482 / 3e-174
ataurora3 (.1)
Lus10005487
335 / 5e-116
AT4G32830 545 / 0.0
ataurora1 (.1)
Lus10006033
335 / 9e-116
AT4G32830 488 / 2e-175
ataurora1 (.1)
Lus10042229
185 / 2e-55
AT5G58380 676 / 0.0
SNF1-RELATED PROTEIN KINASE 3.8, CBL-INTERACTING PROTEIN KINASE 10, SOS3-interacting protein 1 (.1)
Lus10028115
184 / 3e-55
AT1G30270 793 / 0.0
SNF1-RELATED PROTEIN KINASE 3.23, SOS2-like protein kinase 17, LOW-K+-SENSITIVE 1, CBL-interacting protein kinase 23 (.1)
Lus10042816
184 / 4e-55
AT1G30270 794 / 0.0
SNF1-RELATED PROTEIN KINASE 3.23, SOS2-like protein kinase 17, LOW-K+-SENSITIVE 1, CBL-interacting protein kinase 23 (.1)
Lus10024448
179 / 8e-53
AT5G25110 654 / 0.0
SNF1-RELATED PROTEIN KINASE 3.25, CBL-interacting protein kinase 25 (.1)
Lus10007448
178 / 1e-52
AT5G25110 650 / 0.0
SNF1-RELATED PROTEIN KINASE 3.25, CBL-interacting protein kinase 25 (.1)
Lus10021852
173 / 5e-51
AT4G30960 658 / 0.0
SNF1-RELATED PROTEIN KINASE 3.14, CBL-INTERACTING PROTEIN KINASE 6, SOS3-interacting protein 3 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G080500
417 / 6e-149
AT2G45490 470 / 2e-169
ataurora3 (.1)
Potri.006G235000
337 / 8e-117
AT4G32830 553 / 0.0
ataurora1 (.1)
Potri.018G057700
330 / 2e-114
AT4G32830 544 / 0.0
ataurora1 (.1)
Potri.002G177900
189 / 3e-57
AT1G01140 711 / 0.0
SNF1-RELATED PROTEIN KINASE 3.12, PROTEIN KINASE 6, CBL-interacting protein kinase 9 (.1.2.3)
Potri.014G104200
187 / 2e-56
AT1G01140 688 / 0.0
SNF1-RELATED PROTEIN KINASE 3.12, PROTEIN KINASE 6, CBL-interacting protein kinase 9 (.1.2.3)
Potri.006G062800
186 / 7e-56
AT1G30270 753 / 0.0
SNF1-RELATED PROTEIN KINASE 3.23, SOS2-like protein kinase 17, LOW-K+-SENSITIVE 1, CBL-interacting protein kinase 23 (.1)
Potri.018G119200
186 / 8e-56
AT1G30270 749 / 0.0
SNF1-RELATED PROTEIN KINASE 3.23, SOS2-like protein kinase 17, LOW-K+-SENSITIVE 1, CBL-interacting protein kinase 23 (.1)
Potri.001G222600
180 / 8e-54
AT2G26980 752 / 0.0
SNF1-RELATED PROTEIN KINASE 3.17, CBL-interacting protein kinase 3 (.1.2.3.4.5)
Potri.013G156000
179 / 2e-53
AT5G58380 623 / 0.0
SNF1-RELATED PROTEIN KINASE 3.8, CBL-INTERACTING PROTEIN KINASE 10, SOS3-interacting protein 1 (.1)
Potri.006G263500
179 / 4e-53
AT5G25110 568 / 0.0
SNF1-RELATED PROTEIN KINASE 3.25, CBL-interacting protein kinase 25 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0016
PKinase
PF00069
Pkinase
Protein kinase domain
Representative CDS sequence
>Lus10006266 pacid=23181457 polypeptide=Lus10006266 locus=Lus10006266.g ID=Lus10006266.BGIv1.0 annot-version=v1.0
ATGGTGGCCGAAAAACCAGGAACCAGGCGGCAGTGGTCGTTGGCGGACTTTGAGACGGGAAAGCCACTCGGGAAAGGGAAGTTCGGCAGAGTCTATCTCG
CTCGTGAAGTCAAGAGCAAGTACATAGTGGCGCTGAAGGTGATATTCAAGGCACAGATTGAGAAGTACAAGATTCAGAATCAATTGAGGAGAGAAATGGA
GATCCAGACCACTCTCAAACACCCCAACGTGTTACGCCTCTACGGCTGGTTCGACGACGAGGAGAGGATCTTCCTCATTCTCGAGTACGCTCATGGCGGT
GAGCTTTATGGAGTTATGCAGAAACGAGGCCACTTGACTGAGAAGGAAGCTGCTACGTATATTGCGAGTCTTGCGTATGCATTAGCTTATTGCCATGGGA
AGGACGTGATCCACAGAGACATTAAGCCAGAGAATCTGTTGCTGGATCATGAGGGCAGGTTAAAGATTGCAGATTTTGGGTGGTCAGTACAATCGAGAAG
CAAGAGACACACCATGTGCGGAACACTGGATTACTTAGCACCAGAACTTGTAGAAAACAAAGCTCACGATTACACAGTGGATAACTGGACATTAGGAGTG
CTTTGTTACGAATTCCTCTACGGTAACCCTCCTTTCGAGGCAGAGGGGCAGCGAGATACTTTCAGAAGGATCATGAAGGTTGATCTTAACTTCCCTTCTC
ACCCATTCACCTCAGAGGAAGCAAAAGACATTATCAGCAAAGTAAGTTTGTCCCCAAAAAAAACCCCTTTTCGAGCATCTTCCTGTCAAGTTTTGGTTCT
TAACGGGAAATTTGTAAATTGA
AA sequence
>Lus10006266 pacid=23181457 polypeptide=Lus10006266 locus=Lus10006266.g ID=Lus10006266.BGIv1.0 annot-version=v1.0
MVAEKPGTRRQWSLADFETGKPLGKGKFGRVYLAREVKSKYIVALKVIFKAQIEKYKIQNQLRREMEIQTTLKHPNVLRLYGWFDDEERIFLILEYAHGG
ELYGVMQKRGHLTEKEAATYIASLAYALAYCHGKDVIHRDIKPENLLLDHEGRLKIADFGWSVQSRSKRHTMCGTLDYLAPELVENKAHDYTVDNWTLGV
LCYEFLYGNPPFEAEGQRDTFRRIMKVDLNFPSHPFTSEEAKDIISKVSLSPKKTPFRASSCQVLVLNGKFVN
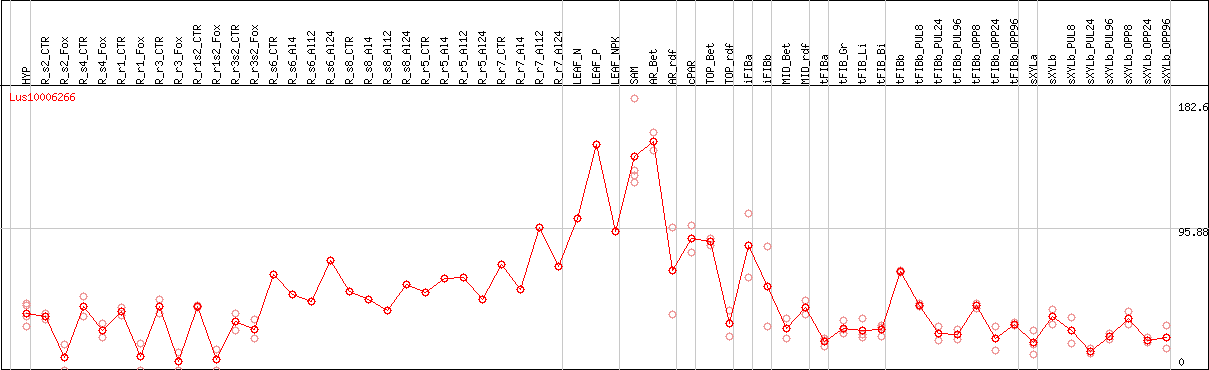
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10006266 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.