External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G22690 378 / 4e-125
unknown protein
AT5G40405 273 / 5e-87
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G62890 267 / 4e-85
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT5G66520 263 / 5e-83
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G26782 260 / 1e-81
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G08070 260 / 8e-81
EMB3102, OTP82
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G21065 253 / 9e-81
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT2G29760 256 / 1e-79
OTP81
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G14820 255 / 4e-79
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G24000 252 / 2e-78
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10029344
266 / 5e-84
AT5G06540 803 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10000706
270 / 7e-84
AT3G12770 878 / 0.0
mitochondrial editing factor 22 (.1)
Lus10024876
268 / 4e-83
AT3G12770 886 / 0.0
mitochondrial editing factor 22 (.1)
Lus10040680
265 / 1e-82
AT2G29760 926 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10038741
251 / 8e-78
AT3G24000 739 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10010909
249 / 3e-77
AT5G66520 537 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10018223
249 / 7e-77
AT2G29760 931 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10016387
243 / 9e-77
AT4G33990 717 / 0.0
embryo defective 2758, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10010385
245 / 4e-76
AT4G37380 758 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G083700
441 / 1e-149
AT3G22690 1050 / 0.0
unknown protein
Potri.018G071800
286 / 4e-91
AT1G08070 606 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.006G252400
273 / 3e-88
AT4G21065 496 / 8e-172
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.007G050200
271 / 3e-86
AT4G37380 817 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.013G044700
274 / 9e-86
AT3G22690 582 / 0.0
unknown protein
Potri.003G031600
268 / 7e-84
AT3G15930 784 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.016G065500
263 / 3e-83
AT5G06540 828 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.018G040100
261 / 2e-81
AT1G08070 974 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.014G191200
258 / 3e-81
AT5G48910 852 / 0.0
LOW PSII ACCUMULATION 66, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.006G105700
259 / 7e-81
AT3G08820 810 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10006599 pacid=23145300 polypeptide=Lus10006599 locus=Lus10006599.g ID=Lus10006599.BGIv1.0 annot-version=v1.0
ATGGAAGTTGCTTCTGCATGTGGGTATCTTGGAGGTCTGGAACTTGCTAAATGGATCTATGTGTACATTCAGAAGAACAAGATTCAGTACGATACCCGGC
TTGGCACAATCTTGATTGATATGTTTGCTAGGTGTGGTGATCCTCGTAGTGCAATGCAGGTCTTCAGTAAGATGGCAAAAAAAGATGTGTCAGCTTGGAC
AGCAGCCATCAGAGCCATGGCCATGGAAGGGAATGGCGAACAAGCTATTGAGCTTTTCAATGAGATGATACGTGAAGGAATTAAACCTGACGAGGTCGTG
TTCGTCAATTTACTGTCGGGACTGAGTCACAGTGGGTTAGTGGAAGAAGGCTGGCAGATTTTCAGGTTAATGGAGGACTCGTTTGGCTTAGAGCCCCAAG
TGGTTCATTATGGTTGCATGGTTGATTTGCTTGGTCGAGCTGGGTTTATTGCAGAAGCAGTCAATCTTATCAAGGACATGCCAATGGAACCAAATGTGGT
TGTTTGGAGTTCTCTTTTAGCTGCCTGCTGGATGCACAACAATATCAATATTGCCACATATGCAGCTGAAAGGATAACCGTACACGATCCTGGAAGACCT
GGAGTCCATGTTCTTTTGTCTAATATCTACGCATCTGCTGGGCAATGGAATGATGTTACCAAAGTCAGGCTACAAATGAAAGAGCTAGGGGTCCAAAAAG
ACCCCGGATCGAGCTCTATTCAGATCGATGGAGAAGTTTATGAGTTCACCTCTGGGACTGAATCTGTATTCCGCACGGAAAAGAACCAAACCGAACCAAT
GTTGGAAGAGATGAGTTGCAGATTGACAGATGCTGGGTTTGTTTCGGATCTCGCAAACGTTCTGCTGGATGTGGATGAGGCAGAAAAACGGTACCTGCTC
AGTAGACACAGTGAGAAGCAAGCTTTGGCTTTGCTCTCATCCGCACAAGCCCAACCATGCCAATACGAGTGA
AA sequence
>Lus10006599 pacid=23145300 polypeptide=Lus10006599 locus=Lus10006599.g ID=Lus10006599.BGIv1.0 annot-version=v1.0
MEVASACGYLGGLELAKWIYVYIQKNKIQYDTRLGTILIDMFARCGDPRSAMQVFSKMAKKDVSAWTAAIRAMAMEGNGEQAIELFNEMIREGIKPDEVV
FVNLLSGLSHSGLVEEGWQIFRLMEDSFGLEPQVVHYGCMVDLLGRAGFIAEAVNLIKDMPMEPNVVVWSSLLAACWMHNNINIATYAAERITVHDPGRP
GVHVLLSNIYASAGQWNDVTKVRLQMKELGVQKDPGSSSIQIDGEVYEFTSGTESVFRTEKNQTEPMLEEMSCRLTDAGFVSDLANVLLDVDEAEKRYLL
SRHSEKQALALLSSAQAQPCQYE
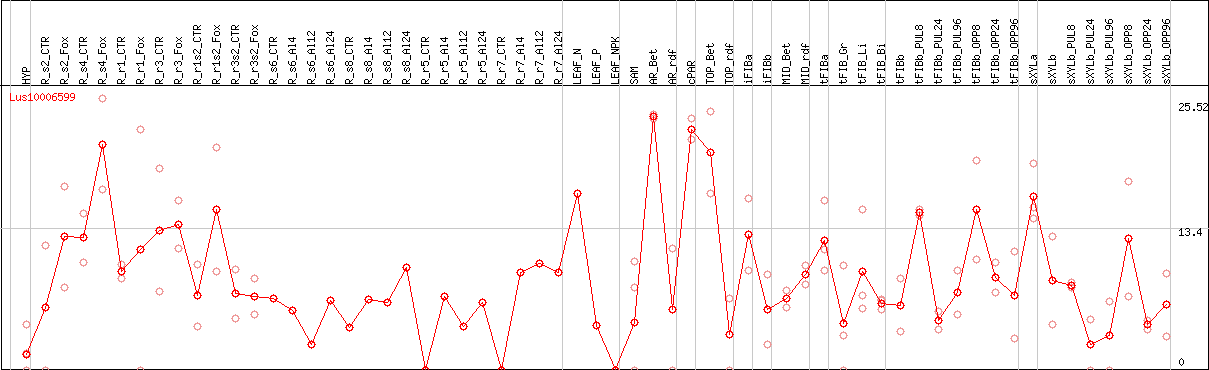
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10006599 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.