External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G61930 111 / 3e-32
Protein of unknown function, DUF584 (.1)
AT1G11700 109 / 2e-31
Protein of unknown function, DUF584 (.1)
AT4G21930 102 / 8e-29
Protein of unknown function, DUF584 (.1)
AT4G21970 79 / 3e-20
Protein of unknown function, DUF584 (.1)
AT4G04630 77 / 2e-19
Protein of unknown function, DUF584 (.1)
AT3G45210 73 / 7e-18
Protein of unknown function, DUF584 (.1)
AT2G28400 71 / 8e-17
Protein of unknown function, DUF584 (.1)
AT5G03230 69 / 3e-16
Protein of unknown function, DUF584 (.1)
AT5G60680 67 / 2e-15
Protein of unknown function, DUF584 (.1)
AT3G15040 62 / 4e-13
Protein of unknown function, DUF584 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G015700
136 / 5e-42
AT1G11700 151 / 5e-46
Protein of unknown function, DUF584 (.1)
Potri.011G003300
135 / 2e-41
AT1G11700 161 / 6e-50
Protein of unknown function, DUF584 (.1)
Potri.011G004100
81 / 1e-20
AT4G04630 125 / 1e-36
Protein of unknown function, DUF584 (.1)
Potri.009G014100
77 / 4e-19
AT3G45210 124 / 2e-36
Protein of unknown function, DUF584 (.1)
Potri.004G014000
75 / 3e-18
AT4G04630 124 / 4e-36
Protein of unknown function, DUF584 (.1)
Potri.016G089900
74 / 6e-18
AT5G03230 115 / 6e-33
Protein of unknown function, DUF584 (.1)
Potri.004G211100
74 / 6e-18
AT5G60680 120 / 1e-34
Protein of unknown function, DUF584 (.1)
Potri.016G076300
70 / 4e-16
AT3G15040 90 / 9e-22
Protein of unknown function, DUF584 (.1)
Potri.006G127900
69 / 8e-16
AT5G03230 122 / 1e-35
Protein of unknown function, DUF584 (.1)
Potri.011G076900
49 / 9e-09
AT1G29640 90 / 4e-24
Protein of unknown function, DUF584 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04520
Senescence_reg
Senescence regulator
Representative CDS sequence
>Lus10006775 pacid=23163534 polypeptide=Lus10006775 locus=Lus10006775.g ID=Lus10006775.BGIv1.0 annot-version=v1.0
ATGACCGCGGCGTCGTCGGCTCCGGTGAACGTGCCGGATTGGGGGAAGATTCTGCGGGTGAGGTCCGCGGAGTCGATGAATGAGATGAGCGACTACGACG
GAGATGATTCGGAAGAGATGATGGTCCCGCCGCATGAGTACCTAGCGCGTGAGTACGCACGGAGAAGGAAAATGGAGGGGGCGTCGGTGGTTGAAGGGGT
TGGCAGGACGTTGAAGGGGCGTGACATGAGGCGCGTGAGGGACGCTGTGTGGAGCCAAACGGGGTTTTACGGCTGA
AA sequence
>Lus10006775 pacid=23163534 polypeptide=Lus10006775 locus=Lus10006775.g ID=Lus10006775.BGIv1.0 annot-version=v1.0
MTAASSAPVNVPDWGKILRVRSAESMNEMSDYDGDDSEEMMVPPHEYLAREYARRRKMEGASVVEGVGRTLKGRDMRRVRDAVWSQTGFYG
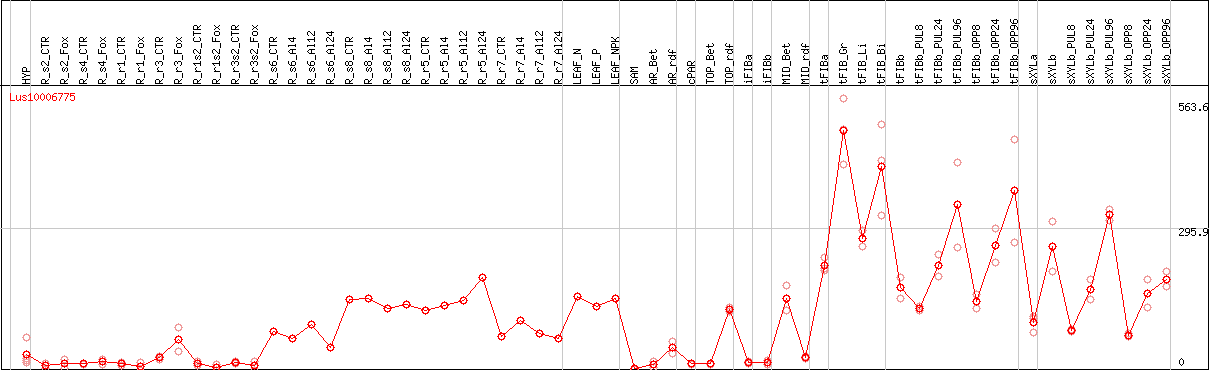
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10006775 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.