Lus10006825 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10006825 pacid=23150770 polypeptide=Lus10006825 locus=Lus10006825.g ID=Lus10006825.BGIv1.0 annot-version=v1.0
ATGCCGAGAAGACCAACAATCCCTACTCCTACAGCTCAAACATGGCCTCAATTTAAGCTATCCATCATTTTCAAGGGTGATATCATGCCGAGAAGACCAA
CAATCCCTACTCCTACAGCTCAAACATGGCCTCAATTTAAGCTATCCATCATTTTCAAGGGTGATATCATGCCGAGAAGACCAACAATCCCTACTCCTAC
AGCTCAAACATGGCCTCAATTTAAGCTATCCATCATTTTCAAGGGTGATATCATGCCGAGAAGACCAACAATCCCTACTCCTACAGCTCAAACATGGCCT
CAATTTAAGCTATCCATCATTTTCAAGGGTGATATCATGCCGAGAAGACCAACAATCCCTACTCCTACAGCTCAAACATGGCCTCAATTTAAGCTATCCA
TCATTTTCAAGGGCTCAGAAAATCGACTGGTTGAGTGGAATTCGAGCACGGATTGCTGTGAGTGGCCTGGTATAAAATGTGATCTCAATGGCCTTGGTCG
TGTTGTTGGACTCAACTTGAGCTTTGAACAGATAATTGTACGAGGTGGAATCGACGATTCTTCGGCTCTGTTTAGTCTTGGATATCTCGAGGAATTGGAT
CTGTCCGACAACTTCTTCAACACAACTATTCCAGGTGCACTTGGCAACCTCATCAATTTGAGGTATCTACACTTATCCGATGCTGGTTTCTATGGTCAAA
TTCCAACTGCCATCTCTCAGCTGACAAGCTTGGTTAGTCTTGATTTGTCATATCAAAATCTAAAGATTGAGAGTACAAATCTGGCGCTACTTGTTCAGAA
TCTCACAAAATTAACCAAGCTTCGACTTGATGGCGTGGACTTATCCACTCAAGGGAAGGAGTGGTTCCATGTCTTATCTTCTTCACTACCAAATCTCAGA
GAGTTGAGCTTGAGCAAATGCAACCTTTCAGCCCCTATGGATGCTCCCTCCCTTTCAAAACTTCAGTCTCTCACGGACCTTTACCTTGATCACAACAATT
TCTCCTCTTTAATCCCCGATATCATCCAAGATTTAAAAAACTTGACAGCCTTGAGAATGTCAAATTGTGGGCTTAGAGGAACATTCCCAAAAAAGATAGT
CCAAAACCTGACTCAACTAGAGTATTTGGATTTCTCACATAACCAATTGGTGGGTAATATACCATATGCTCACTGGGAAAATATTCACAACCTTGTCCAA
GTAGACCTTAGAGACAATTTTTTCAACGGGACCATCCCACCATCTTTATTTTCCATTCCATCTTTGAAAAATATTGATCTCTCCAATAACCAGGTTCAAG
GTCAAATTCCAGAGTTTTCAGCTAATACTTCTTCGACTCTGAATTCTATATTCTTGAATAACAATTTTCTCAACGGAAGCATCCCACCATCTTTATTTTC
CATTCCATCTTTGAAATATATTGATCTCTCCAATAACCAACTAGTAGGTCCCATCCCCCCTTCAATTTTCGAACTTTCATCACTTCATTCCCTCTTCCTT
TCATCTAATAGATTGAGTGGCAAGGTCCAGTTAAGTTGGTTTCAAACACTCCAAAATCTCACTTCCCTTGATCTTTCCTACAACAACCTAACAGTTGATG
CAAATATCAATAAATCCTCTACGAGCTCGCCCGTCTTTCCTCAATTAAGACAATTGCATCTTCCATCGTGCAATCTTAGAACAATTCCCGAGTTGGGCAA
CCAACTAAGTCTTTCCAACCTAGACCTTTCTGACAACAATATCAATGGGGTAATACCTCGTTGGATTATGGATTCAACTACTTTAATATCCCTCAATCTT
TCACACAACAACCTAGTAGGTTTTGAGGTCCAACATTTTGCTTCTACTCCTACATTTTTGGACATGCATGACAACCAACTCCAAGGAAACTTTCCATTCA
TTCCATTTCCATTACCATTATTCGACATATGGTACATAGATTTCTCCCACAATCATCTCAATGGTTCCATTCCAGCCCGAATTTTCAACAATCTTTATTT
GAACTTTCTTTCACTATCAAATAATAGCTTCACAGGAACTATTCCAACAAGTATGTGTGATGCTGTAAGTCTTGACGTCATTAATCTTTCAAACAACCAT
TTGAATGGAACAATCCCATCTTGTCTCATAGAAACCATGAAAGATCTCTCTGTCCTAGATTTGAGTGGGAATATGTTTACTGGCAACATACCAAACACAT
TCCATACTTCATGTCGTTTACAAACTCTAAATTTTCACAACAACCAATTACAAGGAACATTTCCGAAGTCCTTGAAGAACTGCACGAAGTTGGAAGTTTT
AGACATCGGGAACAACAAAATCGCAGGTCCTTTCCCTTGTCTTCAAGGAGAAATCAAATCTACTTTGCGTGTCCTAATTCTTAGGAACAACCTCTTCCAT
GGGAGCATTCAGTGCTCGTATGATGATCAAACTACTTGGCAACAACTCCAAATTGTCGATCTTGCTTCCAACAATTTTTCGGGACCCTTAAATGCCAAAT
ACTTAGATTCATTGAAAGTCATGATGGAAGATGTCTTGACCGCATTCAAGTCAATTGATTTGTCACACAACAATTTTGACGGACCAATACCTCAAGTTAT
CGGAAAATTCAAAGGACTTCAGGTTCTCAACTTGTCTCAGAATGCTTTAATAAGTCAGATCCCATCAACATTCGTCAACTTGTCAAGGTTGGAGTCTCTA
GACCTCTCTGATAACATGCTGAGTGGACATATACCAAAGGAACTAGCGGACTTGACGTTCCTTTCATGCATAAACCTATCAAATAACCATTTAGTTGGCA
GCATCCCAACTGGCAGACAATTTGATACGTTCGAGAACACTTCGTATGGAGGTAATCCAAAACTGTGTGGGAAACCTTTGACTAGATCCTGCACTTCACT
ACTTGATCCGCCGGATGTCATTATTAGACGTGGGAAATCAAATGGAGGAACCAACTGGGACAAGCTTAGCAAAGCATTTGGCTATTTGTTTGGCCTAACT
GTTGTCATGTTGCCAGCTTCGTTATGGCCAAGTTTTCGAGTTTGGTACTGGCCAAAGATCGATTCTCTAATCCTTTGGATGTTCCCAAAGTTATACGTGA
AAGACTGGAATAATCGTCTTAGGAAAAGACAAAGCCAACGACGCACCAAGAAAATTGCGACAAGCCGGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10006825 pacid=23150770 polypeptide=Lus10006825 locus=Lus10006825.g ID=Lus10006825.BGIv1.0 annot-version=v1.0
MPRRPTIPTPTAQTWPQFKLSIIFKGDIMPRRPTIPTPTAQTWPQFKLSIIFKGDIMPRRPTIPTPTAQTWPQFKLSIIFKGDIMPRRPTIPTPTAQTWP
QFKLSIIFKGDIMPRRPTIPTPTAQTWPQFKLSIIFKGSENRLVEWNSSTDCCEWPGIKCDLNGLGRVVGLNLSFEQIIVRGGIDDSSALFSLGYLEELD
LSDNFFNTTIPGALGNLINLRYLHLSDAGFYGQIPTAISQLTSLVSLDLSYQNLKIESTNLALLVQNLTKLTKLRLDGVDLSTQGKEWFHVLSSSLPNLR
ELSLSKCNLSAPMDAPSLSKLQSLTDLYLDHNNFSSLIPDIIQDLKNLTALRMSNCGLRGTFPKKIVQNLTQLEYLDFSHNQLVGNIPYAHWENIHNLVQ
VDLRDNFFNGTIPPSLFSIPSLKNIDLSNNQVQGQIPEFSANTSSTLNSIFLNNNFLNGSIPPSLFSIPSLKYIDLSNNQLVGPIPPSIFELSSLHSLFL
SSNRLSGKVQLSWFQTLQNLTSLDLSYNNLTVDANINKSSTSSPVFPQLRQLHLPSCNLRTIPELGNQLSLSNLDLSDNNINGVIPRWIMDSTTLISLNL
SHNNLVGFEVQHFASTPTFLDMHDNQLQGNFPFIPFPLPLFDIWYIDFSHNHLNGSIPARIFNNLYLNFLSLSNNSFTGTIPTSMCDAVSLDVINLSNNH
LNGTIPSCLIETMKDLSVLDLSGNMFTGNIPNTFHTSCRLQTLNFHNNQLQGTFPKSLKNCTKLEVLDIGNNKIAGPFPCLQGEIKSTLRVLILRNNLFH
GSIQCSYDDQTTWQQLQIVDLASNNFSGPLNAKYLDSLKVMMEDVLTAFKSIDLSHNNFDGPIPQVIGKFKGLQVLNLSQNALISQIPSTFVNLSRLESL
DLSDNMLSGHIPKELADLTFLSCINLSNNHLVGSIPTGRQFDTFENTSYGGNPKLCGKPLTRSCTSLLDPPDVIIRRGKSNGGTNWDKLSKAFGYLFGLT
VVMLPASLWPSFRVWYWPKIDSLILWMFPKLYVKDWNNRLRKRQSQRRTKKIATSR
|
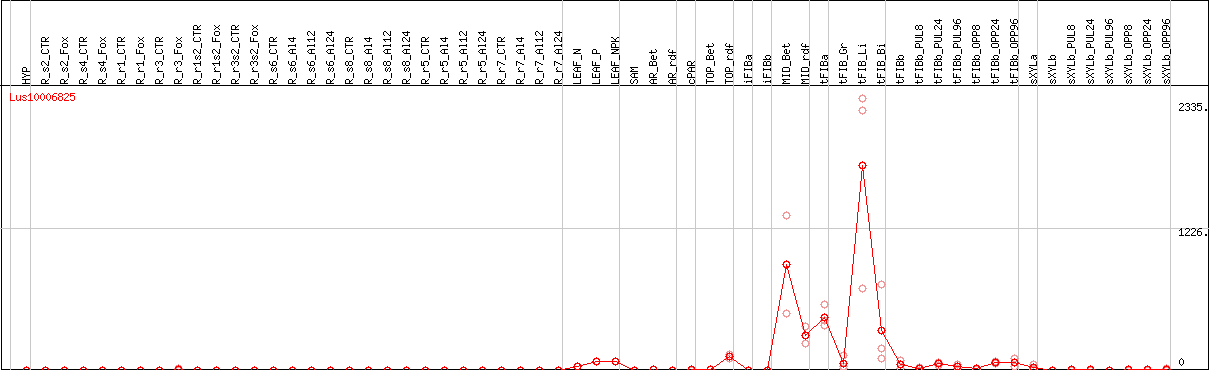
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10006825 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
