External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G57685 67 / 1e-14
LSB1, ATGDU3
LESS SUSCEPTIBLE TO BSCTV 1, ARABIDOPSIS THALIANA GLUTAMINE DUMPER 3, glutamine dumper 3 (.1)
AT4G25760 59 / 2e-11
ATGDU2
glutamine dumper 2 (.1)
AT5G24920 53 / 2e-09
ATGDU5
glutamine dumper 5 (.1)
AT2G24762 51 / 1e-08
ATGDU4
glutamine dumper 4 (.1)
AT4G31730 50 / 3e-08
ATGDU1, GDU1
glutamine dumper 1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10008910
121 / 4e-35
AT5G57685 125 / 6e-37
LESS SUSCEPTIBLE TO BSCTV 1, ARABIDOPSIS THALIANA GLUTAMINE DUMPER 3, glutamine dumper 3 (.1)
Lus10006986
117 / 1e-33
AT5G57685 124 / 2e-36
LESS SUSCEPTIBLE TO BSCTV 1, ARABIDOPSIS THALIANA GLUTAMINE DUMPER 3, glutamine dumper 3 (.1)
Lus10008909
84 / 2e-21
AT5G57685 87 / 2e-23
LESS SUSCEPTIBLE TO BSCTV 1, ARABIDOPSIS THALIANA GLUTAMINE DUMPER 3, glutamine dumper 3 (.1)
Lus10019985
84 / 8e-21
AT5G57685 123 / 3e-36
LESS SUSCEPTIBLE TO BSCTV 1, ARABIDOPSIS THALIANA GLUTAMINE DUMPER 3, glutamine dumper 3 (.1)
Lus10019986
83 / 2e-20
AT5G57685 125 / 7e-37
LESS SUSCEPTIBLE TO BSCTV 1, ARABIDOPSIS THALIANA GLUTAMINE DUMPER 3, glutamine dumper 3 (.1)
Lus10015513
80 / 4e-19
AT5G57685 124 / 2e-36
LESS SUSCEPTIBLE TO BSCTV 1, ARABIDOPSIS THALIANA GLUTAMINE DUMPER 3, glutamine dumper 3 (.1)
Lus10026911
59 / 5e-11
AT4G31730 125 / 6e-37
glutamine dumper 1 (.1)
Lus10020107
56 / 6e-10
AT4G31730 122 / 1e-35
glutamine dumper 1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G173901
82 / 4e-20
AT5G57685 89 / 8e-23
LESS SUSCEPTIBLE TO BSCTV 1, ARABIDOPSIS THALIANA GLUTAMINE DUMPER 3, glutamine dumper 3 (.1)
Potri.006G267900
71 / 1e-15
AT5G57685 83 / 2e-20
LESS SUSCEPTIBLE TO BSCTV 1, ARABIDOPSIS THALIANA GLUTAMINE DUMPER 3, glutamine dumper 3 (.1)
Potri.018G013600
60 / 1e-11
AT4G25760 79 / 2e-19
glutamine dumper 2 (.1)
Potri.004G108800
40 / 6e-05
AT5G24920 57 / 2e-11
glutamine dumper 5 (.1)
PFAM info
Representative CDS sequence
>Lus10006987 pacid=23165411 polypeptide=Lus10006987 locus=Lus10006987.g ID=Lus10006987.BGIv1.0 annot-version=v1.0
ATGGAAGCCACAAGTCAACCCGTCGGAGTAAATCTTCCGGCGGCCACGTCAGCAGCGGAGCCGTCGCCGTGGCATTCCCCTGTTCCGTACTTGTTCGGCG
GATTAGCAGCCATGTTGGGGCTGATTGCCTTCGCGCTGCTGATCCTCGCTTGCTCCTACTGGAAGCTCTCCGGACACCTCGAGAACGGCGAAAACGATGA
TCAGAGGGATCTAGAAGCAGGGGAAGGCGGAGAAACAGAGGAGAAAGGGAAGAAGAAACAATACGAGGAGAAGGTTTTGGTTATAATGGCTGGACAAGTG
AAGCCGACTTTTCTTGCGACTCCGGTGACGAGCAGAAGAACGTCGTCGTTTGGGGACGACGGTGGGAGTGAGAAGACTTGTCGATGTGGGGAGAAATTGG
ACGGCGAGAAACAGGGGAATAGTGAGGAAGAAGAAGAGAATGGAAACAGGGGAGGTTTGAGAAGTGATGATACGACGACGCATTAG
AA sequence
>Lus10006987 pacid=23165411 polypeptide=Lus10006987 locus=Lus10006987.g ID=Lus10006987.BGIv1.0 annot-version=v1.0
MEATSQPVGVNLPAATSAAEPSPWHSPVPYLFGGLAAMLGLIAFALLILACSYWKLSGHLENGENDDQRDLEAGEGGETEEKGKKKQYEEKVLVIMAGQV
KPTFLATPVTSRRTSSFGDDGGSEKTCRCGEKLDGEKQGNSEEEEENGNRGGLRSDDTTTH
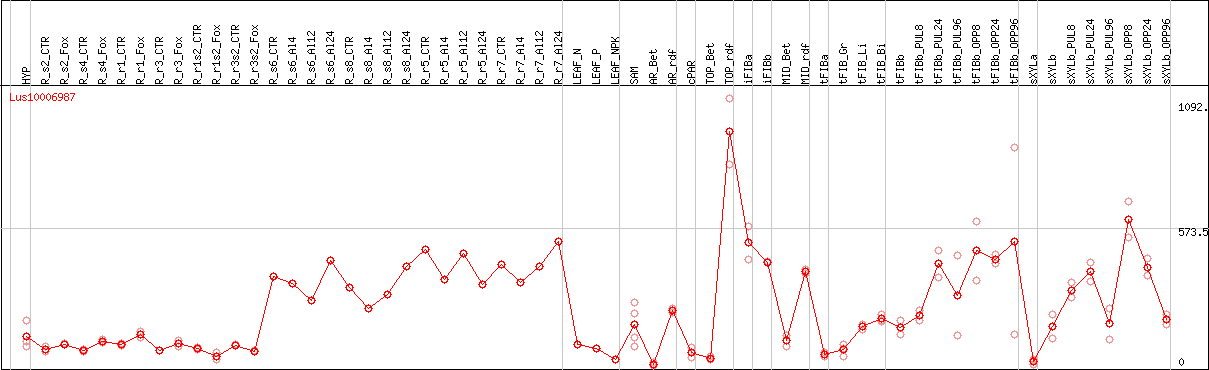
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10006987 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.