External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G57230 226 / 5e-75
MADS
AGL16
AGAMOUS-like 16 (.1.2)
AT4G37940 224 / 2e-74
MADS
AGL21
AGAMOUS-like 21 (.1)
AT2G22630 214 / 2e-70
MADS
AGL17
AGAMOUS-like 17 (.1)
AT2G14210 199 / 3e-64
MADS
AGL44, ANR1
ARABIDOPSIS NITRATE REGULATED 1, AGAMOUS-like 44 (.1)
AT1G26310 152 / 7e-46
MADS
CAL1, AGL10, CAL
CAULIFLOWER, AGAMOUS-like 10, K-box region and MADS-box transcription factor family protein (.1)
AT1G69120 145 / 5e-43
MADS
AGL7, AP1
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
AT4G09960 144 / 1e-42
MADS
AGL11, STK
SEEDSTICK, AGAMOUS-like 11, K-box region and MADS-box transcription factor family protein (.1.2.3.4)
AT2G42830 144 / 1e-42
MADS
AGL5, SHP2
SHATTERPROOF 2, AGAMOUS-like 5, K-box region and MADS-box transcription factor family protein (.1.2)
AT2G45650 144 / 2e-42
MADS
AGL6
AGAMOUS-like 6 (.1)
AT5G60910 143 / 2e-42
MADS
FUL, AGL8
FRUITFULL, AGAMOUS-like 8 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10020477
261 / 7e-90
AT4G37940 195 / 4e-64
AGAMOUS-like 21 (.1)
Lus10017871
144 / 2e-43
AT1G69120 264 / 2e-90
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Lus10021140
144 / 1e-42
AT5G60910 291 / 1e-99
FRUITFULL, AGAMOUS-like 8 (.1.2)
Lus10007983
144 / 2e-42
AT5G60910 298 / 3e-102
FRUITFULL, AGAMOUS-like 8 (.1.2)
Lus10004638
142 / 4e-42
AT3G02310 246 / 2e-82
SEPALLATA 2, AGAMOUS-like 4, K-box region and MADS-box transcription factor family protein (.1)
Lus10034662
142 / 4e-42
AT5G60910 284 / 3e-97
FRUITFULL, AGAMOUS-like 8 (.1.2)
Lus10004637
140 / 2e-41
AT1G69120 318 / 2e-111
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Lus10026679
140 / 2e-41
AT1G69120 338 / 1e-118
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Lus10009481
141 / 4e-41
AT4G09960 274 / 1e-92
SEEDSTICK, AGAMOUS-like 11, K-box region and MADS-box transcription factor family protein (.1.2.3.4)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF01486
K-box
K-box region
PF00319
SRF-TF
SRF-type transcription factor (DNA-binding and dimerisation domain)
Representative CDS sequence
>Lus10007100 pacid=23153246 polypeptide=Lus10007100 locus=Lus10007100.g ID=Lus10007100.BGIv1.0 annot-version=v1.0
ATGGGAAGGGGGAAGACAGTGATAAGGAGGATCGATGACAGAACGAACAGGCAGGTGACTTTCTCAAAGAGAAGAACCGGTTTGATTAAGAAAGCAAAGG
AGCTATCCATCCTTTGTGATGCTCAAATTGCTCTCATTGTTTTCTCCACTACCAACAAGCTCTATCAATTTTCCAGTACCAGTATGATGTCGATCATTGA
TCGTTACACCAGACAGAAAGAAGCTCACAGTCAGCCAGTCAATCCTGCATCAGAGATCAAGTTTTGGCAGAAGGAGGCAGCAAAACTCAAGAGGGAACTG
CAGTCCTTAAATGAATATCACAGACAATTAATGGGAGAAGAAGTATCAGGCTTGAGTGTCAGAGATCTGCAAAATTTGGAAACCCAGCTTGAGATTAGTT
TGAAAGCAATCCGCCTCAAGAAGCACAATACGATGACAGATCAAATAACAGCCTTGACCGCAAAGGGAAATCAATGTTACGAAGAAAATATACAACTTCA
CAAGCAGTTCAATTTCTTCGCCAATGAAAAATCAGCACTCGGTATAAAGGTATATGAAGAAAGGAATGCAGAAGAAACCGATGGAATGGCGAAGGCTGCG
GTCAGTCTGCAATTGACCCAACCTGAACCCCTGAAATTAGGGTAG
AA sequence
>Lus10007100 pacid=23153246 polypeptide=Lus10007100 locus=Lus10007100.g ID=Lus10007100.BGIv1.0 annot-version=v1.0
MGRGKTVIRRIDDRTNRQVTFSKRRTGLIKKAKELSILCDAQIALIVFSTTNKLYQFSSTSMMSIIDRYTRQKEAHSQPVNPASEIKFWQKEAAKLKREL
QSLNEYHRQLMGEEVSGLSVRDLQNLETQLEISLKAIRLKKHNTMTDQITALTAKGNQCYEENIQLHKQFNFFANEKSALGIKVYEERNAEETDGMAKAA
VSLQLTQPEPLKLG
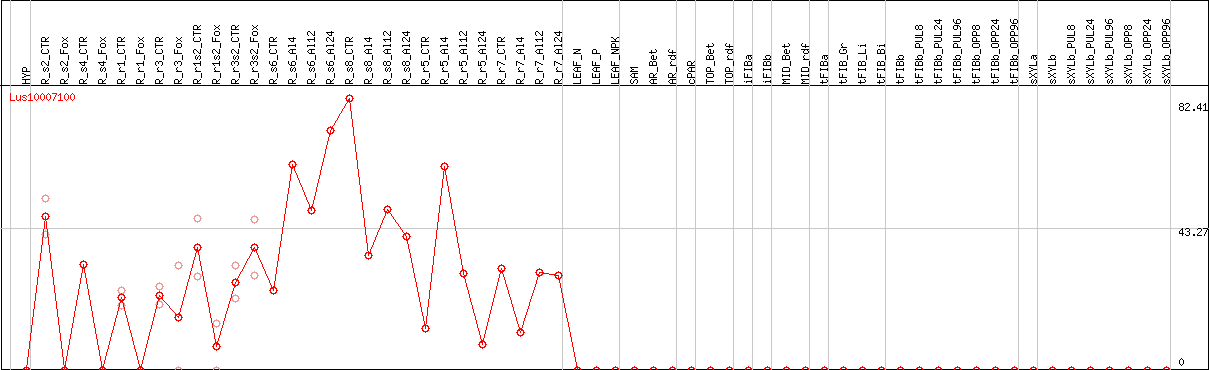
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10007100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.