External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G24140 124 / 2e-34
bHLH
bHLH097, FMA
FAMA, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT5G53210 71 / 6e-15
bHLH
SPCH, bHLH098
SPEECHLESS, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT3G06120 60 / 2e-11
bHLH
MUTE, bHLH045
MUTE, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G72210 59 / 2e-10
bHLH
bHLH096
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G22490 55 / 2e-09
bHLH
bHLH094
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT3G61950 45 / 5e-06
bHLH
bHLH067
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
AT5G65320 45 / 1e-05
bHLH
bHLH099
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT2G46810 44 / 2e-05
bHLH
bHLH070
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G32640 42 / 8e-05
bHLH
JIN1, JAI1, ZBF1, RD22BP1, ATMYC2
JASMONATE INSENSITIVE 1, Basic helix-loop-helix (bHLH) DNA-binding family protein (.1)
AT4G01460 41 / 0.0001
bHLH
bHLH057
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G314400
127 / 1e-35
AT3G24140 399 / 2e-137
FAMA, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.017G054500
118 / 6e-32
AT3G24140 344 / 8e-116
FAMA, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.002G101900
71 / 6e-15
AT1G72210 256 / 4e-84
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.008G202900
69 / 1e-14
AT3G06120 250 / 5e-85
MUTE, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.013G025900
67 / 2e-13
AT1G72210 243 / 2e-78
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.012G031800
67 / 2e-13
AT5G53210 303 / 2e-101
SPEECHLESS, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.005G039800
66 / 3e-13
AT1G72210 231 / 5e-74
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.015G022300
64 / 3e-12
AT5G53210 152 / 1e-42
SPEECHLESS, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.014G106300
63 / 5e-12
AT2G46810 247 / 9e-79
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.005G071100
62 / 1e-11
AT1G72210 177 / 1e-53
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
PFAM info
Representative CDS sequence
>Lus10007138 pacid=23139765 polypeptide=Lus10007138 locus=Lus10007138.g ID=Lus10007138.BGIv1.0 annot-version=v1.0
ATGGCTGAGAACAAATCGTGTCTTGCTGACGTGGAGGTTAAGCTGTTAGGGTTTGATGCAATGATTAAGGTTTTGTCGAGGAGGAGACCTGGTCAGCTGA
TTAAGGCCATTGCTGCTTTGGAGGATTTGCAGCTTACAATCCTGCATACCAACATCACTATCATTGAGCAAACTGTGCTTTATTCCTTTAACGTCAAGGC
TCATCGTTTCTCACACAGAAAACCGTTTACATCTCAGATAAATGAAGTTAGCCTTTGGAGCATTGAGTTGGTGAATGAGGAGCTATCGAATGTGGACGGT
TGTCATTGTGTGATTGAAGAGACACATGGAATCCTTTGTACTTGTAGGCTTGAATCTATGGAGCGCAATTGGGTTAAGGTGAAACCATACCACCTCCACC
ACTTTTGGCGATCATTGGAGTACCGTCTCCCTGTAGACAGTGAGAAGTGGGAAGAGCATGACGCGATAGACCGCGACATCCTCAACGGGAATGGTACTGA
TGATTTGTGA
AA sequence
>Lus10007138 pacid=23139765 polypeptide=Lus10007138 locus=Lus10007138.g ID=Lus10007138.BGIv1.0 annot-version=v1.0
MAENKSCLADVEVKLLGFDAMIKVLSRRRPGQLIKAIAALEDLQLTILHTNITIIEQTVLYSFNVKAHRFSHRKPFTSQINEVSLWSIELVNEELSNVDG
CHCVIEETHGILCTCRLESMERNWVKVKPYHLHHFWRSLEYRLPVDSEKWEEHDAIDRDILNGNGTDDL
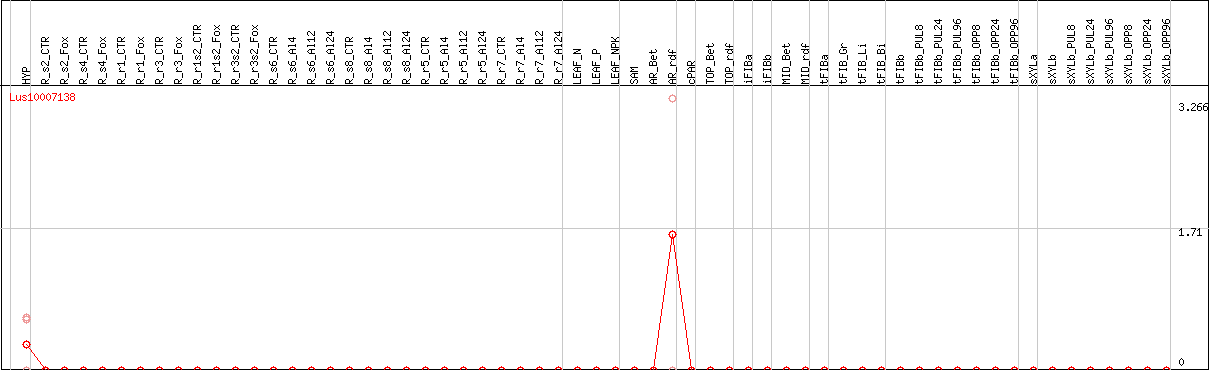
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10007138 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.