External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G08150 205 / 8e-66
HD
BP1, KNAT1
BREVIPEDICELLUS 1, BREVIPEDICELLUS, KNOTTED-like from Arabidopsis thaliana (.1)
AT1G62360 153 / 1e-45
HD
WAM1, WAM, SHL, BUM1, STM
WALDMEISTER 1, WALDMEISTER, SHOOT MERISTEMLESS, SHOOTLESS, BUMBERSHOOT 1, BUMBERSHOOT, KNOX/ELK homeobox transcription factor (.1)
AT1G23380 129 / 5e-37
HD
KNAT6S, KNAT6L, KNAT6
KNOTTED1-like homeobox gene 6 (.1.2)
AT1G70510 122 / 2e-34
HD
ATK1, KNAT2
ARABIDOPSIS THALIANA KN 1, KNOTTED-like from Arabidopsis thaliana 2 (.1)
AT1G62990 72 / 9e-16
HD
IXR11, KNAT7
KNOTTED-like homeobox of Arabidopsis thaliana 7 (.1)
AT5G11060 72 / 2e-15
HD
KNAT4
KNOTTED1-like homeobox gene 4 (.1)
AT5G25220 71 / 5e-15
HD
KNAT3
KNOTTED1-like homeobox gene 3 (.1.2)
AT4G32040 71 / 8e-15
HD
KNAT5
KNOTTED1-like homeobox gene 5 (.1)
AT1G75430 59 / 4e-11
HD
BLH11
BEL1-like homeodomain 11 (.1)
AT1G19700 59 / 9e-11
HD
BEL10, BLH10
BEL1-like homeodomain 10 (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10028244
240 / 2e-80
AT4G08150 370 / 1e-127
BREVIPEDICELLUS 1, BREVIPEDICELLUS, KNOTTED-like from Arabidopsis thaliana (.1)
Lus10030003
147 / 2e-43
AT1G62360 315 / 5e-105
WALDMEISTER 1, WALDMEISTER, SHOOT MERISTEMLESS, SHOOTLESS, BUMBERSHOOT 1, BUMBERSHOOT, KNOX/ELK homeobox transcription factor (.1)
Lus10013037
135 / 2e-39
AT1G23380 417 / 4e-147
KNOTTED1-like homeobox gene 6 (.1.2)
Lus10029125
134 / 6e-39
AT1G23380 417 / 3e-147
KNOTTED1-like homeobox gene 6 (.1.2)
Lus10042102
109 / 4e-31
AT1G23380 155 / 3e-47
KNOTTED1-like homeobox gene 6 (.1.2)
Lus10001238
110 / 3e-29
AT1G23380 250 / 1e-80
KNOTTED1-like homeobox gene 6 (.1.2)
Lus10017824
72 / 4e-16
AT1G62990 269 / 9e-92
KNOTTED-like homeobox of Arabidopsis thaliana 7 (.1)
Lus10003183
72 / 8e-16
AT5G25220 381 / 2e-133
KNOTTED1-like homeobox gene 3 (.1.2)
Lus10001941
73 / 1e-15
AT5G25220 439 / 2e-153
KNOTTED1-like homeobox gene 3 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G113300
205 / 4e-66
AT4G08150 421 / 7e-147
BREVIPEDICELLUS 1, BREVIPEDICELLUS, KNOTTED-like from Arabidopsis thaliana (.1)
Potri.006G270401
153 / 1e-45
AT1G62360 399 / 3e-138
WALDMEISTER 1, WALDMEISTER, SHOOT MERISTEMLESS, SHOOTLESS, BUMBERSHOOT 1, BUMBERSHOOT, KNOX/ELK homeobox transcription factor (.1)
Potri.011G011101
153 / 1e-45
AT1G62360 400 / 7e-139
WALDMEISTER 1, WALDMEISTER, SHOOT MERISTEMLESS, SHOOTLESS, BUMBERSHOOT 1, BUMBERSHOOT, KNOX/ELK homeobox transcription factor (.1)
Potri.004G004700
151 / 4e-45
AT1G62360 385 / 3e-133
WALDMEISTER 1, WALDMEISTER, SHOOT MERISTEMLESS, SHOOTLESS, BUMBERSHOOT 1, BUMBERSHOOT, KNOX/ELK homeobox transcription factor (.1)
Potri.012G087100
140 / 3e-41
AT1G23380 323 / 1e-109
KNOTTED1-like homeobox gene 6 (.1.2)
Potri.015G079100
138 / 2e-40
AT1G23380 309 / 3e-104
KNOTTED1-like homeobox gene 6 (.1.2)
Potri.008G188700
137 / 2e-40
AT1G23380 390 / 1e-136
KNOTTED1-like homeobox gene 6 (.1.2)
Potri.010G043500
136 / 6e-40
AT1G23380 371 / 3e-129
KNOTTED1-like homeobox gene 6 (.1.2)
Potri.005G014200
119 / 3e-33
AT1G23380 250 / 1e-81
KNOTTED1-like homeobox gene 6 (.1.2)
Potri.013G008600
113 / 8e-31
AT1G23380 250 / 1e-81
KNOTTED1-like homeobox gene 6 (.1.2)
PFAM info
Representative CDS sequence
>Lus10007245 pacid=23182158 polypeptide=Lus10007245 locus=Lus10007245.g ID=Lus10007245.BGIv1.0 annot-version=v1.0
ATGGGAGGTTCGTCGGAAGAGGAGCAGGAGAACAGTGGAGGTGAAACAGAACTCCCTGAGATCGATCCAAGAGCGGAAGACAGGGAGCTGAAGAACCATC
TGTTGAGAAAGTACAGTGGATACCTAAGCAGCTTGAAGCAAGAGCTGTCCAAGAAGAAGAAGAAAGGGAAGCTCCCTAAAGAAGCCAGGCAGAAGCTACT
CAGTTGGTGGGAACTTCACTACAAGTGGCCTTACCCTTCGGAGTCGGAGAAAGTGGCATTGGCAGAGACAACAGGGCTAGACCAGAAGCAGATAAACAAC
TGGTTCATAAACCAGAGGAAAAGGCACTGGAAACCATCAGAAGACATGCAGTTCATGGTGATGGATGGACTTCATCACCACCCAGCTCAGAACCCAGCTG
CTGCTCTCTACATGGATGCTCATCATTACATGCCCGATGGTCCATACCGTTTAGGCCCTTAA
AA sequence
>Lus10007245 pacid=23182158 polypeptide=Lus10007245 locus=Lus10007245.g ID=Lus10007245.BGIv1.0 annot-version=v1.0
MGGSSEEEQENSGGETELPEIDPRAEDRELKNHLLRKYSGYLSSLKQELSKKKKKGKLPKEARQKLLSWWELHYKWPYPSESEKVALAETTGLDQKQINN
WFINQRKRHWKPSEDMQFMVMDGLHHHPAQNPAAALYMDAHHYMPDGPYRLGP
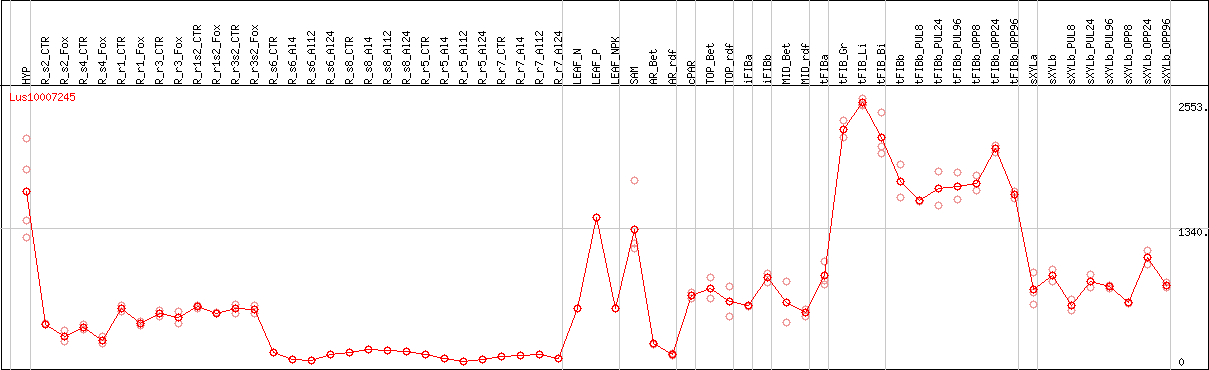
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10007245 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.