External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G42610 236 / 5e-80
LSH10
LIGHT SENSITIVE HYPOCOTYLS 10, Protein of unknown function (DUF640) (.1), Protein of unknown function (DUF640) (.2)
AT4G18610 213 / 1e-70
LSH9
LIGHT SENSITIVE HYPOCOTYLS 9, Protein of unknown function (DUF640) (.1)
AT1G78815 206 / 8e-68
LSH7
LIGHT SENSITIVE HYPOCOTYLS 7, Protein of unknown function (DUF640) (.1)
AT1G07090 204 / 4e-67
LSH6
LIGHT SENSITIVE HYPOCOTYLS 6, Protein of unknown function (DUF640) (.1)
AT5G58500 201 / 6e-66
LSH5
LIGHT SENSITIVE HYPOCOTYLS 5, Protein of unknown function (DUF640) (.1)
AT3G23290 190 / 1e-61
LSH4
LIGHT SENSITIVE HYPOCOTYLS 4, Protein of unknown function (DUF640) (.2)
AT1G16910 186 / 2e-60
LSH8
LIGHT SENSITIVE HYPOCOTYLS 8, Protein of unknown function (DUF640) (.1)
AT2G31160 187 / 3e-60
OBO1, LSH3
ORGAN BOUNDARY 1, LIGHT SENSITIVE HYPOCOTYLS 3, Protein of unknown function (DUF640) (.1)
AT3G04510 182 / 2e-58
LSH2
LIGHT SENSITIVE HYPOCOTYLS 2, Protein of unknown function (DUF640) (.1)
AT5G28490 180 / 1e-57
OBO2, LSH1
ORGAN BOUNDARY 2, LIGHT-DEPENDENT SHORT HYPOCOTYLS 1, Protein of unknown function (DUF640) (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10029219
328 / 4e-116
AT2G42610 238 / 6e-81
LIGHT SENSITIVE HYPOCOTYLS 10, Protein of unknown function (DUF640) (.1), Protein of unknown function (DUF640) (.2)
Lus10029838
236 / 1e-79
AT2G42610 261 / 3e-90
LIGHT SENSITIVE HYPOCOTYLS 10, Protein of unknown function (DUF640) (.1), Protein of unknown function (DUF640) (.2)
Lus10020700
231 / 1e-77
AT2G42610 258 / 1e-88
LIGHT SENSITIVE HYPOCOTYLS 10, Protein of unknown function (DUF640) (.1), Protein of unknown function (DUF640) (.2)
Lus10042565
200 / 1e-65
AT5G28490 233 / 1e-78
ORGAN BOUNDARY 2, LIGHT-DEPENDENT SHORT HYPOCOTYLS 1, Protein of unknown function (DUF640) (.1)
Lus10022023
199 / 2e-65
AT1G07090 226 / 3e-76
LIGHT SENSITIVE HYPOCOTYLS 6, Protein of unknown function (DUF640) (.1)
Lus10023383
182 / 3e-58
AT1G07090 246 / 2e-83
LIGHT SENSITIVE HYPOCOTYLS 6, Protein of unknown function (DUF640) (.1)
Lus10038424
181 / 1e-57
AT1G07090 247 / 9e-84
LIGHT SENSITIVE HYPOCOTYLS 6, Protein of unknown function (DUF640) (.1)
Lus10024901
181 / 2e-57
AT2G31160 249 / 3e-84
ORGAN BOUNDARY 1, LIGHT SENSITIVE HYPOCOTYLS 3, Protein of unknown function (DUF640) (.1)
Lus10022918
181 / 2e-57
AT2G31160 249 / 4e-84
ORGAN BOUNDARY 1, LIGHT SENSITIVE HYPOCOTYLS 3, Protein of unknown function (DUF640) (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G057300
243 / 1e-82
AT2G42610 241 / 2e-82
LIGHT SENSITIVE HYPOCOTYLS 10, Protein of unknown function (DUF640) (.1), Protein of unknown function (DUF640) (.2)
Potri.011G066400
238 / 8e-81
AT2G42610 238 / 5e-81
LIGHT SENSITIVE HYPOCOTYLS 10, Protein of unknown function (DUF640) (.1), Protein of unknown function (DUF640) (.2)
Potri.014G142400
231 / 4e-78
AT2G42610 259 / 1e-89
LIGHT SENSITIVE HYPOCOTYLS 10, Protein of unknown function (DUF640) (.1), Protein of unknown function (DUF640) (.2)
Potri.001G390900
229 / 3e-77
AT1G78815 242 / 2e-82
LIGHT SENSITIVE HYPOCOTYLS 7, Protein of unknown function (DUF640) (.1)
Potri.011G109900
228 / 1e-76
AT2G42610 238 / 3e-81
LIGHT SENSITIVE HYPOCOTYLS 10, Protein of unknown function (DUF640) (.1), Protein of unknown function (DUF640) (.2)
Potri.002G214200
225 / 9e-76
AT2G42610 240 / 5e-82
LIGHT SENSITIVE HYPOCOTYLS 10, Protein of unknown function (DUF640) (.1), Protein of unknown function (DUF640) (.2)
Potri.009G076100
213 / 3e-70
AT1G07090 259 / 2e-88
LIGHT SENSITIVE HYPOCOTYLS 6, Protein of unknown function (DUF640) (.1)
Potri.001G281000
207 / 6e-68
AT1G07090 251 / 4e-85
LIGHT SENSITIVE HYPOCOTYLS 6, Protein of unknown function (DUF640) (.1)
Potri.009G157600
194 / 2e-63
AT5G28490 228 / 7e-77
ORGAN BOUNDARY 2, LIGHT-DEPENDENT SHORT HYPOCOTYLS 1, Protein of unknown function (DUF640) (.1)
Potri.010G070700
188 / 1e-60
AT2G31160 226 / 3e-75
ORGAN BOUNDARY 1, LIGHT SENSITIVE HYPOCOTYLS 3, Protein of unknown function (DUF640) (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04852
DUF640
Protein of unknown function (DUF640)
Representative CDS sequence
>Lus10007272 pacid=23142459 polypeptide=Lus10007272 locus=Lus10007272.g ID=Lus10007272.BGIv1.0 annot-version=v1.0
ATGTCATCATCGGAGCAAAATTTAACGATAGGAGTGGAGGGATCCTCCTCCACTTCCAGAACCAACCACCACCACCATCACAACCCGCCGCCGCTGAGCC
GGTACGAGTCGCAGAAGAGGCGGGACTGGAACACGTTCGGGCAGTACTTGAAGAACCAGAGGCCGCCCGTTTCAGTTTCCCACTGCGGGTCGGTCCACGT
GCTGGACTTCCTCCGATACTTAGACCAGTTCGGGAAGACCAAAGTCCACCTAAACGGCTGCGCCTTCTTCGGGCAGCCGGACCCACCTGCTCCGTGCACC
TGCCCGCTCAGGCAGGCGTGGGGAAGCCTGGACGCGCTCATAGGCAGGCTGAAGGCTGCCTATGAGGAGCACGGCGGGTCTCCCGAGACCAACCCATTCG
GAAACGGGGCGATCCGGGTTTACCTGAGGGAAGTGAGGGAGTGCCAGGCCAAGGCTAGAGGGATTCCGTACAAGAAAAAGAAGAAGAAGAAGGTTATTGG
CCATCATAACAAGGTGGCGGCCGTTGGTAGGGATAATTGTCGTCTTGATGGCGGCGGTGGTGGTAATAAGTCGGCGGCGGCGGCTGCTGCTGCTCCTATG
CCGCAGTGA
AA sequence
>Lus10007272 pacid=23142459 polypeptide=Lus10007272 locus=Lus10007272.g ID=Lus10007272.BGIv1.0 annot-version=v1.0
MSSSEQNLTIGVEGSSSTSRTNHHHHHNPPPLSRYESQKRRDWNTFGQYLKNQRPPVSVSHCGSVHVLDFLRYLDQFGKTKVHLNGCAFFGQPDPPAPCT
CPLRQAWGSLDALIGRLKAAYEEHGGSPETNPFGNGAIRVYLREVRECQAKARGIPYKKKKKKKVIGHHNKVAAVGRDNCRLDGGGGGNKSAAAAAAAPM
PQ
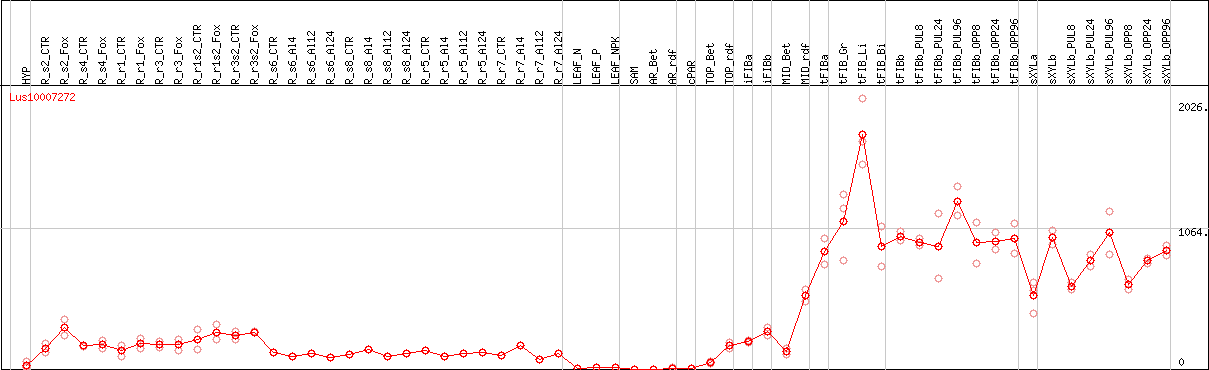
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10007272 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.