External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G45820 575 / 0
PKS18, CIPK20, SnRK3.6
SNF1-RELATED PROTEIN KINASE 3.6, PROTEIN KINASE 18, CBL-interacting protein kinase 20 (.1)
AT5G58380 496 / 8e-174
PKS2, CIPK10, SnRK3.8, SIP1
SNF1-RELATED PROTEIN KINASE 3.8, CBL-INTERACTING PROTEIN KINASE 10, SOS3-interacting protein 1 (.1)
AT4G18700 494 / 4e-173
ATWL4, CIPK12, SnRK3.9
SNF1-RELATED PROTEIN KINASE 3.9, WPL4-LIKE 4, CBL-interacting protein kinase 12 (.1)
AT4G30960 477 / 5e-167
CIPK6, SIP3, SnRK3.14, ATCIPK6
SNF1-RELATED PROTEIN KINASE 3.14, CBL-INTERACTING PROTEIN KINASE 6, SOS3-interacting protein 3 (.1)
AT5G45810 473 / 9e-165
CIPK19, SnRK3.5
SNF1-RELATED PROTEIN KINASE 3.5, CBL-interacting protein kinase 19 (.1)
AT5G07070 466 / 3e-162
CIPK2, SnRK3.2
SNF1-RELATED PROTEIN KINASE 3.2, CBL-interacting protein kinase 2 (.1)
AT5G01810 463 / 9e-162
ATPK10, SnRK3.1, SIP2, PKS3, CIPK15
SNF1-RELATED PROTEIN KINASE 3.1, SOS3-INTERACTING PROTEIN 2, PROTEIN KINASE 10, CBL-interacting protein kinase 15 (.1.2.3)
AT5G10930 461 / 9e-161
CIPK5, SnRK3.24
SNF1-RELATED PROTEIN KINASE 3.24, CBL-interacting protein kinase 5 (.1)
AT5G25110 461 / 7e-160
CIPK25, SnRK3.25
SNF1-RELATED PROTEIN KINASE 3.25, CBL-interacting protein kinase 25 (.1)
AT1G30270 456 / 7e-158
PKS17, ATCIPK23, SnRK3.23, LKS1, CIPK23
SNF1-RELATED PROTEIN KINASE 3.23, SOS2-like protein kinase 17, LOW-K+-SENSITIVE 1, CBL-interacting protein kinase 23 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10042229
521 / 0
AT5G58380 676 / 0.0
SNF1-RELATED PROTEIN KINASE 3.8, CBL-INTERACTING PROTEIN KINASE 10, SOS3-interacting protein 1 (.1)
Lus10022749
517 / 0
AT5G58380 598 / 0.0
SNF1-RELATED PROTEIN KINASE 3.8, CBL-INTERACTING PROTEIN KINASE 10, SOS3-interacting protein 1 (.1)
Lus10014163
514 / 0
AT5G58380 592 / 0.0
SNF1-RELATED PROTEIN KINASE 3.8, CBL-INTERACTING PROTEIN KINASE 10, SOS3-interacting protein 1 (.1)
Lus10021852
502 / 8e-177
AT4G30960 658 / 0.0
SNF1-RELATED PROTEIN KINASE 3.14, CBL-INTERACTING PROTEIN KINASE 6, SOS3-interacting protein 3 (.1)
Lus10036280
501 / 3e-176
AT4G30960 662 / 0.0
SNF1-RELATED PROTEIN KINASE 3.14, CBL-INTERACTING PROTEIN KINASE 6, SOS3-interacting protein 3 (.1)
Lus10024448
476 / 5e-166
AT5G25110 654 / 0.0
SNF1-RELATED PROTEIN KINASE 3.25, CBL-interacting protein kinase 25 (.1)
Lus10007448
472 / 2e-164
AT5G25110 650 / 0.0
SNF1-RELATED PROTEIN KINASE 3.25, CBL-interacting protein kinase 25 (.1)
Lus10015275
473 / 4e-164
AT4G18700 736 / 0.0
SNF1-RELATED PROTEIN KINASE 3.9, WPL4-LIKE 4, CBL-interacting protein kinase 12 (.1)
Lus10028115
462 / 4e-161
AT1G30270 793 / 0.0
SNF1-RELATED PROTEIN KINASE 3.23, SOS2-like protein kinase 17, LOW-K+-SENSITIVE 1, CBL-interacting protein kinase 23 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G067500
594 / 0
AT5G45820 647 / 0.0
SNF1-RELATED PROTEIN KINASE 3.6, PROTEIN KINASE 18, CBL-interacting protein kinase 20 (.1)
Potri.016G133500
530 / 0
AT5G58380 593 / 0.0
SNF1-RELATED PROTEIN KINASE 3.8, CBL-INTERACTING PROTEIN KINASE 10, SOS3-interacting protein 1 (.1)
Potri.019G048800
524 / 0
AT5G58380 539 / 0.0
SNF1-RELATED PROTEIN KINASE 3.8, CBL-INTERACTING PROTEIN KINASE 10, SOS3-interacting protein 1 (.1)
Potri.019G127500
514 / 0
AT5G58380 641 / 0.0
SNF1-RELATED PROTEIN KINASE 3.8, CBL-INTERACTING PROTEIN KINASE 10, SOS3-interacting protein 1 (.1)
Potri.013G156000
505 / 5e-178
AT5G58380 623 / 0.0
SNF1-RELATED PROTEIN KINASE 3.8, CBL-INTERACTING PROTEIN KINASE 10, SOS3-interacting protein 1 (.1)
Potri.006G186200
503 / 5e-177
AT4G30960 649 / 0.0
SNF1-RELATED PROTEIN KINASE 3.14, CBL-INTERACTING PROTEIN KINASE 6, SOS3-interacting protein 3 (.1)
Potri.018G108500
494 / 8e-174
AT4G30960 639 / 0.0
SNF1-RELATED PROTEIN KINASE 3.14, CBL-INTERACTING PROTEIN KINASE 6, SOS3-interacting protein 3 (.1)
Potri.006G263500
475 / 5e-166
AT5G25110 568 / 0.0
SNF1-RELATED PROTEIN KINASE 3.25, CBL-interacting protein kinase 25 (.1)
Potri.011G067600
466 / 4e-161
AT4G18700 720 / 0.0
SNF1-RELATED PROTEIN KINASE 3.9, WPL4-LIKE 4, CBL-interacting protein kinase 12 (.1)
Potri.004G058300
462 / 1e-159
AT4G18700 735 / 0.0
SNF1-RELATED PROTEIN KINASE 3.9, WPL4-LIKE 4, CBL-interacting protein kinase 12 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0573
KA1-like
PF03822
NAF
NAF domain
CL0016
PKinase
PF00069
Pkinase
Protein kinase domain
Representative CDS sequence
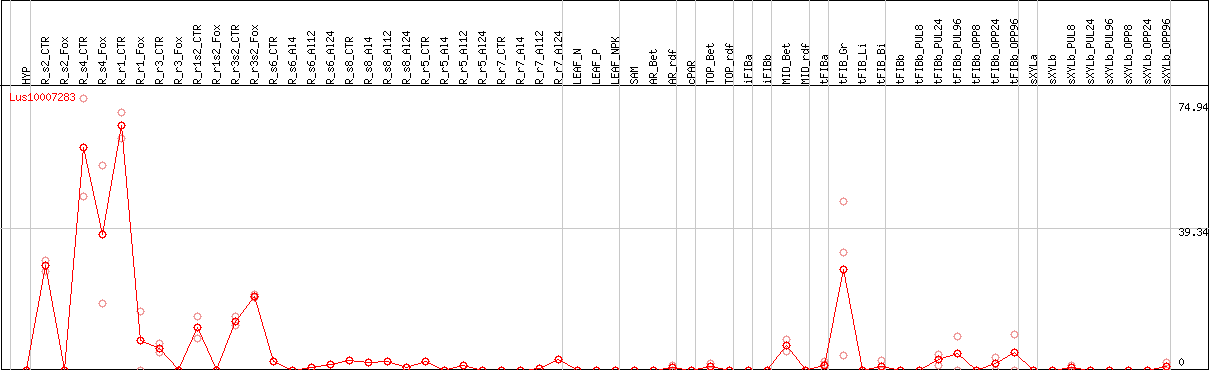
>Lus10007283 pacid=23142465 polypeptide=Lus10007283 locus=Lus10007283.g ID=Lus10007283.BGIv1.0 annot-version=v1.0
ATGGAGAAGAAGAAGCCAGGGGCAATCCTAATGAGCAAATACGAGCTGGGAAAGCTGCTCGGGAAGGGCAACTTCGCCAAAGTGTACCACGCCCGCAACC
TCCTGACGGGCCAAAACGTCGCCGTGAAAGTGATAGACAAGAGCAAGATCCCTGCCGGAATGACGGACCAGATCAAGCGAGAAATCTCCGTCATGAGCCT
CGTCCGCCACCCCAACGTCGTCCACCTCTTCGAGGTCATGGCCAGCAGGTCCAAGATCTACTTCGTCATGGAGCTCGCTCGAGGAGGCGAGCTCTTCAAC
CGGATAATCGCTGCCAAAGGTAGGCTTAAGGAGGAAACAGCGAGAAAATACTTCCAGCAATTGATCGCCGCCGTGGAATTTTGCCACAGCCGGGGAGTCT
ACCACCGGGATTTGAAACCCGAGAATCTCCTGTTGGACGAACAGGGGAATTTGAAGGTTTCGGATTTCGGGTTGAGCGCGATCCGAGACGACTCGACCCG
AAACGGAAACGACGGGCTTTTGCATACCACTTGTGGGACCCCAGCCTACGTTGCTCCTGAGGTGGTTATGAGGAGAGGATATGACGGGGCGAAGACGGAT
GTATGGTCGTGCGGGGTCGTTTTGTTTGTTTTGTTAGCGGGTTATTTGCCGTTTCACGACGCGAATTTGATGGAGATGTATCGGAAAATCGCCCGGGGGA
AATTTAAGTGTCCTAATTGGTTCAATCCGGAGGCTAGGAAGTTGGTTTCGAAGATTCTGGATCCTTCTCCGGCGAGGAGGATTACGATTGCGAGTATCAT
GACGAATTCTTGGTTCCTGAAGGGGTTTAAGAAGATCGAAACGCCGAGGTCGCCTCAGGGGGATGACCGCCGAAATATGCTCCGGGACGTGGAAACGGCA
TTCGACGGTAATATAAAGGATGACGGTGATAGTCCTACTTCGGTACTGCGACCGGCTTTTTACAACGCGTTCGACATCATTGCGCTGTCGAGAGGGTTCG
ACCTGTCGGGGCTGTTTGACAACGGCGGACGGATTAAAGAAAGGGAGTCGAGGTTCATGACGAGGAGGACGGCGGAGGAGATAGTGAATAAGTTCGAGGA
GATTGCGGCGAACGGAGGGTTTGAGGCGAGGAGGGAGGAGGGCGGTGGGACCACCGCGGTGAAGCTGGAAGGGAGGGAAGAAGGGAGGAAAGGGCAGCTG
GTGATTGACGCCGAGATATTCGAGGTGACACCAGCGTTTCACGTGGTGGAGGTAAAGAAAGCGGCGGGGGATACATTGGAGTACGTAGAGTTTTGTAACG
AGGGGTTGAGACCGTCGTTGAAGGATATTGTTTGGGCTTGGCAAGGGAACGTAGTAGAGCAGCCGGCATAG
AA sequence
>Lus10007283 pacid=23142465 polypeptide=Lus10007283 locus=Lus10007283.g ID=Lus10007283.BGIv1.0 annot-version=v1.0
MEKKKPGAILMSKYELGKLLGKGNFAKVYHARNLLTGQNVAVKVIDKSKIPAGMTDQIKREISVMSLVRHPNVVHLFEVMASRSKIYFVMELARGGELFN
RIIAAKGRLKEETARKYFQQLIAAVEFCHSRGVYHRDLKPENLLLDEQGNLKVSDFGLSAIRDDSTRNGNDGLLHTTCGTPAYVAPEVVMRRGYDGAKTD
VWSCGVVLFVLLAGYLPFHDANLMEMYRKIARGKFKCPNWFNPEARKLVSKILDPSPARRITIASIMTNSWFLKGFKKIETPRSPQGDDRRNMLRDVETA
FDGNIKDDGDSPTSVLRPAFYNAFDIIALSRGFDLSGLFDNGGRIKERESRFMTRRTAEEIVNKFEEIAANGGFEARREEGGGTTAVKLEGREEGRKGQL
VIDAEIFEVTPAFHVVEVKKAAGDTLEYVEFCNEGLRPSLKDIVWAWQGNVVEQPA