External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G18960 179 / 2e-56
MADS
AG
AGAMOUS, K-box region and MADS-box transcription factor family protein (.1.2)
AT3G58780 138 / 2e-40
MADS
AGL1, SHP1
SHATTERPROOF 1, AGAMOUS-like 1, K-box region and MADS-box transcription factor family protein (.1.2.3)
AT2G42830 130 / 2e-37
MADS
AGL5, SHP2
SHATTERPROOF 2, AGAMOUS-like 5, K-box region and MADS-box transcription factor family protein (.1.2)
AT4G09960 108 / 4e-29
MADS
AGL11, STK
SEEDSTICK, AGAMOUS-like 11, K-box region and MADS-box transcription factor family protein (.1.2.3.4)
AT1G69120 56 / 4e-09
MADS
AGL7, AP1
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
AT1G26310 52 / 6e-08
MADS
CAL1, AGL10, CAL
CAULIFLOWER, AGAMOUS-like 10, K-box region and MADS-box transcription factor family protein (.1)
AT5G60910 50 / 2e-07
MADS
FUL, AGL8
FRUITFULL, AGAMOUS-like 8 (.1.2)
AT2G45660 43 / 4e-05
MADS
ATSOC1, SOC1, AGL20
SUPPRESSOR OF OVEREXPRESSION OF CO 1, AGAMOUS-like 20 (.1)
AT4G22950 43 / 5e-05
MADS
AGL19
AGAMOUS-like 19 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10029275
186 / 1e-58
AT3G58780 220 / 2e-71
SHATTERPROOF 1, AGAMOUS-like 1, K-box region and MADS-box transcription factor family protein (.1.2.3)
Lus10005302
113 / 2e-30
AT4G09960 330 / 2e-115
SEEDSTICK, AGAMOUS-like 11, K-box region and MADS-box transcription factor family protein (.1.2.3.4)
Lus10008264
110 / 5e-29
AT4G09960 328 / 3e-114
SEEDSTICK, AGAMOUS-like 11, K-box region and MADS-box transcription factor family protein (.1.2.3.4)
Lus10009481
103 / 3e-26
AT4G09960 274 / 1e-92
SEEDSTICK, AGAMOUS-like 11, K-box region and MADS-box transcription factor family protein (.1.2.3.4)
Lus10028215
48 / 3e-07
AT3G57230 164 / 4e-52
AGAMOUS-like 16 (.1.2)
Lus10034662
49 / 6e-07
AT5G60910 284 / 3e-97
FRUITFULL, AGAMOUS-like 8 (.1.2)
Lus10017870
48 / 1e-06
AT1G69120 212 / 3e-69
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Lus10006715
48 / 1e-06
AT4G22950 223 / 1e-73
AGAMOUS-like 19 (.1)
Lus10017871
47 / 2e-06
AT1G69120 264 / 2e-90
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF01486
K-box
K-box region
Representative CDS sequence
>Lus10007324 pacid=23142493 polypeptide=Lus10007324 locus=Lus10007324.g ID=Lus10007324.BGIv1.0 annot-version=v1.0
ATGGGTGCAGCACCAACTGCATGGAAGGCATTGATCAATGCTAGTTGTGATTCTTTCTTAAAGGTTAAAGACACAATAGAGAGGTACAAGAAAGCCAATT
CAGACTCTACAAACACTGGATCTGTTGCTGAAGCAAATGCTCAGTACTATCAGGAGGAAGCAGCAAAGCTGAGGCAGAAAATTGGGAATCTGCTGAATAC
AAACAGGAAACTTGCTATGTTTGTCAACCATAACAACAACGACGGTGGCGGTGGGGGATCTTCTTCTAGACATATGCCTGGTGAATCACTAGGAGGGCTC
GGTCCCAAGGATCTCAAGAACTTGGAGAGCAAACTTGAGAAAGGAATCAGCAGGATCAGATCCAAAAAGAATGAACTGCTGTTTTCGGAGATAGAGTATA
TGCAGAAGAGGGAAGTTGATCTGCACAACAACAACCAGCTTCTTCGTGCAAAGATAGCTGATAATGAGAGGAAGCAGCAGAACATGAGCTTGATGCCAGG
AGGAGGAGGAGGTGGTGGAGGGAGTACTAGCTACGAGATTGTGCACCCTCATCAGCATCAGCAATTTGATTCCCGAAACTACTTCCAAGTGAATGCTTTA
CAACCCACCAACCATTACCCTCAACAAGACCATATGTCACTTCAGTTGGGGTATAAAGATTAA
AA sequence
>Lus10007324 pacid=23142493 polypeptide=Lus10007324 locus=Lus10007324.g ID=Lus10007324.BGIv1.0 annot-version=v1.0
MGAAPTAWKALINASCDSFLKVKDTIERYKKANSDSTNTGSVAEANAQYYQEEAAKLRQKIGNLLNTNRKLAMFVNHNNNDGGGGGSSSRHMPGESLGGL
GPKDLKNLESKLEKGISRIRSKKNELLFSEIEYMQKREVDLHNNNQLLRAKIADNERKQQNMSLMPGGGGGGGGSTSYEIVHPHQHQQFDSRNYFQVNAL
QPTNHYPQQDHMSLQLGYKD
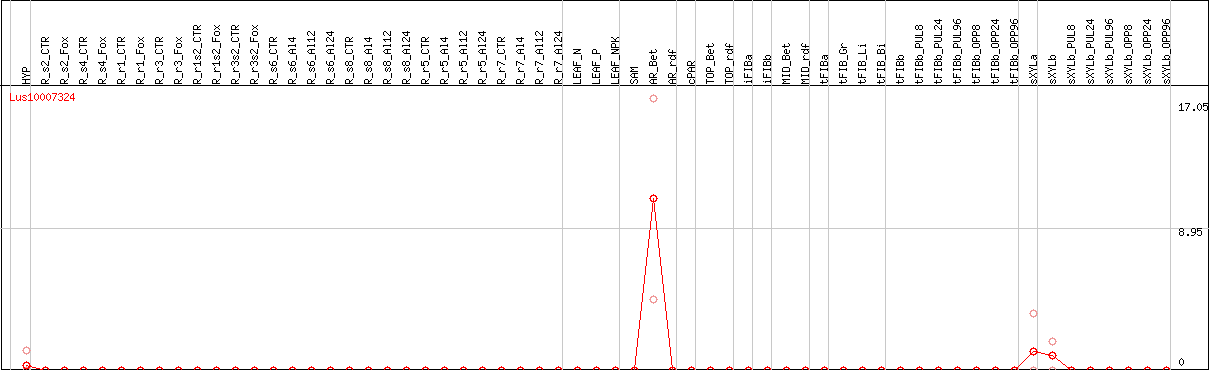
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10007324 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.