External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G65690 231 / 1e-75
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
AT5G36970 219 / 3e-71
NHL25
NDR1/HIN1-like 25 (.1)
AT1G54540 159 / 9e-48
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
AT2G27080 117 / 2e-31
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1), Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.2)
AT5G21130 107 / 1e-27
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
AT3G52470 73 / 2e-15
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
AT5G53730 71 / 2e-14
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
AT5G06320 71 / 3e-14
NHL3
NDR1/HIN1-like 3 (.1)
AT3G11650 70 / 7e-14
NHL2
NDR1/HIN1-like 2 (.1)
AT3G11660 67 / 3e-13
NHL1
NDR1/HIN1-like 1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10020783
391 / 1e-138
AT5G36970 229 / 2e-75
NDR1/HIN1-like 25 (.1)
Lus10031888
160 / 4e-48
AT1G54540 236 / 2e-78
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Lus10031317
148 / 2e-43
AT1G54540 231 / 1e-76
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Lus10021288
74 / 2e-15
AT2G35980 237 / 2e-79
YELLOW-LEAF-SPECIFIC GENE 9, ARABIDOPSIS NDR1/HIN1-LIKE 10, Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Lus10021287
72 / 1e-14
AT2G35980 236 / 7e-79
YELLOW-LEAF-SPECIFIC GENE 9, ARABIDOPSIS NDR1/HIN1-LIKE 10, Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Lus10016962
72 / 1e-14
AT2G35980 237 / 3e-79
YELLOW-LEAF-SPECIFIC GENE 9, ARABIDOPSIS NDR1/HIN1-LIKE 10, Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Lus10037638
71 / 3e-14
AT1G17620 201 / 4e-64
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Lus10015622
67 / 6e-13
AT1G17620 196 / 5e-62
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Lus10000157
66 / 9e-13
AT3G52470 164 / 3e-51
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.015G148200
248 / 3e-82
AT5G36970 227 / 2e-74
NDR1/HIN1-like 25 (.1)
Potri.012G145500
244 / 1e-80
AT5G36970 238 / 2e-78
NDR1/HIN1-like 25 (.1)
Potri.017G154000
180 / 1e-55
AT1G54540 242 / 1e-80
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Potri.004G067100
163 / 4e-49
AT1G54540 203 / 3e-65
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Potri.009G158900
141 / 2e-40
AT2G27080 259 / 8e-87
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1), Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.2)
Potri.004G197600
115 / 1e-30
AT2G27080 264 / 8e-89
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1), Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.2)
Potri.013G135800
100 / 3e-25
AT1G65690 112 / 4e-30
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Potri.003G039400
83 / 2e-18
AT1G17620 208 / 1e-66
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Potri.009G003800
76 / 1e-16
AT5G22870 179 / 4e-57
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
Potri.001G200700
74 / 2e-15
AT1G17620 168 / 2e-51
Late embryogenesis abundant (LEA) hydroxyproline-rich glycoprotein family (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0159
E-set
PF03168
LEA_2
Late embryogenesis abundant protein
Representative CDS sequence
>Lus10007364 pacid=23166254 polypeptide=Lus10007364 locus=Lus10007364.g ID=Lus10007364.BGIv1.0 annot-version=v1.0
ATGGCCGACCAGCACAAGAAGATTCACCCACTGAACTCCAACAATCAAAACGACGACGTAGAAGCGGCCGCCGCGTCCTCCTCCTCATCAGACGCTCCCC
ACCATGCTCCAACAGCACCTTTAGTGCCCCGTGGATCATCCAAATCCGACCACGGCGACCCGACCCGGAAAGCTCCGAGTCAAGTACCACCGTACCCTCC
ACCAACATACCACGGGGCGAAGCGGCCGAAAAGACGCCGTTCCACCTGTTGCCGATGCTTCTGTTGGATATTCTCAATCCTGCTCTTCCTAGTCATAGCG
GTGGGGGCCACAGTGGGGATCCTCTTCCTAATCTTCCGCCCCAAGCTACCTGACGTGTCAATCGACCGGCTGCAAATCGCGCGGTTCGGCCTGACACAAA
CCACCCCTGCCACGCTGTCCGCCACGTTTGATGTCACCATCACGGCTAAGAACCCGAACAAGAGGATCGGGATATACTACGAAGGAGGGAGCAGGATCGG
AGTTTCGTTCGCGGGTACGCAGCTCTGCCAGGGGGCGCTGCCGAAGTTCTACCAGGGCCACCGGAACACGACGGTGCTGAACGTCCCGCTGACTGGAGAA
AGCGCGGACGCGGCGGCGCTGTTTGGGGAGATGCAGAGTGAGCAGCAGAGGAATGGGTTCGTGCCGTTGGATTTGAGGGTTAGGCAGCCTGTGAGGATTA
AGCTCGGGAAGCTGAAGCTGCCTAAGATTAAGTTTGTGGTGAGGTGTCGGCTTAACGTGGATAGCTTGACCGCGAATAATAATATCCGGATTAGGAACAG
AAGTTGTAAGTTCAGGCTGAGGCTCTGA
AA sequence
>Lus10007364 pacid=23166254 polypeptide=Lus10007364 locus=Lus10007364.g ID=Lus10007364.BGIv1.0 annot-version=v1.0
MADQHKKIHPLNSNNQNDDVEAAAASSSSSDAPHHAPTAPLVPRGSSKSDHGDPTRKAPSQVPPYPPPTYHGAKRPKRRRSTCCRCFCWIFSILLFLVIA
VGATVGILFLIFRPKLPDVSIDRLQIARFGLTQTTPATLSATFDVTITAKNPNKRIGIYYEGGSRIGVSFAGTQLCQGALPKFYQGHRNTTVLNVPLTGE
SADAAALFGEMQSEQQRNGFVPLDLRVRQPVRIKLGKLKLPKIKFVVRCRLNVDSLTANNNIRIRNRSCKFRLRL
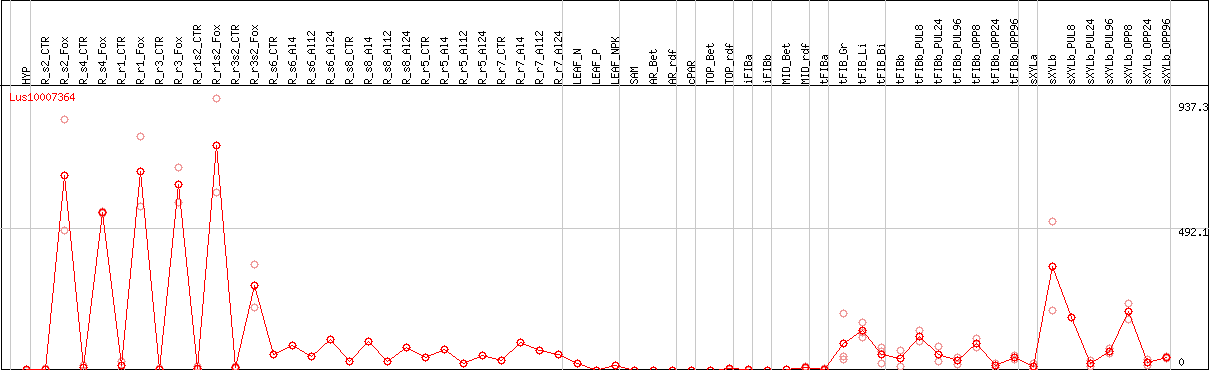
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10007364 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.