Lus10007461 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||
| PFAM info | |||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10007461 pacid=23178391 polypeptide=Lus10007461 locus=Lus10007461.g ID=Lus10007461.BGIv1.0 annot-version=v1.0
ATGGCTCCATTCCCTCTTACTCTCCTCCATTCTCTCTTCTTCTTCGCCCTCTTATCATCATCCGCCGCCATACCCGATTCCTCTTCTTCGTATTCCGATG
CCGATGCGGATTCCGATTCCGCTTCCTACCTTTCTCTCTTCCATCAAGACTACTCTCCTCCCTCCCCTCCTCCTCCGCCTCCCCACCCCCCTCCCCTTTC
ATGCACCGACGACCTCCACGGAATCGGCTCCTTGGACAACACTTGCCAGATAATCTCCGATGTCAACCTCACCGCCGACGTCTACATCCAGGGCCAAGGC
AACTTCTACATACACCCTGGCGTCAGCTTCAGCTGTCCTTCCCCTGGCTGTTCAATCACCCTCAACATCTCCGGCAACCTCACCCTCGCTCCCAACTCCT
CCATCGTCGCCGGTGCCTTTGAACTCTTCGCCTCCAATGCCTCCTTTCTTAACGGCACCTTCGTCAACACCACCGGCTTGGCTGGGGATCCGCCTCCCCA
AACCAGTGGCACGCCTCAAGGTGTCGATGGTGCTGGTGGAGGCCATGGCGGTCGTGGCGCTGCTTGCTTAGTCGACCTCACCAAGCTCCCCGAAGATATT
TGGGGTGGGGATGCATACTCATGGGCTAGTTTGCAGGAGCCTTGTAGTTATGGGAGCAAAGGTGGAACTACCAGTAACGAGGAGGATTACGGTGGGGGTG
GCGGTGGAATGGTGAGGATGACCATTGATGATTATATAGAGGTTGATGGGCTTGTTTTAGCTGATGGAGGGGACGGTGGTACCAAGGGAGGCGGTGGTTC
TGGCGGTAGCATTTACATCAAGGCTTATAAAATGACGGGTAGTGGCGAGATTAGTGCTTGTGGAGGAAGTGGTTTTGCTGGAGGAGGTGGAGGAAGAATT
TCTGTGGACATTTTCAGTAGGCATGATGATCCTCGAGTATTTGTGCATGGAGGAAGTAGTCGTGGCTGTCCGGAAAACGCTGGGGCTGCTGGGACCCTAT
ATGATGCTGTTCCACGAAGTCTTATTGTTAGCAATGACAATCTGTCAACGAATACAGACACTCTTCTCCTGGACTTTCCTAATCAACCCCTTTGGACGAA
CATTTATGTCCGCAATCATGCTCGTGCCACTGTTCCATTGCTCTGGAGTCGTGTCCAGGTGCAAGGACAGATAAGTTTGCTGTCTGGCGGACTACTGAGT
TTTGGGCTAGCTCACTACGCTACTTCTGAATTTGAACTATTGGCTGAAGAACTATTAATGAGCGACTCTGTAATGAAGGTTTACGGAGCTCTACGCATGA
CTGTGAAAATGTTTTTGATGTGGAATTCCGAAATGCTCATAGATGGAGGTGAAGATGCTGCTGTTGCTACTTCCTTGCTTGAAGCCAGTAATTTAGTTGT
TCTGAGGGAGTCATCGGTCATACAGTCTAATGCGAATCTGGGTGTTCATGGACAAGGTTTGTTGAATTTGTCTGGACCAGGTGACAGGATTGATGCCCAA
CGTCTTGTCCTATCATTGTTCTACGGTGTTCATGTTGGTCCTGGATCTGTTTTACGTGGTCCTCTGGAGAATGCAACTAGTGATGCTATTACACCACAAC
TACATTGTGACTTTGAAGATTGCCCTGTTGAACTACTTCATCCGCCTGAAGACTGCAACGTGAATTCATCACTGTCCTTTACTCTCCAGATATGCCGAGT
TGAGGATATTACTGTCGAGGGACTAGTAGAAGGATCTGTTGTCCATTTCCATCGTGCAAGAACTATATCAGTCCAACCTTCAGGAATAATTAGTGCATCT
GGAACAGGCTGCACAGGCGGTGTGGGAAGAGGTAATTTCTCTAGTGATGGTGTTGGTAGTGGTGGAGGCCACGGTGGCAAAGGTGGGCTAGGATGTTACA
ATGATAGTTGCATTGATGGCGGCATTTCATACGGAAACCCAGTCTTTCCATGTGAACTTGGTAGTGGCAGTGGGTACGGAAGTTTGGCTGGCTCAACTGC
AGGGGGTGGGATCATAGTTATGGGTTCTCTGGAGCATCCATTGTCAAGCTTGTCTGTTAAAGGATCAGTCAGAGCTGATGGAGAAAGTTTTCAACAAACT
ATTAAACGTAGGGAGTTCGCTGTTATGAATAGTAGTGGTTGGGGACCTGGGGGTGGATCTGGGGGAAGCATACTTATGTTCTTGAGGACGATAGAACTTG
CTGAGACTGCTCTACTTTCAAGTTCTGGTGGATATGGTAGCCCACAAGGGAGTGGTGGAGGTGGAGGAGGGAGGATTCACTTTCACTGGGCGGATATTCC
AGCTGGTGACGTGTACCAGCCAATTGCCAGTGTGAAAGGGGGCATTCACTCTAGGGGAGGGCTAGGTAAGGATGGAGGTGGTTCTGGGGGAAACGGAACA
GTAACTGGAAAAGCCTGCCCGAAAGGACTATATGGAGTATTTTGCAAGGAATGTCCAGCTGGAACTTACAAGAACGACACTGGATCTGACGAAGCCCTTT
GCCGATCATGTCCTCCTAATGAGCTTCCTAATCGTGCTCTATACATTTCTGTCAGAGGTGGTATTGCTGAAACACCATGTCCATATAGATGTATTTCCGA
CAGATTTCACATGCCCCACTGCTATACGGCCCTTGAAGAGTTGATTTACACATTTGGTGGACCATGGTTATTTGGACTTCTTCTTGTGGGTCTACTCATT
CTTTTGGCACTGGTACTAAGTGTTGCCCGGATGAAATTTGTTGGTGGTGATGAACTACCTGGCCCAGCTCCCACCCAACACGGTTCTCAAATAGATCACT
CCTTTCCGTTTTTGGAATCATTGAATGAGGTGCTAGAAACAAACAGAGCTGAGGAGTCCCAGAGTCATGTGCACAGGATGTATTTTCTGGGTCCTAATTC
TTTCAGTGAACCTTGGCATCTTCCACACTCACCCCCTGAGCAAATAAAAGAGACAGTATACGAAGGCGCATTTAATACATTTGTCGATGAGATTAATGCA
ATAGCTACGTATCAATGGTGGGAAGGTGCCATCCACAGTATCCTTTCTGTAACTTTATATCCCTTGGCATGGTCATGGCAGCATTGGCGTCGCAGAATCA
AACTGCAAAGACTTCGGGAGTTTGTTCGATCAGAGTATGATCATGCTTGCTTACGCTCTTGCCGGTCCAGGGCTCTCTATGAAGGGATTAAGGTTGCTGC
CACTTCTGATTTAATGCTTGCATATGTTGACTTCTTCCTCGGTGGAGATGAGAAGAGAAGTGACCTTCCTCCTCGACTGTATCAACGATTTCCAATGTCT
ATAATTTTTGGGGGTGATGGCAGCTATATTACTCCATTCTCTATTCAAAGTGATAACATTGTCAGCAGTCTCATGGGCCAGCTGGTCGCTCCCACCACAT
GGTATCGAATGGTGGCTGGTCTGAATGCACAACTTCGATTAGTTCGTCGAGGACGGCTGAGAGTAACTTTCAAGCCAGTCATCAAATGGCTGGAAACTCA
TGCTAACCCAGCTTTGAAAGTGTACCAAGTACAAGTTGATCTTGTGAGGCTTCAGGCTACAGCTTGTGGTTATCGCCAGTATGGACTTTTGGTTTATGCT
GCTGATGATGAAAACCAGTATGCATCTAGTGACAGTCTTGAGGATACAAAGCCAACCACACAACATTCACGCATGAAGAAGACGGAAGGAGAATATGCAT
TTGGTCATTTGAGCGAGCCTATTCTCTTAACCCGATCTGATAGCAGCGAGAGTTATTTGAGGCGGATGAAGACTCATGGCTGTGTGATTGATGCTGACAA
TTTAAAAACACTTGAAGAGAAGAGAGATATTTTTTACATTCTCTCTCTCTTAGTTTATAATACTAAACCAGTTGGCCACCAGGATCTTGTTGGTTTGGTC
ATCTCAATGCTGCTTCTAGGAGATTTTAGCTTAGTGTTGCTTACTTTGCTCCAGCTGTATTCAATTTCACTGGGTGATGTTTTCCTGGTTTTGTTCATCC
TGCCTCTGGGTATTCTTCTGCCATTTCCTGCTGGGATCAATGCTTTATTCAGTCATGGACATAGACGTTCAGCTGGGCTTGCGCGTATCTATGCACTGTG
GAATATTACATCATTGATGAATGTGGTTGTTGCTTTTGTGTGCGGGTACGTCCACTACATCAGTCAATCGTCATCAAGCAAGAAGTTCCCCTTTCAGCCG
TGGAATATCAGCATGGATGAGAGTGAATGGTGGATATTCCCGGGGGCACTGGTTATATGTAAAATGGTGCAGTCCCAGCTAGTGAATTGGCATGTGGGAA
ATCTAGAGATTCAAGACCGTTCCCTGTACAGTAATGATTTCCAGCTGTTCTGGCAGTCTTGA
|
||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10007461 pacid=23178391 polypeptide=Lus10007461 locus=Lus10007461.g ID=Lus10007461.BGIv1.0 annot-version=v1.0
MAPFPLTLLHSLFFFALLSSSAAIPDSSSSYSDADADSDSASYLSLFHQDYSPPSPPPPPPHPPPLSCTDDLHGIGSLDNTCQIISDVNLTADVYIQGQG
NFYIHPGVSFSCPSPGCSITLNISGNLTLAPNSSIVAGAFELFASNASFLNGTFVNTTGLAGDPPPQTSGTPQGVDGAGGGHGGRGAACLVDLTKLPEDI
WGGDAYSWASLQEPCSYGSKGGTTSNEEDYGGGGGGMVRMTIDDYIEVDGLVLADGGDGGTKGGGGSGGSIYIKAYKMTGSGEISACGGSGFAGGGGGRI
SVDIFSRHDDPRVFVHGGSSRGCPENAGAAGTLYDAVPRSLIVSNDNLSTNTDTLLLDFPNQPLWTNIYVRNHARATVPLLWSRVQVQGQISLLSGGLLS
FGLAHYATSEFELLAEELLMSDSVMKVYGALRMTVKMFLMWNSEMLIDGGEDAAVATSLLEASNLVVLRESSVIQSNANLGVHGQGLLNLSGPGDRIDAQ
RLVLSLFYGVHVGPGSVLRGPLENATSDAITPQLHCDFEDCPVELLHPPEDCNVNSSLSFTLQICRVEDITVEGLVEGSVVHFHRARTISVQPSGIISAS
GTGCTGGVGRGNFSSDGVGSGGGHGGKGGLGCYNDSCIDGGISYGNPVFPCELGSGSGYGSLAGSTAGGGIIVMGSLEHPLSSLSVKGSVRADGESFQQT
IKRREFAVMNSSGWGPGGGSGGSILMFLRTIELAETALLSSSGGYGSPQGSGGGGGGRIHFHWADIPAGDVYQPIASVKGGIHSRGGLGKDGGGSGGNGT
VTGKACPKGLYGVFCKECPAGTYKNDTGSDEALCRSCPPNELPNRALYISVRGGIAETPCPYRCISDRFHMPHCYTALEELIYTFGGPWLFGLLLVGLLI
LLALVLSVARMKFVGGDELPGPAPTQHGSQIDHSFPFLESLNEVLETNRAEESQSHVHRMYFLGPNSFSEPWHLPHSPPEQIKETVYEGAFNTFVDEINA
IATYQWWEGAIHSILSVTLYPLAWSWQHWRRRIKLQRLREFVRSEYDHACLRSCRSRALYEGIKVAATSDLMLAYVDFFLGGDEKRSDLPPRLYQRFPMS
IIFGGDGSYITPFSIQSDNIVSSLMGQLVAPTTWYRMVAGLNAQLRLVRRGRLRVTFKPVIKWLETHANPALKVYQVQVDLVRLQATACGYRQYGLLVYA
ADDENQYASSDSLEDTKPTTQHSRMKKTEGEYAFGHLSEPILLTRSDSSESYLRRMKTHGCVIDADNLKTLEEKRDIFYILSLLVYNTKPVGHQDLVGLV
ISMLLLGDFSLVLLTLLQLYSISLGDVFLVLFILPLGILLPFPAGINALFSHGHRRSAGLARIYALWNITSLMNVVVAFVCGYVHYISQSSSSKKFPFQP
WNISMDESEWWIFPGALVICKMVQSQLVNWHVGNLEIQDRSLYSNDFQLFWQS
|
DESeq2's median of ratios [FLAX]
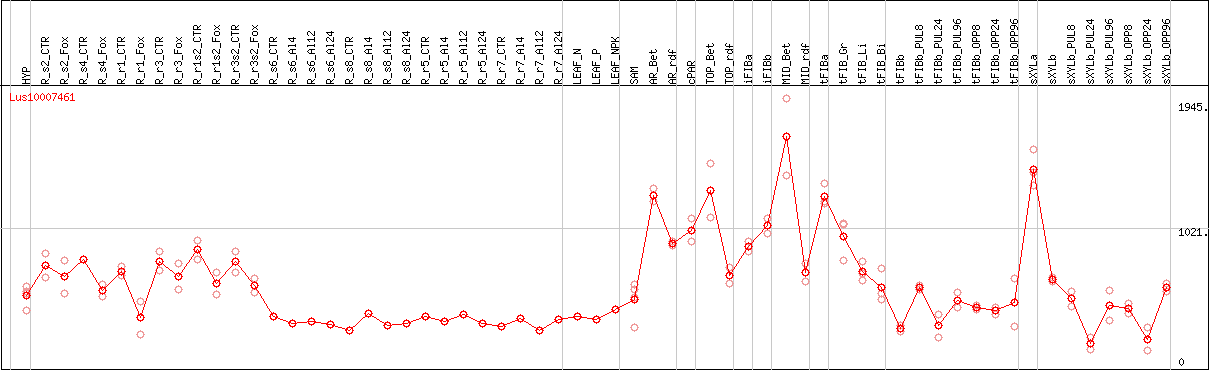
Coexpressed genes
Lus10007461 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
