External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G40320 229 / 5e-75
TBL33
TRICHOME BIREFRINGENCE-LIKE 33 (.1)
AT3G11030 215 / 2e-69
TBL32
TRICHOME BIREFRINGENCE-LIKE 32 (.1)
AT1G73140 149 / 7e-44
TBL31
Plant protein of unknown function (DUF828) (.1)
AT5G01620 146 / 9e-43
TBL35
TRICHOME BIREFRINGENCE-LIKE 35 (.1.2.3)
AT5G01360 143 / 1e-41
TBL3
TRICHOME BIREFRINGENCE-LIKE 3, Plant protein of unknown function (DUF828) (.1), Plant protein of unknown function (DUF828) (.2)
AT2G38320 130 / 4e-37
TBL34
TRICHOME BIREFRINGENCE-LIKE 34 (.1)
AT3G55990 130 / 1e-36
TBL29, ESK1
TRICHOME BIREFRINGENCE-LIKE 29, ESKIMO 1, Plant protein of unknown function (DUF828) (.1)
AT3G12060 128 / 2e-35
TBL1
TRICHOME BIREFRINGENCE-LIKE 1, Plant protein of unknown function (DUF828) (.1)
AT2G40150 125 / 6e-35
TBL28
TRICHOME BIREFRINGENCE-LIKE 28 (.1)
AT2G40160 125 / 6e-35
TBL30
Plant protein of unknown function (DUF828) (.1), Plant protein of unknown function (DUF828) (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017428
300 / 3e-103
AT2G40320 540 / 0.0
TRICHOME BIREFRINGENCE-LIKE 33 (.1)
Lus10009839
249 / 2e-82
AT2G40320 678 / 0.0
TRICHOME BIREFRINGENCE-LIKE 33 (.1)
Lus10040953
243 / 4e-80
AT2G40320 675 / 0.0
TRICHOME BIREFRINGENCE-LIKE 33 (.1)
Lus10042845
149 / 1e-43
AT1G73140 559 / 0.0
Plant protein of unknown function (DUF828) (.1)
Lus10012229
144 / 4e-43
AT5G01620 509 / 0.0
TRICHOME BIREFRINGENCE-LIKE 35 (.1.2.3)
Lus10002858
143 / 2e-42
AT5G01620 510 / 0.0
TRICHOME BIREFRINGENCE-LIKE 35 (.1.2.3)
Lus10028141
144 / 4e-42
AT1G73140 563 / 0.0
Plant protein of unknown function (DUF828) (.1)
Lus10027813
144 / 6e-42
AT5G01360 570 / 0.0
TRICHOME BIREFRINGENCE-LIKE 3, Plant protein of unknown function (DUF828) (.1), Plant protein of unknown function (DUF828) (.2)
Lus10005042
142 / 2e-41
AT5G01360 560 / 0.0
TRICHOME BIREFRINGENCE-LIKE 3, Plant protein of unknown function (DUF828) (.1), Plant protein of unknown function (DUF828) (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G073300
263 / 4e-88
AT2G40320 694 / 0.0
TRICHOME BIREFRINGENCE-LIKE 33 (.1)
Potri.010G184000
256 / 2e-85
AT2G40320 692 / 0.0
TRICHOME BIREFRINGENCE-LIKE 33 (.1)
Potri.016G125500
153 / 2e-45
AT2G38320 541 / 0.0
TRICHOME BIREFRINGENCE-LIKE 34 (.1)
Potri.016G125600
149 / 7e-44
AT5G01620 657 / 0.0
TRICHOME BIREFRINGENCE-LIKE 35 (.1.2.3)
Potri.001G376700
143 / 9e-42
AT1G73140 640 / 0.0
Plant protein of unknown function (DUF828) (.1)
Potri.016G119100
143 / 1e-41
AT5G01360 578 / 0.0
TRICHOME BIREFRINGENCE-LIKE 3, Plant protein of unknown function (DUF828) (.1), Plant protein of unknown function (DUF828) (.2)
Potri.008G070000
143 / 2e-41
AT3G55990 530 / 0.0
TRICHOME BIREFRINGENCE-LIKE 29, ESKIMO 1, Plant protein of unknown function (DUF828) (.1)
Potri.010G187600
139 / 1e-39
AT3G55990 660 / 0.0
TRICHOME BIREFRINGENCE-LIKE 29, ESKIMO 1, Plant protein of unknown function (DUF828) (.1)
Potri.010G187300
136 / 7e-39
AT3G55990 511 / 3e-179
TRICHOME BIREFRINGENCE-LIKE 29, ESKIMO 1, Plant protein of unknown function (DUF828) (.1)
Potri.008G069900
135 / 2e-38
AT3G55990 686 / 0.0
TRICHOME BIREFRINGENCE-LIKE 29, ESKIMO 1, Plant protein of unknown function (DUF828) (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
Representative CDS sequence
>Lus10007529 pacid=23176503 polypeptide=Lus10007529 locus=Lus10007529.g ID=Lus10007529.BGIv1.0 annot-version=v1.0
ATGGCGATGAGGAGTATGCTCCGGTGGGTTAGGAAGAATGTTGACCGGAAGAGTGGGCGGGTTTTTTTCACCGGCATGTCTCCTACTCATGGCAATAGCG
CGGAATGGGGAGGGCAGGCCGGGGGCAACTGCTACAACGAAACATCACCGATCACGAACAACGCAACGTACTGGGGATCAGACAGCAAGAGGAGCATAAT
GCAAGTGATCGGCGACGAGTTCCGAAAATCGAAATTCCCGGTGACGTTCCTCAACATAACGCAGCTCTCCAACTACAGGAAAGACGCCCACACTTCTATC
TACAAGAAACAGTGGAGCCCACTAACGCCGGAGCAACTGGCCAACCCGGTCAGCTACTCTGACTGCGTCCACTGGTGCTTGCCTGGCCTACAAGATACAT
GGAATGAGCTCCTTTTTGCCAAGCTTTTCTACCCCTGA
AA sequence
>Lus10007529 pacid=23176503 polypeptide=Lus10007529 locus=Lus10007529.g ID=Lus10007529.BGIv1.0 annot-version=v1.0
MAMRSMLRWVRKNVDRKSGRVFFTGMSPTHGNSAEWGGQAGGNCYNETSPITNNATYWGSDSKRSIMQVIGDEFRKSKFPVTFLNITQLSNYRKDAHTSI
YKKQWSPLTPEQLANPVSYSDCVHWCLPGLQDTWNELLFAKLFYP
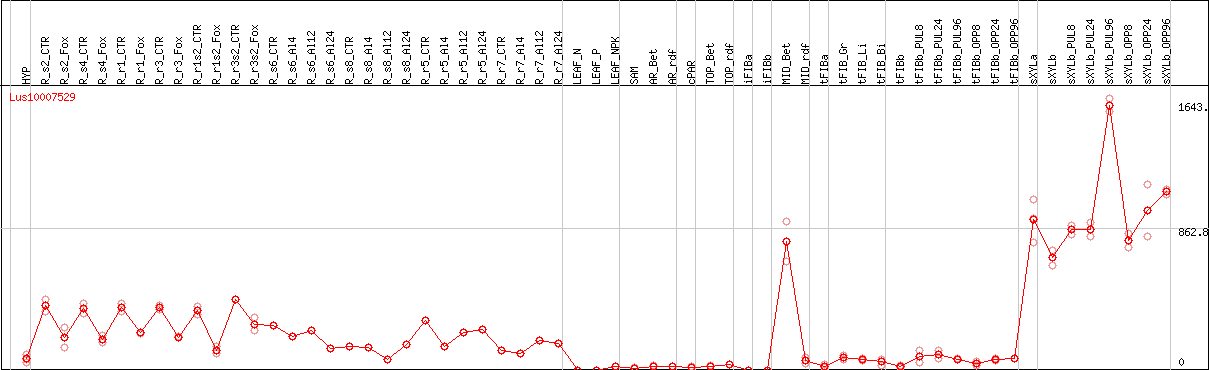
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10007529 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.