Lus10007586 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10007586 pacid=23148098 polypeptide=Lus10007586 locus=Lus10007586.g ID=Lus10007586.BGIv1.0 annot-version=v1.0
ATGGACCAGCGTCAAAAACTCAGAGATCCACTCCTCGCCGACGGAGATGATGATCGGAAGCCGCTCAGAGGCGCCGCCGGCGATCGGTTCAGGGGATTCG
CCAAATGGGTTCTCGAGATGTTGATGTGGGCAATCTTCATTGTGTGGCTCACTCTCGTCTTCGTCTTCCCAACTGAGCTGGGCAACGATTTGGTCTTCAA
GAAGTGGACTGCCCTCACCAAGACTCCTCTCTTCGGCATCACAGGGAGCTTGTTGTTGCTGCTGGCCGGACCGGTGCTGCTCATCGCACTCCTGGCCGCG
GCCCATCTCATCATTTCCGGGGAAGGGGAGCATCACCTTTACCAGAAGAAGAGCTGGAAGTACCCGAGGCCCCAGCTATGGACATTCCCGGTGATCGTGG
ATGGACCACTCGGGGTTGTATCGGCGACAGAGCTTATCGGAGTTGTGCTCTTCGTTGGCTACATTGTATGGGCTTTTTACCTCTACACCGTCCGGAACCT
CGCTCTCTTGTCTTCCTACACCCTCACTCTCCCACAATTCAGTTTGTTTCTGATGGAGCTCACGGGGCTACGATTGGGAATGATCGGGATGTTCTGCTTG
GCGTTTCTCTTCCTCCCGATCTCGAGAGGATCGATGCTCCTTCGACTCATCGACATCCCTGTAGAGCACGCAACGAGATACCATGTTTGGTTGGGACACA
TAACCATGTTGTTGTTCACTCTCCATGGACTGTTCTATGTCGTCGCTTGGGCTGTTCAGGGAAAGCTCTTAGAGGAAATATTGGAGTGGAAGAACATTGG
GATATCCAACCTGGCAGGGTTGATTTCCCTCGTGGCAGGTCTGTTGATGTGGGTGACATCACTAAACCCAGTCAGGAGATGGAACTTTGAGCTCTTCTTC
TACACACATCAACTCTACATTGTCTTTGTTCTCTTCTTGGCCTTCCACGTCGGGGACTTCCTTTTCACCATCGCTGCCGGTGGGATCTTCCTCTTCATCC
TCGATCGATTTCTCCGGTTCTGCCAGTCCAGGAAGACCGTTGATGTCGTCTCTGCCAAATGCTTCCCCTGTGGCACTGTTGAGCTGACCCTCTCCAAGCC
CGCAAATCTAAGGTACAATGCTCTGAGTTTCATCTTCATACAAGTCAGGGAACTGTCCTTGCTGCAATGGCACCCTTTCAGTGTTTCATCCAGTCCTTTA
GAAGGCAAGAATCACATATCAGTTCTCATCAAAGTCCTCGGGGAGTGGACTGACAAGCTCAAAGATACCATCAAGATCAATACCTCGACAGTCAACAACG
ATCCTGCATCTCGAAAGATCACAGCTTCTGTCGAAGGTCCATACGGGCACGAGTCACCTTACCACTTGATGTATGAGCACCTTGTTCTAGTAGCAGGAGG
CATAGGGATCTCACCCTTCATGGCAATCTTGAGTGATGTCTTCCACAGAGTCAATGATGGCAAACCCTGCCTACCAAGAGAAATCTTGGTAGTGTGGGCC
GTCAAGAAGTCCGAAGAGATCGGACTTCTGTCAGCGATGAATCTCGACCGCATATGTTCAGATCTCTATGACAGGCTCAATTTGCGTATCCACATCTACG
TTACTCGAGAGTCAGATCCTCCACTGGAAGAAGGTAAAGTGTCGACCGAGGCATCAGGGGTTTCGGCCACCAGGTGCCCCTTCCCGATAGGGAACGACAT
GTCGGTCCTGGTTGGTACGGGCGATAATGTATGGTCCGGACTGTATATTTTGTCGTCGACGTTGGGTTTCATCCTTTCGCTCGCTATGACTGACGCGTAT
TACATCAACCCATATGGGGTAACATCATGGTGGTACAAGGGGCTTCTCCTTGTGGCGTGCATGGTTGGAAGCGTGGTTGTTTTCGGCGGTGCGGTTGTTG
CATTGTGGCATCTCTGGGGTAGAAAAGCTTCAACCTTGGAGCAAGATGATGAGGAAACAATGACCAAAGTTGATGAAGGTTTGCCCAATAGTACTCCTAA
CGGGACAAGGATTGTTGAAGATTTGCTTGCTTCAAGTTCAACCTCCGTCAACTATGGTTCAAGACCAGACTTCAAAGAAATAATGCAAAGTATATCGAAG
GACTGGGGGCATGTTGATGTTGGTGTGATGGTATGTGGACCTCCAACTCTTCAGTCGACCGTCGCCAAGGAAATTAGGGCACACAATTTCAGAAGGAGCT
CTGGTGAAACTGTGTTCCATTTCAATAGCCACAGTTTTGATCTTTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10007586 pacid=23148098 polypeptide=Lus10007586 locus=Lus10007586.g ID=Lus10007586.BGIv1.0 annot-version=v1.0
MDQRQKLRDPLLADGDDDRKPLRGAAGDRFRGFAKWVLEMLMWAIFIVWLTLVFVFPTELGNDLVFKKWTALTKTPLFGITGSLLLLLAGPVLLIALLAA
AHLIISGEGEHHLYQKKSWKYPRPQLWTFPVIVDGPLGVVSATELIGVVLFVGYIVWAFYLYTVRNLALLSSYTLTLPQFSLFLMELTGLRLGMIGMFCL
AFLFLPISRGSMLLRLIDIPVEHATRYHVWLGHITMLLFTLHGLFYVVAWAVQGKLLEEILEWKNIGISNLAGLISLVAGLLMWVTSLNPVRRWNFELFF
YTHQLYIVFVLFLAFHVGDFLFTIAAGGIFLFILDRFLRFCQSRKTVDVVSAKCFPCGTVELTLSKPANLRYNALSFIFIQVRELSLLQWHPFSVSSSPL
EGKNHISVLIKVLGEWTDKLKDTIKINTSTVNNDPASRKITASVEGPYGHESPYHLMYEHLVLVAGGIGISPFMAILSDVFHRVNDGKPCLPREILVVWA
VKKSEEIGLLSAMNLDRICSDLYDRLNLRIHIYVTRESDPPLEEGKVSTEASGVSATRCPFPIGNDMSVLVGTGDNVWSGLYILSSTLGFILSLAMTDAY
YINPYGVTSWWYKGLLLVACMVGSVVVFGGAVVALWHLWGRKASTLEQDDEETMTKVDEGLPNSTPNGTRIVEDLLASSSTSVNYGSRPDFKEIMQSISK
DWGHVDVGVMVCGPPTLQSTVAKEIRAHNFRRSSGETVFHFNSHSFDL
|
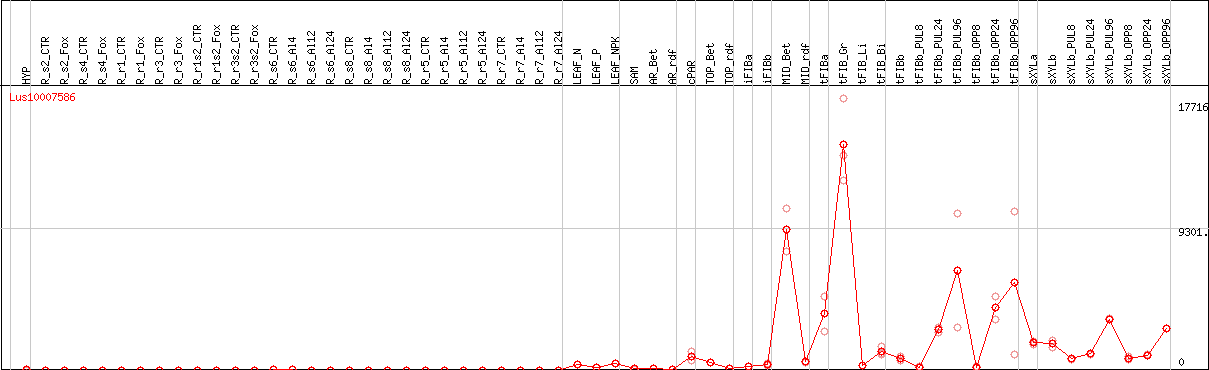
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10007586 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
