Lus10007661 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10007661 pacid=23140312 polypeptide=Lus10007661 locus=Lus10007661.g ID=Lus10007661.BGIv1.0 annot-version=v1.0
ATGTCTTGGGGATTAGGATGGAAGCGGCCGTCGGAGATCTTCCGCTTAAGTCTTAACTATGGCACGGAGGAGTCCTGGGACGATCCAATCCGTACATCGA
GCTCCTCGTCGTGTTCTTCGTCGTCAACGATACCGCCGCTGGATCAGGATGGTGGGTTCCGCGTCGATTTGGATTGGACGGCCGGAGAAGACGAGGATCA
GTTGGCCCTGAGGTTGCTGTCGCAGCTCACGGTGGCACTCCCGATGCCGCAGGATTGTGTGACGGTGGAATTGGCGACTGGAGATGAGGAAGGGGCGGAG
GAAGGAAATGTCAGAGTAGAGATGAATGTGGTGAAGAGGAGGGAGCCGTTGAGAGGGATCACGATCTCGAAGGCCGGCTCTGGACAGCAGAGTGATGGCA
TCGGTGTTTTGACGAGATTACTGAGGTCCAATTTAGCTTCTGCTGGAATTGGGGATGGTTCTGATTTTTCTTGTGCCGATCACTGGAAGACTGTCACCTT
GCTTAACCTATGCGGCTGTGGTTTGTCGACATTGCCGCCTGAGCTACTACAATTGCCAACCCTCGAGAAGCTTTTTCTTGACAATAATAGACTTACTTTT
TTGCCGCCTGAGCTTGGGGAGCTGAAAACTTTGAAAGTTCTCAGTGTTGATTACAACTTGTTGGTTTCTGTTCCTGTGGAATTGAGGCAGTGTGTTCAAC
TCGTTGAGTTGTCATTGGAACACAACAAGCTTGTCCGTCCTCTTCTTGATTTCAGGGCCATGGCCGAGTTACAAATTCTTAGGCTATTTGGCAACCCTCT
GGAATTTCTTCCTGAGATTTTACCACTCCACAAACTTCGCCACTTGACCCTTGCAAATATCAGGATTGTTTCTGATGAAAATTTAAGATCTGTGACAGTG
CAGATAGAGGGGGAGAACAATTCTTATTTTGGAGCATCTAGACATAAGTTAAGTGCATTTTTTTCTCTTATATTCCGCTTCTCATCATGCCATCACTCTC
TGCTAGCATCTGCACTGGCAAAGATAGTGAAAGACCAAGGTAACCGAGCAGTAATTGGTAAAGATGAGAATGCAGTGCGACAGCTCATTAGTATGATCAG
TAGCGACGACCGTTATGTGGTTGAGCAAGCATGTTCTGCACTGTCCACTCTTGCTGGAGATGTTTCTATGGCAATGCAGTTGATTAAATGCGATATCATG
CAGCCTATTGAGACAGTGCTAAGATCTGTTTCCCAGGAAGAAGTCATTTCTGTACTGCAAGTTGTGTCAACGTTAGCTTTTGCATCAGATAGAGTGGCCC
AGAAGATGTTAACCAAGGAATTAATAAGATCATTGAAAGTTCTCTGTGGTCATAAAAACCCAGAGTTGCAAAAGTCGGCTTTGTTTGTGGTGGGTAATCT
GGCGTTCTGTTTGGAAAATAGACGTATTTTGGTTACTTCTGAAAGTCTGCGGGACTTGCTATTGCGGTTGACAGTTACATCTGAACCTCGTGTGAACAAA
GCTGCTGCTCGTGCCTTGGCCATTCTTGGAGAGAATGAAAATTTGAGACGTGCCGTAAAGGGAAGACCTGTGACCAAGCAAGGACTGCGCATCCTATCAA
TGGATGGAGGTGGAATGAAAGGTCTGGCCACCGTACAGATTCTCAAGGCAATTGAAAAGGGAACTGGAAAACGAATACATGAGCTGTTTGACCTTATATG
CGGCACATCAACGGGTGGAATGTTGGCTGTTGCTCTTGGTATAAAGCTAATGTCCCTGGATCAATGCGAGGAAATATACAAAAATCTTGGGAAACTTGTG
TTTGCTGAACCTGTGCCCAAGGATAACGAAGCTGCATCATGGAGAGAAAAGTTAGATCAGCTTTACAAAAGTTCATCCCAGAGTTTTAGAGTTGTCGTAC
ACGGATCCAAACACAGCGCAGATGAGTTTGAGAGACTGTTAAAGGAAATGTGTGCTGATGGGGATGGAGATCTATTAATAGACTCAGCAGTGAAATGCAT
TCCAAAAGTTTTTACCGTCTCAACTTTGGTTAGCACGATGCCGGCTCAGCCATTCATATTTCGCAACTATCAGTACCCAGTTGGGACACCAGAAGTGCCA
ACCACTCCTACAGAGGGATCTGGTATCCCTGTGTTAGGATCACCTTTTGCTGGTAGCCAAATTGGATACAGGCGCAGTGCTTCTGTTGGGAGTTGTAAAC
ACCATGTTTGGCAAGCAATAAGAGCGTCATCTGCTGCACCGTATTATCTTGATGACTATTCTGATGACATAAACCGCTGGCAAGATGGTGCTATTGTGGC
CAATAATCCTACCATATTTGCCATAAGAGAAGCACAGCTTCTCTGGCCTGACACAAGAATCGACTGTCTTGTTTCGGTTGGGTGTGGTTCTGTTCCAACA
AAGCCCCGGAAAGGTGGTTGGCGCTATCTAGACACTGGGCAAGTTCTTATCGAGAGTGCATGCTCCGTGGACCGAGTGGAAGAAGCCTTAAGTACATTAC
TCCCCATGCTTCCTGAGATACAGTATTTTCGTTTCAACCCAGTCGATGAACGCTGCGATATGGAATTGGATGAGACCGACCCTGCTGTATGGCTAAAGTT
GGAAGCATCAGTCGAAGAATACATCCTAGAGAGTGCTGAAGCTTTCAAGAATGTCTGTGAGAGACTGGTGTTGCCATTCCAAAATGATGACAAGTTCTCA
GACCACGTGAAAACTCAGCAACTTCCTAGAGTAAAGACGTTCACTACTGGTGGGAGTGGACCCACTCTCGGTTGGAGGCGTAATGTTCTACTTGTTGAAG
CTCTTCACTGTCCTGATTCAGGAAGAGTTATGCACCATGCCCGGGCACTCGAGTCGTTTTGTGCACGCAACAGCATAAAAATATCTCATGTTCATAGCAC
ATCTGGAATTTCGAAGACGGTACCAACCTCAATCCCATCACCCTTCACCTCACCGCTAATCACCGGAAGCTTTCCTTCAAGCCCACTTCTATACAGTCCC
GACTTTGCCTCCCAAAGAATGGGTCGAATTGAAATGGTTCCACCTTTAAGCTTGGATGGAACTCAATCTCTAAGAGGGGGATCATCGCCACCAATGTCCC
CTGCCGCACACAGACAGTTCTCTTTACCCGTTCAATCATTGCATGAAAAGCTGCAGCATGCACCGCAAGTGGGGATCGTGCACTTAGCCCTTCAGAACGA
TTCATCTGGCTCGATTTTAAGTTGGCAGAATGACGTATTTGTGGTTGCTGAACCCGGAGAACTCGCTGACAAGTTTCTACAAAGTGTGAAGCTAAGTTTA
TTGTCCATGATGCGGTGCCACCGACGGAAGGTAACATCGTTCCTTTCCAAGATATCAACGGTTGCTGATCTCGTTAGCTCCAGACCGTACTTCCAAGTTG
GAAATGTCGTCCACCGTTACATAGGCCGGCAGACACAGGTGATGGAAGATGACCAAGAAATTGCAGCATACATGTTTCGAAGAACGGTTCCTTCAATTCA
TTTGACACCTGATGATGTCCGATGGATGGTCGGAGCTTGGAGAGACCGGATCATCATCTGCACGGGAACTTATGGCCCTTCCCAGACGCTAATCAAAGCC
TTCATGGACTCAGGTGCAAAAGCCGTCATATGTCCTTCCGCAGACCCGATGGACATTCCGGTGACAGCCGCCAGCTTGTCAGGTGACTTCAACTTGGAGA
ACGGCGGGAAGTTCGAGATCGGAGAAGAGGAAGCAGAAGACGAAGACATCGAGCCTCCCAGTCCGGCAAGCGACTGGGAAGACAGTGATCCAGAGAAGGC
CTGTGGGGGGTCGAACTCGATGGGGTTTTGGGACGATGACGAAGAGAAGCTAGCGCAGTTTCTAAGTCAGGTGTATGATTCGTTGTTCCGGGAAGGTGCC
AGCGTGGATGTAGCCTTACGTACTGCTCTGGCTTCACATCGTAGACTGAGGTATTCGTACCATCTCCCCGGTTTGAAACCAACAGCCTCCTGCAGCTCCA
CGCCAGCACTGAATCTCCCCGAGCAGATGATGTCCGCCGCAGCAGCAGCTCCGGTGGCGGTCGGGACACGTGGCACAGTGGGTTCGCTAGTTCGGAAAGA
GATCGACTACTTTACCAGCTTGGACACGACCGGAAGCACTAGTGCTCGGAAATTTGCTGCTCCAGGGAGTATTAATGGTGGTGGTACTGCTACGTCTTCT
TCTAGAAGATTTTGGCCGTGGAGAATTAGCTGGAGAAGAAAGAGATGTGGGAGCAAGTTGCTACATCCGGGGATGTGCGCCACCGTGGATGTTGGTGGTG
ATTCTGGTGTCGGGATCTCCGGCCGGTTCAGTTATAGAGTTCTTAAGCATGATTATGAGTTAGATATGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10007661 pacid=23140312 polypeptide=Lus10007661 locus=Lus10007661.g ID=Lus10007661.BGIv1.0 annot-version=v1.0
MSWGLGWKRPSEIFRLSLNYGTEESWDDPIRTSSSSSCSSSSTIPPLDQDGGFRVDLDWTAGEDEDQLALRLLSQLTVALPMPQDCVTVELATGDEEGAE
EGNVRVEMNVVKRREPLRGITISKAGSGQQSDGIGVLTRLLRSNLASAGIGDGSDFSCADHWKTVTLLNLCGCGLSTLPPELLQLPTLEKLFLDNNRLTF
LPPELGELKTLKVLSVDYNLLVSVPVELRQCVQLVELSLEHNKLVRPLLDFRAMAELQILRLFGNPLEFLPEILPLHKLRHLTLANIRIVSDENLRSVTV
QIEGENNSYFGASRHKLSAFFSLIFRFSSCHHSLLASALAKIVKDQGNRAVIGKDENAVRQLISMISSDDRYVVEQACSALSTLAGDVSMAMQLIKCDIM
QPIETVLRSVSQEEVISVLQVVSTLAFASDRVAQKMLTKELIRSLKVLCGHKNPELQKSALFVVGNLAFCLENRRILVTSESLRDLLLRLTVTSEPRVNK
AAARALAILGENENLRRAVKGRPVTKQGLRILSMDGGGMKGLATVQILKAIEKGTGKRIHELFDLICGTSTGGMLAVALGIKLMSLDQCEEIYKNLGKLV
FAEPVPKDNEAASWREKLDQLYKSSSQSFRVVVHGSKHSADEFERLLKEMCADGDGDLLIDSAVKCIPKVFTVSTLVSTMPAQPFIFRNYQYPVGTPEVP
TTPTEGSGIPVLGSPFAGSQIGYRRSASVGSCKHHVWQAIRASSAAPYYLDDYSDDINRWQDGAIVANNPTIFAIREAQLLWPDTRIDCLVSVGCGSVPT
KPRKGGWRYLDTGQVLIESACSVDRVEEALSTLLPMLPEIQYFRFNPVDERCDMELDETDPAVWLKLEASVEEYILESAEAFKNVCERLVLPFQNDDKFS
DHVKTQQLPRVKTFTTGGSGPTLGWRRNVLLVEALHCPDSGRVMHHARALESFCARNSIKISHVHSTSGISKTVPTSIPSPFTSPLITGSFPSSPLLYSP
DFASQRMGRIEMVPPLSLDGTQSLRGGSSPPMSPAAHRQFSLPVQSLHEKLQHAPQVGIVHLALQNDSSGSILSWQNDVFVVAEPGELADKFLQSVKLSL
LSMMRCHRRKVTSFLSKISTVADLVSSRPYFQVGNVVHRYIGRQTQVMEDDQEIAAYMFRRTVPSIHLTPDDVRWMVGAWRDRIIICTGTYGPSQTLIKA
FMDSGAKAVICPSADPMDIPVTAASLSGDFNLENGGKFEIGEEEAEDEDIEPPSPASDWEDSDPEKACGGSNSMGFWDDDEEKLAQFLSQVYDSLFREGA
SVDVALRTALASHRRLRYSYHLPGLKPTASCSSTPALNLPEQMMSAAAAAPVAVGTRGTVGSLVRKEIDYFTSLDTTGSTSARKFAAPGSINGGGTATSS
SRRFWPWRISWRRKRCGSKLLHPGMCATVDVGGDSGVGISGRFSYRVLKHDYELDM
|
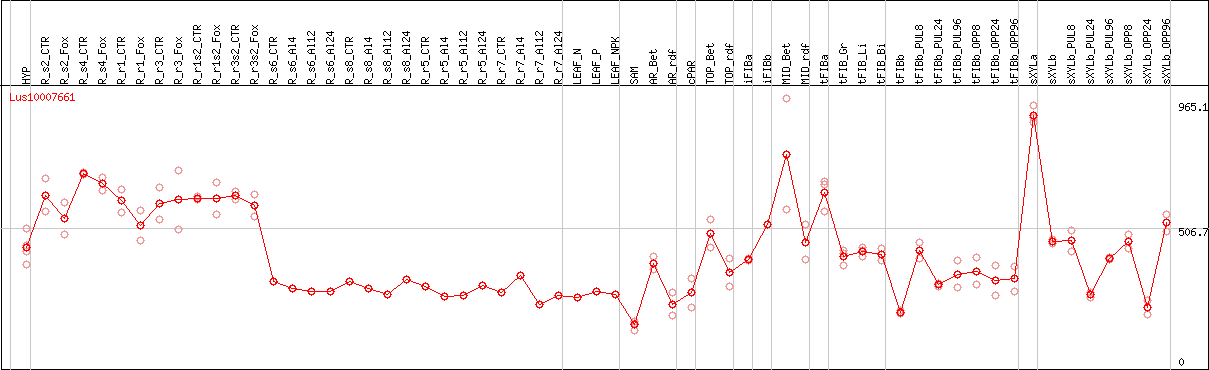
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10007661 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
