External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G15130 323 / 6e-106
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G68930 214 / 5e-64
pentatricopeptide (PPR) repeat-containing protein (.1)
AT4G15720 197 / 1e-58
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G33680 197 / 7e-58
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G18750 197 / 3e-57
DOT4
DEFECTIVELY ORGANIZED TRIBUTARIES 4, Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT4G14050 194 / 3e-57
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT4G33170 197 / 4e-57
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G20730 191 / 1e-56
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G13650 195 / 2e-56
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G11290 193 / 3e-56
CRR22
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10030124
536 / 0
AT3G15130 613 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10015898
210 / 1e-62
AT2G13600 899 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10009269
208 / 2e-62
AT2G13600 874 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10004892
202 / 3e-59
AT2G27610 998 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10027534
195 / 3e-58
AT3G20730 474 / 2e-162
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10002054
199 / 9e-58
AT1G74600 930 / 0.0
organelle transcript processing 87, pentatricopeptide (PPR) repeat-containing protein (.1)
Lus10036029
192 / 2e-57
AT5G27110 577 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10040504
193 / 5e-57
AT3G13770 904 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10024220
195 / 1e-56
AT1G74600 924 / 0.0
organelle transcript processing 87, pentatricopeptide (PPR) repeat-containing protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G244500
387 / 6e-131
AT3G15130 891 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.010G136200
218 / 1e-65
AT1G68930 997 / 0.0
pentatricopeptide (PPR) repeat-containing protein (.1)
Potri.013G006700
205 / 1e-61
AT1G56570 732 / 0.0
PENTATRICOPEPTIDE REPEAT PROTEIN FOR GERMINATION ON NaCl, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.007G104700
206 / 3e-61
AT2G13600 834 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.013G006800
205 / 5e-61
AT2G33680 780 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.007G068900
202 / 7e-61
AT3G20730 561 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.003G191000
203 / 8e-60
AT1G11290 1178 / 0.0
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.005G010900
194 / 9e-58
AT1G56570 666 / 0.0
PENTATRICOPEPTIDE REPEAT PROTEIN FOR GERMINATION ON NaCl, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.004G111300
197 / 2e-57
AT3G02330 1077 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.007G021700
193 / 2e-56
AT3G47840 796 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10007683 pacid=23165681 polypeptide=Lus10007683 locus=Lus10007683.g ID=Lus10007683.BGIv1.0 annot-version=v1.0
ATGATAGCAGCGTACAATCTTACAGGGAAGGGTAAGAAAGCTCTGGGTGTGTTCAGGAAAATGAGAGAAGATGGACAAGTTCCAGATGAGTTTACACTTA
CAAGTGCCTTGAAGGCCTGTACCAAGCTCGGTGCTGCTAGAGAAGCAGCGCAGATTCATGCATCATCGGTCCCTGGTGGCTTTCATTATTCCGACAAGGC
TATTGTTGCTGGTTACTTGATAGATTCGTATGTGAAATGTGGGGAGCTGTTGCAAGCTCGACGAGTATTTGATCAGATTGAGAAGAAACATATAATATCG
TGGAGTGCACTGATTCTCGGCTATGCTCAAGAAGAAAAGTTAGAAGAAGCCATGGAGCTGTTTAAGCTGCTAAGACTTGGTACAATTGAAGTTGACGAAT
ATGTGCTGTCCAGTATGATGGCCATCTTTGCAGATTTTGCCCTGGTGGAGCAAGGCAGGCAAGTTCATGGTTTTAGCCTTAAGATCCCCTCTGGTTTGTG
GATTTCGATATGTAATTCAGCTCTAGATATGTATCTCAAATGTGGGATGATTGAAGATGCTAAGAAACTGTTTGGTGCAATGTCAGCGAGAGACGTGGTT
TCCTGGACTGTAATGATCACTGGCTACGGAAAGCATGGTCTTGGGAAAGAGGCAGTTTATCTCTTCAACAGAATGCTATCGCAAAATATTAAGCCTGATG
ATGTTATCTACTTGGCCTTTCTCTTAGCCTGCAGCCATTCAGGGTTGGTGGAAGAAGGTGAAGAGTACTTATCCAAGTTATGCAGCGATGAGGGGATCAA
ACTTAGAGTTGAGCATTATGCCTGCATGGTTGATCTCGTGGGCCGTGTCCGAAGATTAAACTAA
AA sequence
>Lus10007683 pacid=23165681 polypeptide=Lus10007683 locus=Lus10007683.g ID=Lus10007683.BGIv1.0 annot-version=v1.0
MIAAYNLTGKGKKALGVFRKMREDGQVPDEFTLTSALKACTKLGAAREAAQIHASSVPGGFHYSDKAIVAGYLIDSYVKCGELLQARRVFDQIEKKHIIS
WSALILGYAQEEKLEEAMELFKLLRLGTIEVDEYVLSSMMAIFADFALVEQGRQVHGFSLKIPSGLWISICNSALDMYLKCGMIEDAKKLFGAMSARDVV
SWTVMITGYGKHGLGKEAVYLFNRMLSQNIKPDDVIYLAFLLACSHSGLVEEGEEYLSKLCSDEGIKLRVEHYACMVDLVGRVRRLN
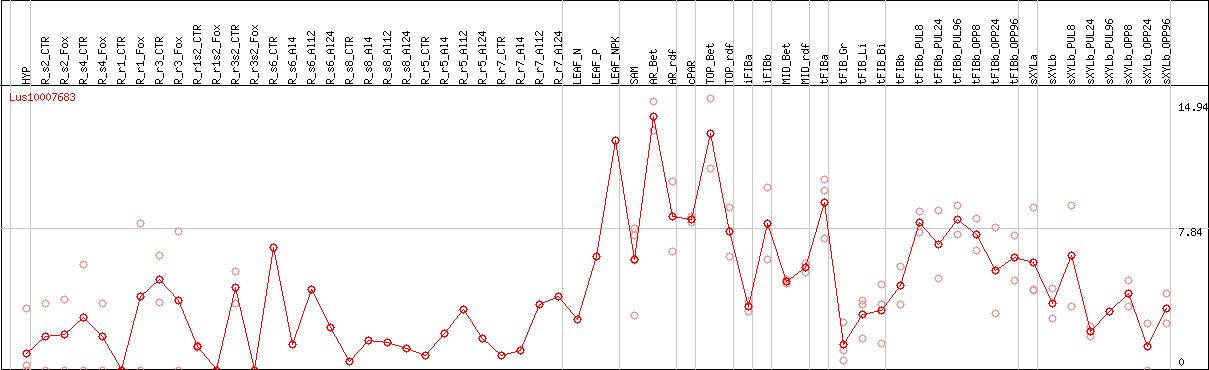
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10007683 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.